No project description provided
Project description
SynOmicsBench
SynOmicsBench is a unified benchmarking framework for synthetic data generation (SDG) for clinical transcriptomic cancer cohorts.
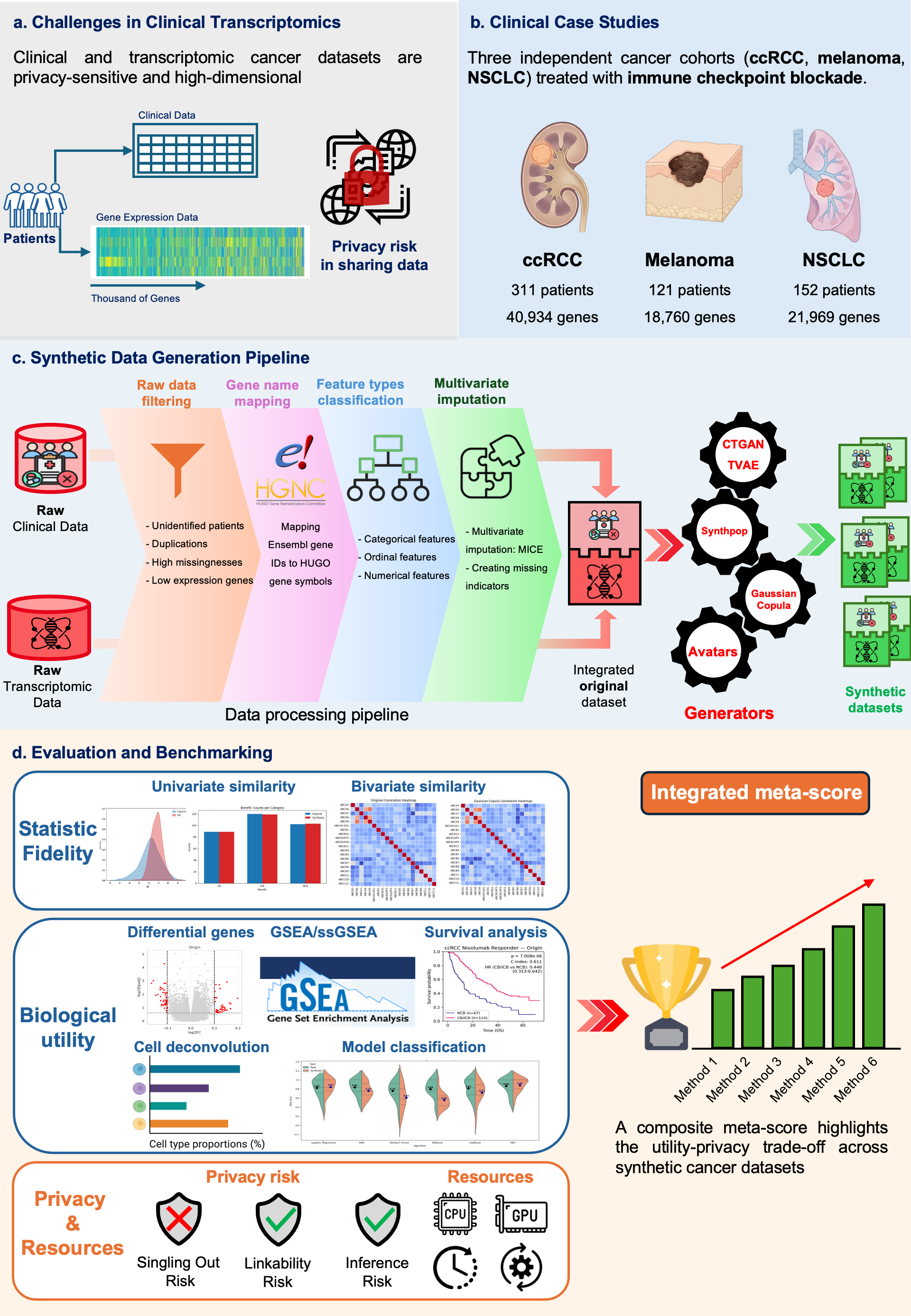
Achieving a trade-off between biological utility and patient privacy is critical for secure data sharing when applying transcriptomic clinical datasets to artificial intelligence in precision oncology. Here, we present the SynOmicsBench framework. SynOmicsBench combines standardized preprocessing with multidimensional evaluation, prioritizing downstream biological validation alongside statistical fidelity and attack-based privacy assessment. This work provides a reproducible decision-support tool for method selection and promotes biologically informed, privacy-aware adoption of synthetic data in precision oncology.
