THUNDER: Tile-level Histopathology image UNDERstanding benchmark
Project description
Tile-level Histopathology image Understanding benchmark

Published at NeurIPS 2025 Datasets and Benchmarks Track (Spotlight)
We introduce THUNDER, a comprehensive benchmark designed to rigorously compare foundation models across various downstream tasks in computational pathology. THUNDER enables the evaluation and analysis of feature representations, robustness, and uncertainty quantification of these models across different datasets. Our benchmark encompasses a diverse collection of well-established datasets, covering multiple cancer types, image magnifications, and varying image and sample sizes. We propose an extensive set of tasks aimed at thoroughly assessing the capabilities and limitations of foundation models in digital pathology.
⚡ Paper: THUNDER: Tile-level Histopathology image UNDERstanding benchmark
⚡ Homepage/Documentation: THUNDER docs
⚡ Leaderboards: THUNDER leaderboards
News
- 2025-11-14: GIGAPATH, and KAIKO-ViT model variants (S/8, S/16, B/8, B/16) are now supported in THUNDER. The downstream performance of GIGAPATH, KAIKO-S/16 and KAIKO-B/16 were integrated into the up-to-date rank-sum leaderboard and the SPIDER Leaderboard.
- 2025-11-11: You can now run any task in thunder on a custom dataset. Check the tutorial to learn more about this feature.
- 2025-10-30: DINOv3 model variants (B, S, L) are now supported in THUNDER. Their downstream performance was integrated along with SPIDER results into a new up-to-date rank-sum leaderboard.
- 2025-10-06: As requested (https://github.com/MICS-Lab/thunder/issues/1), a new zero-shot classification task to evaluate VLMs was included into THUNDER. Example command:
thunder benchmark keep spider_breast zero_shot_vlm. See the dedicated zero-shot classification leaderboard. - 2025-09-30: Patch-level SPIDER datasets have been integrated into THUNDER. See the dedicated SPIDER leaderboard.
- 2025-09-18: THUNDER was accepted to NeurIPS 2025 Datasets & Benchmarks Track as a Spotlight presentation!
Overview
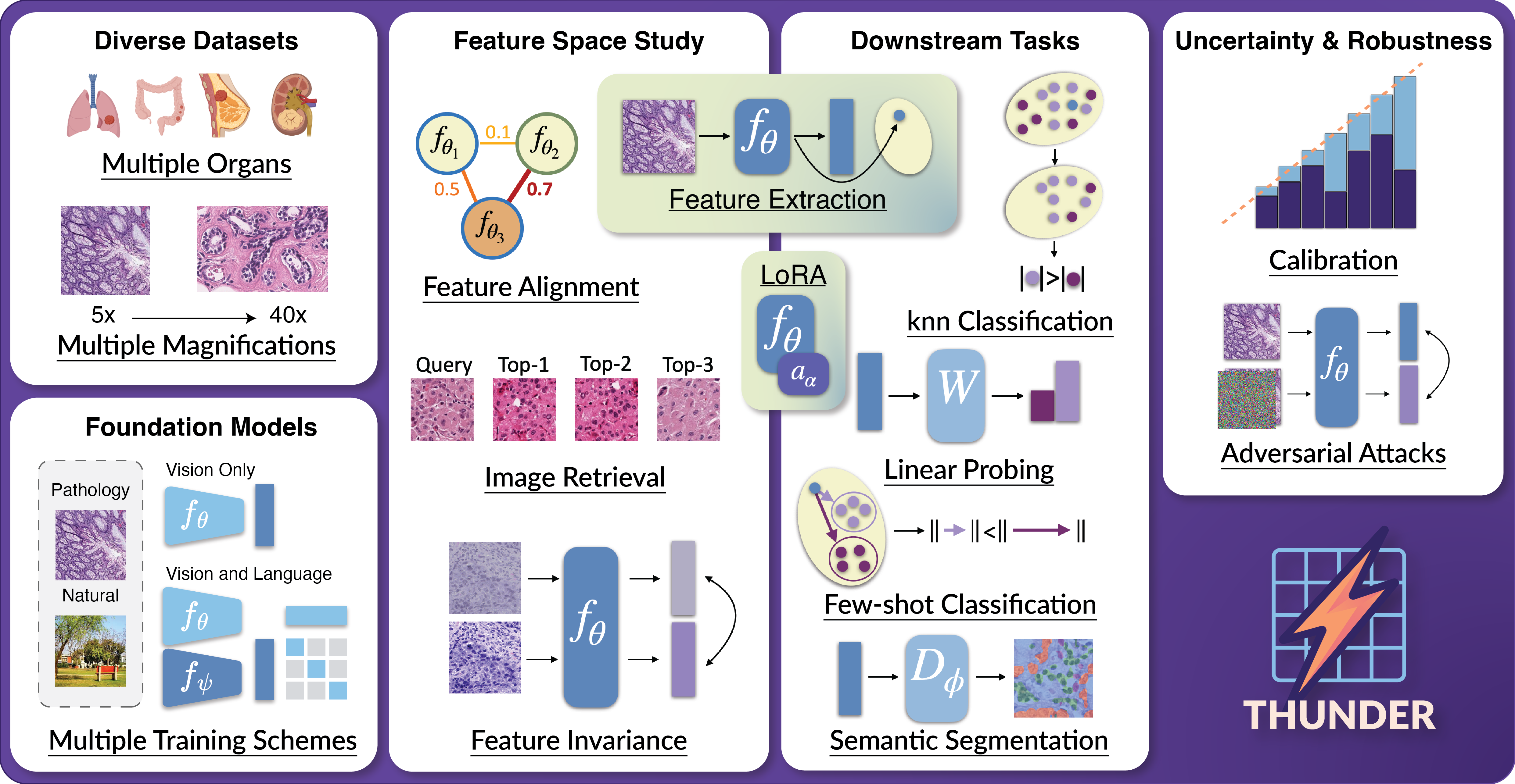
We propose a benchmark to compare and study foundation models across three axes: (i) downstream task performance, (ii) feature space comparisons, and (iii) uncertainty and robustness. Our current version integrates 23 foundation models, vision-only, vision-language, trained on pathology or natural images, on 16 datasets covering different magnifications and organs. THUNDER also supports the use of new user-defined models for direct comparisons.
Usage
To learn more about how to use thunder, please visit our documentation.
