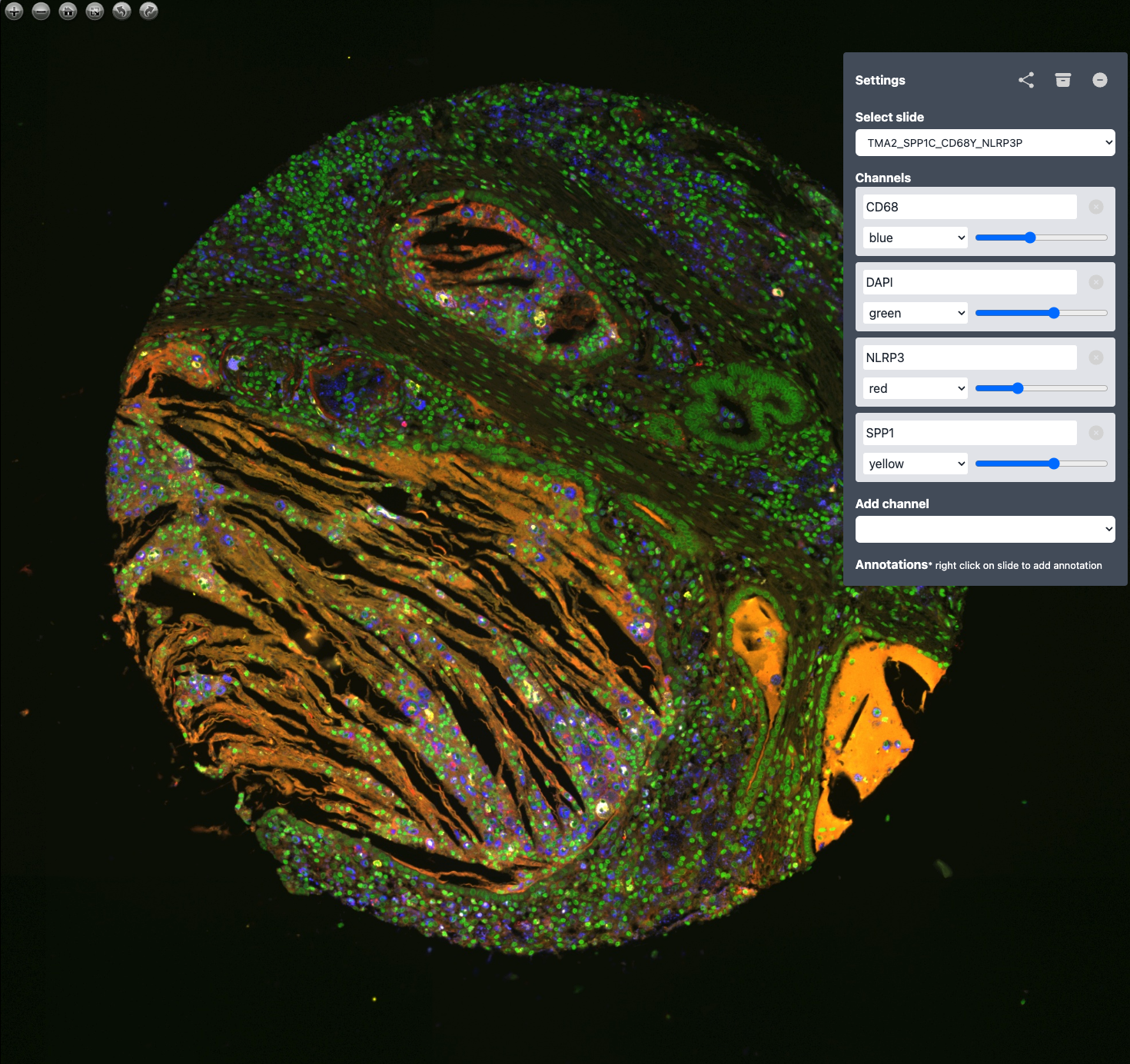
Standalone multi-channel OME-TIFF viewer with a built-in web UI.
