TIPP3 - A Phylogeney-Based Abundance Profiling Tool
Project description
- Developer:
Chengze Shen
News
1.21.2025 -
1.20.2025 - TIPP3 now is feature complete for abundance profiling, for both the more accurate TIPP3 mode or the fast TIPP3-fast mode. By default, TIPP3-fast is used.
12.17.2024 - TIPP3 now can handle both fasta (.fasta or .fa) and fastq (.fastq or .fq) files as inputs, and they can be in gzip format (e.g., queries.fasta.gz or queries.fq.gz).
TODO list
1.20.2025 - Add a script for downloading the latest reference package to a user specified directory.
Method Overview
TIPP3 is a metagenomic profiling method that solves the following problems:
- Taxonomic identification
Input: A query read q
Output: The taxonomic lineage of q (if identified)
- Abundance profiling
Input: A set Q of query reads
Output: An abundance profile estimated on Q
TIPP3 continues the TIPP-family methods (prior methods: TIPP and TIPP2), which use a marker gene database to identify the taxonomic lineage of input reads (if the read comes from a marker gene). See the pipeline below for the TIPP3 workflow.
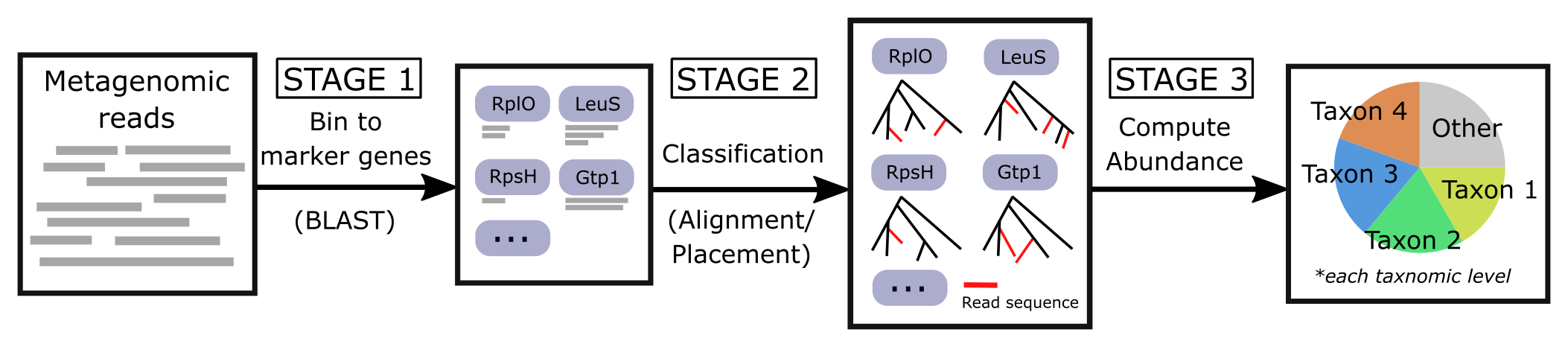
Publication(s) |
|---|
(TIPP3) Shen, Chengze, Eleanor Wedell, Mihai Pop, and Tandy Warnow, “TIPP3 and TIPP3-fast: improved abundance profiling in metagenomics.” TBD. |
(TIPP2) Nguyen, Nam, Siavash Mirarab, Bo Liu, Mihai Pop, and Tandy Warnow, “TIPP: Taxonomic identification and phylogenetic profiling.” Bioinformatics, 2014. https://doi.org/10.1093/bioinformatics/btu721 |
(TIPP) Shah, Nidhi, Erin K. Molloy, Mihai Pop, and Tandy Warnow, “TIPP2: metagenomic taxonomic profiling using phylogenetic markers.” Bioinformatics, 2020. https://doi.org/10.1093/bioinformatics/btab023 |
Note and Acknowledgment
TIPP3 includes and uses:
pplacer (v1.1.alpha19).
Batch-SCAMPP (v1.0.0).
Installation
TIPP3 was tested on Python 3.8 to 3.12.
There are two ways to install and use TIPP3: with PyPI (pip install) or directly with this GitHub repository. If you have any difficulties installing or running TIPP3, please contact Chengze Shen (chengze5@illinois.edu).
External Requirements
BLAST is a hard requirement to run TIPP3. The software will automatically look for blastn in the $PATH environment variable. If you have not installed BLAST, you can find the latest version from https://ftp.ncbi.nlm.nih.gov/blast/executables/blast+/LATEST/.
Install with PyPI (pip)
TODO
Install from source files
Requirements
python>=3.7 configparser>=5.0.0 DendroPy>=4.5.2 numpy>=1.21.6 psutil>=5.0.0 setuptools>=60.0.0 treeswift>=1.1.28 witch-msa>=1.0.7 lz4>=4.3.2
Installation Steps
# 1. Install via GitHub repo
git clone https://github.com/c5shen/TIPP3.git
# 2. Install all requirements
pip3 install -r requirements.txt
# 3. Execute tipp3.py executable for the first time with "-h" to see
# allowed commandline parameters and example usages
# Running TIPP3 for the first time will also create the main config
# file at "~/.tipp3/main.config", which stores the default behavior
# for running TIPP3 (including all binary executable paths)
python3 tipp3.py [-h]main.config
main.config file will be created the first time running TIPP3 at the user root directory (~/.tipp3/main.config). This file stores the default behavior for running TIPP3 and the paths to all binary executables that TIPP3 need to use.
user-specified config file
In addition, a user can specify a customized config file with -c or --config-file parameter option when running TIPP3 (e.g., tipp3.py -c user.config). The user.config file will override settings from main.config (if overlaps). Command-line arguments still have the highest priority and will override both config files, if any parameters overlap.
Usage
The general command to run TIPP3 is listed below. By default, TIPP3-fast is run, which is significantly faster than the more accurate TIPP3 mode. See Examples below for how to customize the TIPP3 pipeline.
# (Optional) change the logging level to DEBUG for more verbose logging
export TIPP_LOGGING_LEVEL=debug
# TIPP3 supports the following formats for "-i [query reads]"
# XXX.fasta[.gz, .gzip]
# XXX.fa[.gz, .gzip]
# XXX.fastq[.gz, .gzip]
# XXX.fq[.gz, .gzip]
python3 tipp3.py -r [reference package path] -i [query reads] -d [output directory]Examples
Some examples of TIPP3 usage can be found at the bottom of the help text running:
python3 tipp3.py -hAll of the following examples can be found in the examples/run.sh bash script, with example data stored under examples/data. The default example data used is a small set of Illumina short reads denoted as illumina.small.queries.fasta.
Scenario 1
(TIPP3-fast) Use BLAST for query alignment, and Batch-SCAMPP (bscampp) for query placement. Keep all temporary files during the run.
python3 tipp3.py -i examples/illumina.small.queries.fasta \
--reference-package [reference package dir] --outdir tipp3_scenario1 \
--alignment-method blast --placement-method bscampp \
-t 16 --keeptempScenario 2
Use BLAST for query alignment, and pplacer with the taxtastic package for query placement (pplacer-taxtastic). Keep all temporary files.
python3 tipp3.py -i examples/illumina.small.queries.fasta \
--reference-package [reference package dir] --outdir tipp3_scenario1 \
--alignment-method blast --placement-method pplacer-taxtastic \
-t 16 --keeptempScenario 3
(TIPP3) Use WITCH for query alignment, and pplacer-taxtastic for query placement. Keep all temporary files.
python3 tipp3.py -i examples/illumina.small.queries.fasta \
--reference-package [reference package dir] --outdir tipp3_scenario1 \
--alignment-method witch --placement-method pplacer-taxtastic \
-t 16 --keeptempProject details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file tipp3-0.1a0.tar.gz.
File metadata
- Download URL: tipp3-0.1a0.tar.gz
- Upload date:
- Size: 8.3 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.0.1 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e8da8ab04571961f2bf958906fe812c99af25cfe7c0cbe2f6a3710faa2e75fbb
|
|
| MD5 |
8d595e2e911f0811efbb445ec5981834
|
|
| BLAKE2b-256 |
049c8f08d20dbb3b35186d149f993b7454041cd358b0f5b36f29fcbf8bcb297d
|
Provenance
The following attestation bundles were made for tipp3-0.1a0.tar.gz:
Publisher:
python-publish.yml on c5shen/TIPP3
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
tipp3-0.1a0.tar.gz -
Subject digest:
e8da8ab04571961f2bf958906fe812c99af25cfe7c0cbe2f6a3710faa2e75fbb - Sigstore transparency entry: 164289156
- Sigstore integration time:
-
Permalink:
c5shen/TIPP3@7fc171487de3bcc2dcbf7551f6ee07d6247e9a4c -
Branch / Tag:
refs/tags/v0.1a - Owner: https://github.com/c5shen
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@7fc171487de3bcc2dcbf7551f6ee07d6247e9a4c -
Trigger Event:
release
-
Statement type:
File details
Details for the file tipp3-0.1a0-py3-none-any.whl.
File metadata
- Download URL: tipp3-0.1a0-py3-none-any.whl
- Upload date:
- Size: 8.4 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.0.1 CPython/3.12.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d81909777f57b36b7023e94aeffef0114ce5f04af344a2685f4d7ad272e2eee5
|
|
| MD5 |
64016e5cbc54b80355a6c71578220f18
|
|
| BLAKE2b-256 |
206a8e0710c3c2b5124961409bd58a0c5d4519cd1924942d07feb96a99b7b349
|
Provenance
The following attestation bundles were made for tipp3-0.1a0-py3-none-any.whl:
Publisher:
python-publish.yml on c5shen/TIPP3
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
tipp3-0.1a0-py3-none-any.whl -
Subject digest:
d81909777f57b36b7023e94aeffef0114ce5f04af344a2685f4d7ad272e2eee5 - Sigstore transparency entry: 164289157
- Sigstore integration time:
-
Permalink:
c5shen/TIPP3@7fc171487de3bcc2dcbf7551f6ee07d6247e9a4c -
Branch / Tag:
refs/tags/v0.1a - Owner: https://github.com/c5shen
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@7fc171487de3bcc2dcbf7551f6ee07d6247e9a4c -
Trigger Event:
release
-
Statement type:












