Synthetic data generator and evaluator!
Project description
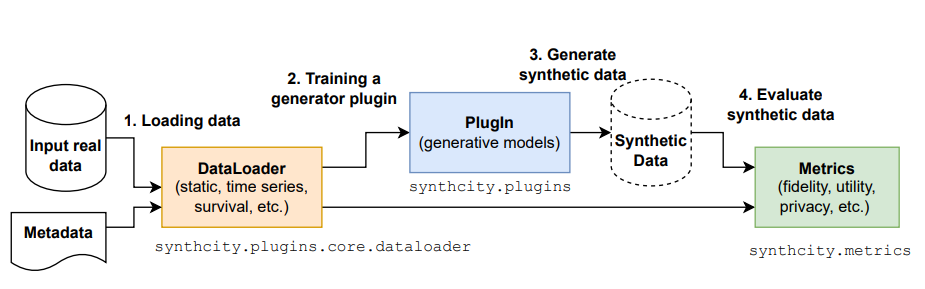
synthcity
A library for generating and evaluating synthetic tabular data.
Features:
- :key: Easy to extend pluginable architecture.
- :cyclone: Several evaluation metrics for correctness and privacy.
- :fire: Several reference models, by type:
- General purpose: GAN-based (AdsGAN, CTGAN, PATEGAN, DP-GAN),VAE-based(TVAE, RTVAE), Normalizing flows, Bayesian Networks(PrivBayes, BN), Random Forrest (arfpy), LLM-based (GReaT).
- Time Series & Time-Series Survival generators: TimeGAN, FourierFlows, TimeVAE.
- Static Survival Analysis: SurvivalGAN, SurVAE.
- Privacy-focused: DECAF, DP-GAN, AdsGAN, PATEGAN, PrivBayes.
- Domain adaptation: RadialGAN.
- Images: Image ConditionalGAN, Image AdsGAN.
- :book: Read the docs !
- :airplane: Checkout the tutorials!
Please note: synthcity does not handle missing data and so these values must be imputed first HyperImpute can be used to do this.
:rocket: Installation
The library can be installed from PyPI using
$ pip install synthcity
or from source, using
$ pip install .
Other library extensions:
- Install the library with unit-testing support
pip install synthcity[testing]
- Install the library with GOGGLE support
pip install synthcity[goggle]
- Install the library with ALL the extensions
pip install synthcity[all]
:boom: Sample Usage
Generic data
- List the available general-purpose generators
from synthcity.plugins import Plugins
Plugins(categories=["generic", "privacy"]).list()
- Load and train a tabular generator
from sklearn.datasets import load_diabetes
from synthcity.plugins import Plugins
X, y = load_diabetes(return_X_y=True, as_frame=True)
X["target"] = y
syn_model = Plugins().get("adsgan")
syn_model.fit(X)
- Generate new synthetic tabular data
syn_model.generate(count = 10)
- Benchmark the quality of the plugins
# third party
from sklearn.datasets import load_diabetes
# synthcity absolute
from synthcity.benchmark import Benchmarks
from synthcity.plugins.core.constraints import Constraints
from synthcity.plugins.core.dataloader import GenericDataLoader
X, y = load_diabetes(return_X_y=True, as_frame=True)
X["target"] = y
loader = GenericDataLoader(X, target_column="target", sensitive_columns=["sex"])
score = Benchmarks.evaluate(
[
(f"example_{model}", model, {}) # testname, plugin name, plugin args
for model in ["adsgan", "ctgan", "tvae"]
],
loader,
synthetic_size=1000,
metrics={"performance": ["linear_model"]},
repeats=3,
)
Benchmarks.print(score)
Static Survival analysis
- List the available generators dedicated to survival analysis
from synthcity.plugins import Plugins
Plugins(categories=["generic", "privacy", "survival_analysis"]).list()
- Generate new data
from lifelines.datasets import load_rossi
from synthcity.plugins.core.dataloader import SurvivalAnalysisDataLoader
from synthcity.plugins import Plugins
X = load_rossi()
data = SurvivalAnalysisDataLoader(
X,
target_column="arrest",
time_to_event_column="week",
)
syn_model = Plugins().get("survival_gan")
syn_model.fit(data)
syn_model.generate(count=10)
Time series
- List the available generators
from synthcity.plugins import Plugins
Plugins(categories=["generic", "privacy", "time_series"]).list()
- Generate new data
# synthcity absolute
from synthcity.plugins import Plugins
from synthcity.plugins.core.dataloader import TimeSeriesDataLoader
from synthcity.utils.datasets.time_series.google_stocks import GoogleStocksDataloader
static_data, temporal_data, horizons, outcome = GoogleStocksDataloader().load()
data = TimeSeriesDataLoader(
temporal_data=temporal_data,
observation_times=horizons,
static_data=static_data,
outcome=outcome,
)
syn_model = Plugins().get("timegan")
syn_model.fit(data)
syn_model.generate(count=10)
Images
Note : The architectures used for generators are not state-of-the-art. For other architectures, consider extending the suggest_image_generator_discriminator_arch method from the convnet.py module.