An API and command line interface (CLI) are provided to allow users to download datasets, models, and run benchmarks. The API is designed to be user-friendly and allows for easy integration into existing workflows. The CLI provides a convenient way to access the same functionality from the command line.
[!IMPORTANT] Downloading supported foundation models: you will have to visit the Huggingface URL of supported models you wish to use in order to accept usage conditions.
List of Huggingface URLs
- UNI: https://huggingface.co/MahmoodLab/UNI
- UNI2-h: https://huggingface.co/MahmoodLab/UNI2-h
- Virchow: https://huggingface.co/paige-ai/Virchow
- Virchow2: https://huggingface.co/paige-ai/Virchow2
- H-optimus-0: https://huggingface.co/bioptimus/H-optimus-0
- H-optimus-1: https://huggingface.co/bioptimus/H-optimus-1
- CONCH: https://huggingface.co/MahmoodLab/CONCH
- TITAN/CONCHv1.5: https://huggingface.co/MahmoodLab/TITAN
- Phikon: https://huggingface.co/owkin/phikon
- Phikon2: https://huggingface.co/owkin/phikon-v2
- Hibou-b: https://huggingface.co/histai/hibou-b
- Hibou-L: https://huggingface.co/histai/hibou-L
- Midnight-12k: https://huggingface.co/kaiko-ai/midnight
- KEEP: https://huggingface.co/Astaxanthin/KEEP
- QuiltNet-B-32: https://huggingface.co/wisdomik/QuiltNet-B-32
- PLIP: https://huggingface.co/vinid/plip
- MUSK: https://huggingface.co/xiangjx/musk
- DINOv2-B: https://huggingface.co/facebook/dinov2-base
- DINOv2-L: https://huggingface.co/facebook/dinov2-large
- ViT-B: https://huggingface.co/google/vit-base-patch16-224-in21k
- ViT-L: https://huggingface.co/google/vit-large-patch16-224-in21k
- CLIP-B: https://huggingface.co/openai/clip-vit-base-patch32
- CLIP-L: https://huggingface.co/openai/clip-vit-large-patch14
API Usage
When using the API you can run the following code to download datasets, models and run a benchmark:
from thunder import benchmark
benchmark("phikon", "break_his", "knn")
CLI Usage
When using the CLI you can run the following command to see all available options,
thunder --help
In order to reproduce the above example you can run the following command:
thunder benchmark phikon break_his knn
Extracting embeddings with any supported foundation model (API Usage)
We also provide a get_model_from_name function through our API to extract embeddings using any foundation model we support on your own data. Below is an example if you want to get the Pytorch callable, transforms and function to extract embeddings for uni2h:
from thunder.models import get_model_from_name
model, transform, get_embeddings = get_model_from_name("uni2h", device="cuda")
Installing thunder
Code tested with Python 3.10. To replicate, you can create the following conda environment and activate it,
conda create -n thunder_env python=3.10
conda activate thunder_env
To install thunder run one of the following commands:
From PyPi
pip install thunder-bench
From Source
pip install -e . # install the package in editable mode
pip install . # install the package
Before running thunder, ensure that the environment variable THUNDER_BASE_DATA_FOLDER is defined. This variable specifies the path where outputs, foundation models, and datasets will be stored. You can set it by running:
export THUNDER_BASE_DATA_FOLDER="/path/to/your/data/folder"
Replace /path/to/your/data/folder with your desired storage directory.
If you want to use the CONCH and MUSK models, you should install them as follows:
pip install git+https://github.com/Mahmoodlab/CONCH.git # CONCH
pip install git+https://github.com/lilab-stanford/MUSK.git # MUSK
Citation
@article{marza2025thunder,
title={{THUNDER}: Tile-level Histopathology image UNDERstanding benchmark},
author={Marza, Pierre and Fillioux, Leo and Boutaj, Sofi{\`e}ne and Mahatha, Kunal and Desrosiers, Christian and Piantanida, Pablo and Dolz, Jose and Christodoulidis, Stergios and Vakalopoulou, Maria},
journal={Neural Information Processing Systems (NeurIPS) D&B Track},
year={2025}
}
Acknowledgments
This work has been partially supported by ANR-23-IAHU-0002, ANR-21-CE45-0007, ANR-23-CE45-0029, and the Health Data Hub (HDH) as part of the second edition of the France-Québec call for projects Intelligence Artificielle en santé. It was performed using computational resources from the Mésocentre computing center of Université Paris-Saclay, CentraleSupélec and École Normale Supérieure Paris-Saclay supported by CNRS and Région Île-de-France, and from GENCI-IDRIS (Grant 2025-AD011016068).
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file thunder_bench-0.2.0.tar.gz.
File metadata
- Download URL: thunder_bench-0.2.0.tar.gz
- Upload date:
- Size: 94.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.17
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dec10f135d28a6c3e757ba72b0e7821d7481b52097ad87e73df012f50d6a38f6
|
|
| MD5 |
d281c9f42cee8659818694d454432320
|
|
| BLAKE2b-256 |
f63fe1fac50a00a407a8e3d8a697c0117b1abfde3b06c0a4fc53b1eccf46052f
|
File details
Details for the file thunder_bench-0.2.0-py3-none-any.whl.
File metadata
- Download URL: thunder_bench-0.2.0-py3-none-any.whl
- Upload date:
- Size: 140.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.17
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f0b3276930bc8d2d859be51aa4cdfc243f40d3dfc139b721ecb1146cf353054b
|
|
| MD5 |
6a8b95a49a8757eba4460fa533038464
|
|
| BLAKE2b-256 |
39541787a2f6c2eaae4e1f11209e42e08b8f4460f227f9294ae0eb1861cc9e05
|

