🔬 Framework Overview
SynOmicsBench compares synthetic data generation methods using a standardized pipeline that combines:
- Standardized Preprocessing: Automated data filtering, harmonization, and integration.
- Multidimensional Evaluation: Assessing Statistical Fidelity, Downstream Biological Utility, and Privacy Risk.
- State-of-the-Art SDG Methods: Native support for CTGAN, TVAE, Gaussian Copula, Synthpop, and Avatars (K5/K10).
🛠 Installation
SynOmicsBench can be installed in three different ways depending on your environment. Python 3.12+ is required.
Option 1: From PyPI (Recommended)
pip install synomicsbench
Option 2: From Source (GitHub)
We recommend using uv for fast, reliable dependency management. This method uses the provided uv.lock file to ensure reproducible installations.
git clone https://github.com/trinhthechuong/SynOmicsBench.git
cd SynOmicsBench
# With uv (Fastest)
uv sync
source .venv/bin/activate
# Or with traditional pip
pip install -e .
Option 3: Pre-built Container (Apptainer/Singularity)
For HPC environments or reproducible workflows, you can pull our fully prepared Apptainer container which contains all dependencies (including heavy ML frameworks and R):
# Pull the latest SynOmicsBench container
apptainer pull synomicsbench.sif oras://ghcr.io/trinhthechuong/synomicsbench:latest
# Verify the container is working and the package is ready
apptainer exec synomicsbench.sif python -c "import synomicsbench; print('OK: SynOmicsBench is ready!')"
(To use the container for your scripts, simply mount your directories via --bind and run your Python scripts using apptainer exec)
🚀 Quick Start
Here is a brief example of how to generate synthetic data with Gaussian Copula and evaluate its statistical fidelity:
import pandas as pd
from synomicsbench.synthesizer.GaussianCopulasynthesizer import GaussianCopulasynthesizer
from synomicsbench.processing.metadata import MetaData
from synomicsbench.metrics.fidelity.UnivariateSimilarity import UnivariateSimilarity
# 1. Load Data & Prepare Metadata
original_data = pd.read_csv("your_clinical_transcriptomic_data.csv")
ordinal_features = ["Mstage", "Tx_Start_ECOG", "numPriorTherapies"]
metadata = MetaData.get_metadata(data=original_data, ordinal_features=ordinal_features)
# 2. Generate Synthetic Data
synth = GaussianCopulasynthesizer(output_path="./results", metadata=metadata)
synthetic_data = synth.generate(
data=original_data,
n_samples=original_data.shape[0]
)
# 3. Evaluate Fidelity
evaluator = UnivariateSimilarity(output_dir="./evaluation_results")
score = evaluator.get_univariate_score(
original_data=original_data,
synthetic_data=synthetic_data,
metadata=metadata,
save=True
)
print(f"Overall Fidelity Score: {score:.4f}")
📚 Documentation
For complete API references, tutorials, and full benchmarking results, check out the SynOmicsBench Official Documentation:
- Getting Started: Step-by-step setup guides.
- Preprocessing Pipeline: Harmonizing multimodal data.
- SDG Methods: Deep dive into generation models.
- Evaluation Framework: Understand our metrics for Privacy and Biological signal preservation.
📝 Citation
If you use SynOmicsBench in your research, please cite:
Trinh, T. C., Woillard, J. B., Uguzzoni, G., & Battail, C. (2024). A unified benchmark of synthetic data generation for clinical and transcriptomic cancer data. (Manuscript in preparation)
📄 License
This project is open-sourced under the MIT License.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file synomicsbench-1.0.2.tar.gz.
File metadata
- Download URL: synomicsbench-1.0.2.tar.gz
- Upload date:
- Size: 283.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
585b03c36d17ddfb091c3fd0578b9e92901a036ad3eb28e1b1b7381a73820e05
|
|
| MD5 |
a7b7625d52faa1ced7400c6be0eecfac
|
|
| BLAKE2b-256 |
1dd937cc82ccdf69a7693abcdd1061aed8f38d125aafd892e58108093aba20da
|
File details
Details for the file synomicsbench-1.0.2-py3-none-any.whl.
File metadata
- Download URL: synomicsbench-1.0.2-py3-none-any.whl
- Upload date:
- Size: 104.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3ff1547b99f4c6afca394e8b374a7b50a25d7cef0002727759efa0068b0b6ab6
|
|
| MD5 |
ab311381c36f8924ec1f82b63806751d
|
|
| BLAKE2b-256 |
7d139410497d4372064498ddb0b9212534a03f922f5ac9dd817aa3ec72693670
|