- List the available generators
from synthcity.plugins import Plugins
Plugins(categories=["images"]).list()
- Generate new data
from synthcity.plugins import Plugins
from synthcity.plugins.core.dataloader import ImageDataLoader
from torchvision import datasets
dataset = datasets.MNIST(".", download=True)
loader = ImageDataLoader(dataset).sample(100)
syn_model = Plugins().get("image_cgan")
syn_model.fit(loader)
syn_img, syn_labels = syn_model.generate(count=10).unpack().numpy()
print(syn_img.shape)
Serialization
- Using save/load methods
from synthcity.utils.serialization import save, load
from synthcity.plugins import Plugins
syn_model = Plugins().get("adsgan")
buff = save(syn_model)
reloaded = load(buff)
assert syn_model.name() == reloaded.name()
- Saving and loading models from disk
from sklearn.datasets import load_diabetes
from synthcity.utils.serialization import save_to_file, load_from_file
from synthcity.plugins import Plugins
X, y = load_diabetes(return_X_y=True, as_frame=True)
X["target"] = y
syn_model = Plugins().get("adsgan", n_iter=10)
syn_model.fit(X)
save_to_file('./adsgan_10_epochs.pkl', syn_model)
reloaded = load_from_file('./adsgan_10_epochs.pkl')
assert syn_model.name() == reloaded.name()
- Using the Serializable interface
from synthcity.plugins import Plugins
syn_model = Plugins().get("adsgan")
buff = syn_model.save()
reloaded = Plugins().load(buff)
assert syn_model.name() == reloaded.name()
📓 Tutorials
Tutorial 0: Getting started with tabular data
Tutorial 1: Writing a new plugin
Tutorial 2: Benchmarking models
Tutorial 3: Generating Survival Analysis data
Tutorial 4: Generating Time Series
Tutorial 5: Generating Data with Differential Privacy Guarantees
Tutorial 6: Practice using Custom Time series data
Tutorial 7: Image generation
Tutorial 8: hyperparameter optimization
Tutorial 9: dealing with missing data
🔑 Methods
Bayesian methods
| Method | Description | Reference |
|---|---|---|
| bayesian_network | The method represents a set of random variables and their conditional dependencies via a directed acyclic graph (DAG), and uses it to sample new data points | pgmpy |
| privbayes | A differentially private method for releasing high-dimensional data. | PrivBayes: Private Data Release via Bayesian Networks |
Generative adversarial networks(GANs)
| Method | Description | Reference |
|---|---|---|
| adsgan | A conditional GAN framework that generates synthetic data while minimize patient identifiability that is defined based on the probability of re-identification given the combination of all data on any individual patient | Anonymization Through Data Synthesis Using Generative Adversarial Networks (ADS-GAN) |
| pategan | The methos uses the Private Aggregation of Teacher Ensembles (PATE) framework and applies it to GANs, allowing to tightly bound the influence of any individual sample on the model, resulting in tight differential privacy guarantees and thus an improved performance over models with the same guarantees. | PATE-GAN: Generating Synthetic Data with Differential Privacy Guarantees |
| ctgan | A conditional generative adversarial network which can handle tabular data. | Modeling Tabular data using Conditional GAN |
Variational autoencoders(VAE)
| Method | Description | Reference |
|---|---|---|
| tvae | A conditional VAE network which can handle tabular data. | Modeling Tabular data using Conditional GAN |
| rtvae | A robust variational autoencoder with β divergence for tabular data (RTVAE) with mixed categorical and continuous features. | Robust Variational Autoencoder for Tabular Data with β Divergence |
Normalizing Flows
| Method | Description | Reference |
|---|---|---|
| nflow | Normalizing Flows are generative models which produce tractable distributions where both sampling and density evaluation can be efficient and exact. | Neural Spline Flows |
Graph neural networks
| Method | Description | Reference |
|---|---|---|
| goggle | GOGGLE: Generative Modelling for Tabular Data by Learning Relational Structure | Paper |
Diffusion models
| Method | Description | Reference |
|---|---|---|
| ddpm | TabDDPM: Modelling Tabular Data with Diffusion Models. | Paper |
Random Forest models
| Method | Description | Reference |
|---|---|---|
| arfpy | Adversarial Random Forests for Density Estimation and Generative Modeling | Paper |
LLM-based models
| Method | Description | Reference |
|---|---|---|
| GReaT | Language Models are Realistic Tabular Data Generators | Paper |
Static Survival analysis methods
| Method | Description | Reference |
|---|---|---|
| survival_gan | SurvivalGAN is a generative model that can handle survival data by addressing the imbalance in the censoring and time horizons, using a dedicated mechanism for approximating time to event/censoring from the input and survival function. | --- |