Project description
TissueViewer
Standalone multi-channel OME-TIFF viewer with a built-in web UI.
Point it at a file or a folder and open your browser.
Contents
- What it is
- Supported formats
- Installation
- Quick start
- Command-line reference
- Environment variables
- Configuration file
- Filename conventions
- Programmatic use
- Troubleshooting
- Development
- Docker migration
- License
- Acknowledgements
What it is
TissueViewer is a zero-config viewer for high-resolution multi-channel microscopy images. It serves deep-zoom tiles over HTTP and ships with a pre-built OpenSeadragon front-end that lets you colourise channels, adjust gains and windowing, and save per-slide settings.
Supported formats
- OME-TIFF (
.ome.tif,.ome.tiff) — any bit depth, any number of channels, with or without a built-in pyramid.
Support for additional formats (plain TIFF, OME-Zarr) is planned as
pluggable handlers; see src/tissueviewer/formats/ for the registry.
Installation
pip install tissueviewer
Or from a source checkout:
git clone https://github.com/davidvi/tissueviewer.git
cd tissueviewer
pip install .
Requires Python 3.10 or later. On macOS / Linux / Windows.
Quick start
Open a single file:
tissueviewer /path/to/slide.ome.tiff
Browse a folder of slides (root folder becomes the public location):
tissueviewer /path/to/slides
Browse a nested tree of folders:
tissueviewer /path/to/slides --recursive --open
Use a config file:
tissueviewer --config config.yaml
Check every file in a directory can be opened, then exit:
tissueviewer /path/to/slides --validate
Command-line reference
usage: tissueviewer [TARGET] [options]
positional:
TARGET Path to an .ome.tif(f) file or a directory of them.
options:
-r, --recursive Include nested subdirectories.
-c, --config FILE YAML configuration file.
-p, --port N Listen port (default 8000).
-H, --host HOST Bind address (default 127.0.0.1).
-o, --open Open the default web browser after start.
-w, --watch Rescan TARGET on filesystem changes.
--validate Verify every OME-TIFF, then exit.
--no-save Disable writing .sample.json sidecars.
--log-level LEVEL DEBUG | INFO | WARNING | ERROR (default INFO).
-V, --version Show version and exit.
-h, --help Show help and exit.
TARGET is optional if either --config is given or TV_SLIDE_DIR is set.
Environment variables
Kept for backward compatibility with the Docker deployment. CLI and config values take precedence.
| Variable | Effect |
|---|---|
TV_SLIDE_DIR |
Default data directory (equivalent to positional TARGET). |
TV_HOST |
Default bind address. |
TV_PORT |
Default listen port. |
TV_SAVE |
false/0/no disables sample.json writes. |
TV_LOG_LEVEL |
Default log level. |
Configuration file
# config.yaml
data_dir: /srv/tissue-viewer/slides
recursive: true
host: 0.0.0.0
port: 8000
save_enabled: false
log_level: INFO
colors:
- red
- green
- blue
tissueviewer --config config.yaml
Precedence (highest wins): CLI flags → environment variables → YAML → built-in defaults.
Filename conventions
All recognized slide files end in .ome.tif or .ome.tiff (case-insensitive).
For a slide named patient42.ome.tiff, two sidecar files may appear next
to it:
| File | Who writes it | Purpose |
|---|---|---|
patient42.sample.json |
the web UI | Per-slide channel colours / gains / annotations. |
patient42.metadata.json |
TissueViewer itself | Cache of OME-XML metadata; regenerated on mtime. |
Delete them freely to reset.
Locations
The web UI groups slides into locations. The mapping:
- Single-file mode → the file lives under the
publiclocation. - Directory mode (no
--recursive) → files at the root →public; every immediate subdirectory → its own location named after the subdirectory. - Directory mode with
--recursive→ files at the root still go topublic; files in nested subdirectories are grouped under a slash-joined location path like2024/batch_A.