| survival_ctgan | SurvivalGAN version using CTGAN | --- |
| survae | SurvivalGAN version using VAE | --- |
| survival_nflow | SurvivalGAN version using normalizing flows | --- |
Time-Series and Time-Series Survival Analysis methods
| Method | Description | Reference |
|---|---|---|
| timegan | TimeGAN is a framework for generating realistic time-series data that combines the flexibility of the unsupervised paradigm with the control afforded by supervised training. Through a learned embedding space jointly optimized with both supervised and adversarial objectives, the network adheres to the dynamics of the training data during sampling. | Time-series Generative Adversarial Networks |
| fflows | FFlows is an explicit likelihood model based on a novel class of normalizing flows that view time-series data in the frequency-domain rather than the time-domain. The method uses a discrete Fourier transform (DFT) to convert variable-length time-series with arbitrary sampling periods into fixed-length spectral representations, then applies a (data-dependent) spectral filter to the frequency-transformed time-series. | Generative Time-series Modeling with Fourier Flows |
Privacy & Fairness
| Method | Description | Reference |
|---|---|---|
| decaf | Machine learning models have been criticized for reflecting unfair biases in the training data. Instead of solving this by introducing fair learning algorithms directly, DEACF focuses on generating fair synthetic data, such that any downstream learner is fair. Generating fair synthetic data from unfair data - while remaining truthful to the underlying data-generating process (DGP) - is non-trivial. DECAF is a GAN-based fair synthetic data generator for tabular data. With DECAF, we embed the DGP explicitly as a structural causal model in the input layers of the generator, allowing each variable to be reconstructed conditioned on its causal parents. This procedure enables inference time debiasing, where biased edges can be strategically removed to satisfy user-defined fairness requirements. | DECAF: Generating Fair Synthetic Data Using Causally-Aware Generative Networks |
| privbayes | A differentially private method for releasing high-dimensional data. | PrivBayes: Private Data Release via Bayesian Networks |
| dpgan | Differentially Private GAN | Differentially Private Generative Adversarial Network |
| adsgan | A conditional GAN framework that generates synthetic data while minimize patient identifiability that is defined based on the probability of re-identification given the combination of all data on any individual patient | Anonymization Through Data Synthesis Using Generative Adversarial Networks (ADS-GAN) |
| pategan | The methos uses the Private Aggregation of Teacher Ensembles (PATE) framework and applies it to GANs, allowing to tightly bound the influence of any individual sample on the model, resulting in tight differential privacy guarantees and thus an improved performance over models with the same guarantees. | PATE-GAN: Generating Synthetic Data with Differential Privacy Guarantees |
Domain adaptation
| Method | Description | Reference |
|---|---|---|
| radialgan | Training complex machine learning models for prediction often requires a large amount of data that is not always readily available. Leveraging these external datasets from related but different sources is, therefore, an essential task if good predictive models are to be built for deployment in settings where data can be rare. RadialGAN is an approach to the problem in which multiple GAN architectures are used to learn to translate from one dataset to another, thereby allowing to augment the target dataset effectively and learning better predictive models than just the target dataset. | RadialGAN: Leveraging multiple datasets to improve target-specific predictive models using Generative Adversarial Networks |
Images
| Method | Description | Reference |
|---|---|---|
| image_cgan | Conditional GAN for generating images | --- |
| image_adsgan | The AdsGAN method adapted for image generation | --- |
Debug methods
| Method | Description | Reference |
|---|---|---|
| marginal_distributions | A differentially private method that samples from the marginal distributions of the training set | --- |
| uniform_sampler | A differentially private method that uniformly samples from the [min, max] ranges of each column. | --- |
| dummy_sampler | Resample data points from the training set | --- |
:zap: Evaluation metrics
The following table contains the available evaluation metrics:
- Sanity checks
| Metric | Description | Values |
|---|---|---|
| data_mismatch | Average number of columns with datatype(object, real, int) mismatch between the real and synthetic data | 0: no datatype mismatch. 1: complete data type mismatch between the datasets. |
| common_rows_proportion | The proportion of rows in the real dataset leaked in the synthetic dataset. | 0: there are no common rows between the real and synthetic datasets. 1: all the rows in the real dataset are leaked in the synthetic dataset. |
| nearest_syn_neighbor_distance | Average distance from the real data to the closest neighbor in the synthetic data | 0: all the real rows are leaked in the synthetic dataset. 1: all the synthetic rows are far away from the real dataset. |
| close_values_probability | The probability of close values between the real and synthetic data. | 0: there is no chance to have synthetic rows similar to the real. 1 means that all the synthetic rows are similar to some real rows. |
| distant_values_probability | Average distance from the real data to the closest neighbor in the synthetic data | 0: no chance to have rows in the synthetic far away from the real data. 1: all the synthetic datapoints are far away from the real data. |
- Statistical tests
| Metric | Description | Values |
|---|---|---|
| inverse_kl_divergence | The average inverse of the Kullback–Leibler Divergence | 0: the datasets are from different distributions. 1: the datasets are from the same distribution. |
| ks_test | The Kolmogorov-Smirnov test | 0: the distributions are totally different. 1: the distributions are identical. |
| chi_squared_test | The p-value. A small value indicates that we can reject the null hypothesis and that the distributions are different. | 0: the distributions are different 1: the distributions are identical. |
| max_mean_discrepancy | Empirical maximum mean discrepancy. | 0: The distributions are the same. 1: The distributions are totally different. |
| jensenshannon_dist | The Jensen-Shannon distance (metric) between two probability arrays. This is the square root of the Jensen-Shannon divergence. | 0: The distributions are the same. 1: The distributions are totally different. |
| wasserstein_dist | Wasserstein Distance is a measure of the distance between two probability distributions. | 0: The distributions are the same. |
| prdc | Computes precision, recall, density, and coverage given two manifolds. | --- |
| alpha_precision | Evaluate the alpha-precision, beta-recall, and authenticity scores. | --- |
| survival_km_distance | The distance between two Kaplan-Meier plots(survival analysis). | --- |
| fid | The Frechet Inception Distance (FID) calculates the distance between two distributions of images. | --- |
- Synthetic Data quality
| Metric | Description | Values |
|---|---|---|
| performance.xgb | Train an XGBoost classifier/regressor/survival model on real data(gt) and synthetic data(syn), and evaluate the performance on the test set. | 1 for ideal performance, 0 for worst performance |
| performance.linear | Train a Linear classifier/regressor/survival model on real data(gt) and the synthetic data and evaluate the performance on test data. | 1 for ideal performance, 0 for worst performance |
| performance.mlp | Train a Neural Net classifier/regressor/survival model on the real data and the synthetic data and evaluate the performance on test data. | 1 for ideal performance, 0 for worst performance |
| performance.feat_rank_distance | Train a model on the synthetic data and a model on the real data. Compute the feature importance of the models on the same test data, and compute the rank distance between the importance(kendalltau or spearman) | 1: similar ranks in the feature importance. 0: uncorrelated feature importance |
| detection_gmm | Train a GaussianMixture model to differentiate the synthetic data from the real data. | 0: The datasets are indistinguishable. 1: The datasets are totally distinguishable. |
| detection_xgb | Train an XGBoost model to differentiate the synthetic data from the real data. | 0: The datasets are indistinguishable. 1: The datasets are totally distinguishable. |
| detection_mlp | Train a Neural net to differentiate the synthetic data from the real data. | 0: The datasets are indistinguishable. 1: The datasets are totally distinguishable. |
| detection_linear | Train a Linear model to differentiate the synthetic data from the real data. | 0: The datasets are indistinguishable. 1: The datasets are totally distinguishable. |
- Privacy metrics
Quasi-identifiers : pieces of information that are not of themselves unique identifiers, but are sufficiently well correlated with an entity that they can be combined with other quasi-identifiers to create a unique identifier.
| Metric | Description | Values |
|---|---|---|
| k_anonymization | The minimum value k which satisfies the k-anonymity rule: each record is similar to at least another k-1 other records on the potentially identifying variables. | Reported on both the real and synthetic data. |
| l_diversity | The minimum value l which satisfies the l-diversity rule: every generalized block has to contain at least l different sensitive values. | Reported on both the real and synthetic data. |
| kmap | The minimum value k which satisfies the k-map rule: every combination of values for the quasi-identifiers appears at least k times in the reidentification(synthetic) dataset. | Reported on both the real and synthetic data. |
| delta_presence | The maximum re-identification risk for the real dataset from the synthetic dataset. | 0 for no risk. |
| identifiability_score | The re-identification score on the real dataset from the synthetic dataset. | --- ] |
| sensitive_data_reidentification_xgb | Sensitive data prediction from the quasi-identifiers using an XGBoost. | 0 for no risk. |
| sensitive_data_reidentification_mlp | Sensitive data prediction from the quasi-identifiers using a Neural Net. | 0 for no risk. |
:mag: Use-cases of synthetic data beyond fidelity & privacy
The following table contains research projects and papers that have been completed using Synthcity. Feel free to explore them and be inspired to use Synthcity in your own research project!
| Project | Description | Code | Paper |
|---|---|---|---|
| Synthetic data capturing nuances of real data | Ensuring that synthetic data mirrors the complex nuances of real-world data is a challenging task. This paper addresses this issue by exploring the potential of integrating data-centric AI techniques which profile the data to guide the synthetic data generation process. | Code | NeurIPS 2023 paper |
| Model Evaluation/Testing | This paper aim to improve model evaluation with synthetic test data | Code | NeurIPS 2023 paper |
| Generative uncertainty | This paper explores how the generative process affects the downstream ML task. | Code | ICML 2023 paper |
| Benchmarking Synthcity | This paper employs synthcity's benchmarking pipeline to compare different SOTA methods across multiple openml datasets. The paper aims to introduce the wider ML community to Synthcity. | Code | NeurIPS 2023 paper |
:hammer: Tests
Install the testing dependencies using
pip install .[testing]
The tests can be executed using
pytest -vsx
Contributing to Synthcity
We want to make contributing to Synthcity is as easy and transparent as possible. We hope to collaborate with as many people as we can.
Development installation
First create a new environment. It is recommended that you use conda. This can be done as follows:
conda create -n your-synthcity-env python=3.9
conda activate your-synthcity-env
Python versions 3.7, 3.8, 3.9, and 3.10 are all compatible, but it is best to use the most up to date version you can, as some models may not support older python versions.
To get the development installation with all the necessary dependencies for linting, testing, auto-formatting, and pre-commit etc. run the following:
git clone https://github.com/vanderschaarlab/synthcity.git
cd synthcity
pip install -e .[testing]
Please check that the pre-commit is properly installed for the repository, by running:
pre-commit run --all
This checks that you are set up properly to contribute, such that you will match the code style in the rest of the project. This is covered in more detail below.
Our Development Process
Code Style
We believe that having a consistent code style is incredibly important. Therefore Synthcity imposes certain rules on the code that is contributed and the automated tests will not pass, if the style is not adhered to. These tests passing is a requirement for a contribution being merged. However, we make adhering to this code style as simple as possible. First, all the libraries required to produce code that is compatible with Synthcity's Code Style are installed in the step above when you set up the development environment. Secondly, these libraries are all triggered by pre-commit, so once you are set-up, you don't need to do anything. When you run git commit, any simple changes to enforce the style will run automatically and other required changes are explained in the stdout for you to go through and fix.
Synthcity uses the black and flake8 code formatter to enforce a common code style across the code base. No additional configuration should be needed (see the black documentation for advanced usage).
Also, Synthcity uses isort to sort imports alphabetically and separate into sections.
Type Hints
Synthcity is fully typed using python 3.7+ type hints. This is enforced for contributions by mypy, which is a static type-checker.
Tests
To run the tests, you can either use pytest (again, installed with the testing extra above).
The following testing command is good for checking your code,as it skips the tests that take a long time to run.
pytest -vvvsx -m "not slow" --durations=50
But the full test suite can be run with the following command.
pytest -vvvs --durations=50
Some plugins may be included in the library as extras, the associated tests for these need to be run separately, e.g. the goggle plugin can be tested with the below command:
pytest -vvvs -k goggle --durations=50
Pull Requests
We actively welcome your pull requests.
- Fork the repo and create your branch from
main. - If you have added code that should be tested, add tests in the same style as those already present in the repo.
- If you have changed APIs, document the API change in the PR.
- Ensure the test suite passes.
- Make sure your code passes the pre-commit, this will be required in order to commit and push, if you have properly installed pre-commit, which is included in the testing extra.
Issues
We use GitHub issues to track public bugs. Please ensure your description is clear and has sufficient instructions to be able to reproduce the issue.
License
By contributing to Synthcity, you agree that your contributions will be licensed under the LICENSE file in the root directory of this source tree. You should therefore, make sure that if you have introduced any dependencies that they also are covered by a license that allows the code to be used by the project and is compatible with the license in the root directory of this project.
Citing
If you use this code, please cite the associated paper:
@misc{https://doi.org/10.48550/arxiv.2301.07573,
doi = {10.48550/ARXIV.2301.07573},
url = {https://arxiv.org/abs/2301.07573},
author = {Qian, Zhaozhi and Cebere, Bogdan-Constantin and van der Schaar, Mihaela},
keywords = {Machine Learning (cs.LG), Artificial Intelligence (cs.AI), FOS: Computer and information sciences, FOS: Computer and information sciences},
title = {Synthcity: facilitating innovative use cases of synthetic data in different data modalities},
year = {2023},
copyright = {Creative Commons Attribution 4.0 International}
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distributions
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file synthcity-0.2.12-py3-none-macosx_10_14_x86_64.whl.
File metadata
- Download URL: synthcity-0.2.12-py3-none-macosx_10_14_x86_64.whl
- Upload date:
- Size: 430.8 kB
- Tags: Python 3, macOS 10.14+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.11.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a8b12118a957df1d15072d58f829f800e37f54d466d8250f8238c9e8eedf1ab9
|
|
| MD5 |
6bafbc71032c08c3cd455960548100ab
|
|
| BLAKE2b-256 |
fd36d3a2cc965b4d184a3deb958471676b0f60ec3a3933f0554179001d041ed6
|
File details
Details for the file synthcity-0.2.12-py3-none-any.whl.
File metadata
- Download URL: synthcity-0.2.12-py3-none-any.whl
- Upload date:
- Size: 434.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.9.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7421a1f98d76f302967f32274b552323e9d63db0419fb652ac978fe96e860b3b
|
|
| MD5 |
b1f05dee11887cd05b6ee93591d60b1c
|
|
| BLAKE2b-256 |
aa290266c383bd38d1caffa857dba43ebe23332edc66cf766f45a7b5cc9ffac5
|