Programmatic use
create_app builds a regular FastAPI instance, suitable for any ASGI
server (uvicorn, hypercorn, daphne).
from tissueviewer import Config, create_app
app = create_app(Config(data_dir="/srv/slides", recursive=True))
uvicorn my_module:app --host 0.0.0.0 --port 8000
Troubleshooting
ModuleNotFoundError: imagecodecs— installimagecodecsor reinstalltissueviewer; many OME-TIFFs use LZW/JPEG/Zstd compression.- Port already in use —
tissueviewer --port 8001. - Blank tiles / wrong colours — delete the slide's
.sample.jsonto reset settings, then reload. - Apple Silicon wheels —
imagecodecsandopencv-python-headlessship arm64 wheels; no local build required. - Zarr 2 vs 3 — both are supported. The pin is
zarr>=2.15,<4. - OpenCV GUI warning —
tissueviewerdepends onopencv-python-headless; a warning aboutcv2.imshownot being available is normal and harmless.
Development
See CONTRIBUTING.md for local setup, how to rebuild the Vue UI, and how to cut a release.
Docker migration
TissueViewer used to ship as a Docker image with a Java-based
bioformats2raw conversion pipeline. That mode is gone in 0.2; see
MIGRATION.md for guidance.
License
Creative Commons Attribution-NonCommercial 4.0 International — see LICENSE.md.
Acknowledgements
TissueViewer uses OpenSeadragon for high-performance tiled image visualisation.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file tissueviewer-0.2.0.tar.gz.
File metadata
- Download URL: tissueviewer-0.2.0.tar.gz
- Upload date:
- Size: 721.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9bb7ddefffc66bac1a9e85ddb29c5eb884bd68ccd15a75f0909d6084e5fa2ddf
|
|
| MD5 |
0f5f4fe833036c1ec8213d03fd8652f1
|
|
| BLAKE2b-256 |
1b1d17507664f2fdcc1c931c55c0d0b4d153feb9cd275e02bb029fb79fbdfca4
|
Provenance
The following attestation bundles were made for tissueviewer-0.2.0.tar.gz:
Publisher:
push.yml on davidvi/tissueviewer
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
tissueviewer-0.2.0.tar.gz -
Subject digest:
9bb7ddefffc66bac1a9e85ddb29c5eb884bd68ccd15a75f0909d6084e5fa2ddf - Sigstore transparency entry: 1340722250
- Sigstore integration time:
-
Permalink:
davidvi/tissueviewer@f687b8222ed41b046a6ba49df989bfe7dc43c728 -
Branch / Tag:
refs/heads/main - Owner: https://github.com/davidvi
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
push.yml@f687b8222ed41b046a6ba49df989bfe7dc43c728 -
Trigger Event:
push
-
Statement type:
File details
Details for the file tissueviewer-0.2.0-py3-none-any.whl.
File metadata
- Download URL: tissueviewer-0.2.0-py3-none-any.whl
- Upload date:
- Size: 729.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ef3cae6ffe90098b9b935d4561c6c2d6e2e6c7013761696dcd3505c1958d9513
|
|
| MD5 |
35cbc1111fabbb7aebc0453b80dc194d
|
|
| BLAKE2b-256 |
31a00f7f633db3d4deb827654ef41f0ef1340c8d99ad7660468672a948dba898
|
Provenance
The following attestation bundles were made for tissueviewer-0.2.0-py3-none-any.whl:
Publisher:
push.yml on davidvi/tissueviewer
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
tissueviewer-0.2.0-py3-none-any.whl -
Subject digest:
ef3cae6ffe90098b9b935d4561c6c2d6e2e6c7013761696dcd3505c1958d9513 - Sigstore transparency entry: 1340722252
- Sigstore integration time:
-
Permalink:
davidvi/tissueviewer@f687b8222ed41b046a6ba49df989bfe7dc43c728 -
Branch / Tag:
refs/heads/main - Owner: https://github.com/davidvi
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
push.yml@f687b8222ed41b046a6ba49df989bfe7dc43c728 -
Trigger Event:
push
-
Statement type: