Tools for medical image processing with PyTorch
Project description
Tools like TorchIO are a symptom of the maturation of medical AI research using deep learning techniques.
Jack Clark, Policy Director at OpenAI (link).
| Package |



|
| CI |



|
| Code |


|
| Tutorials |

|
| Community |


|
(Queue for patch-based training)
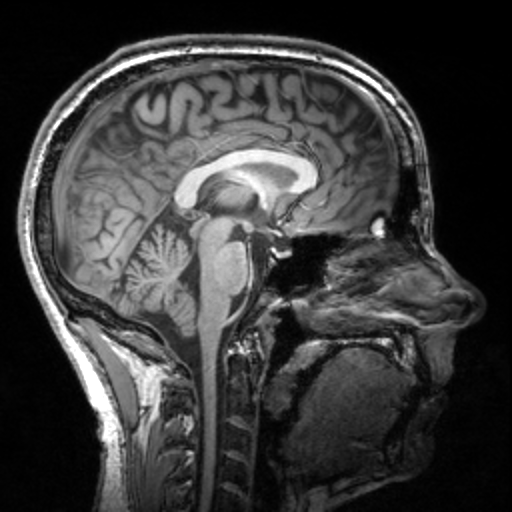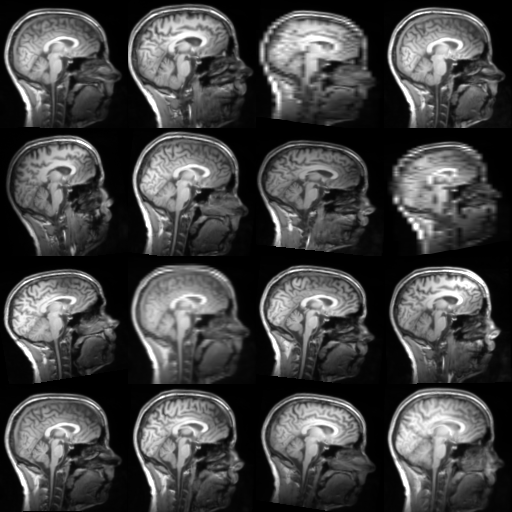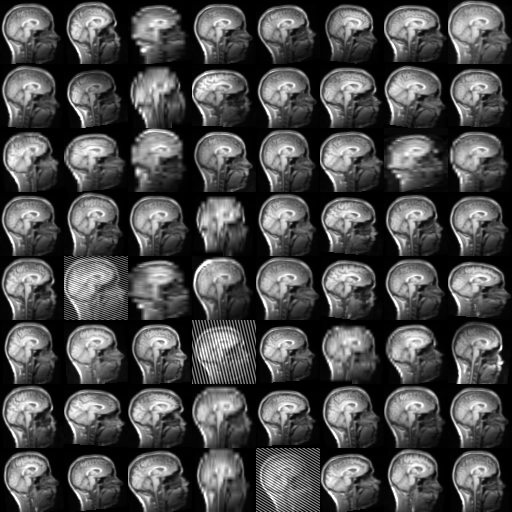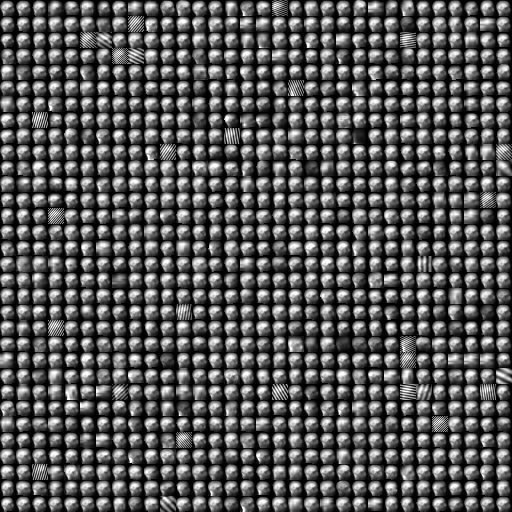
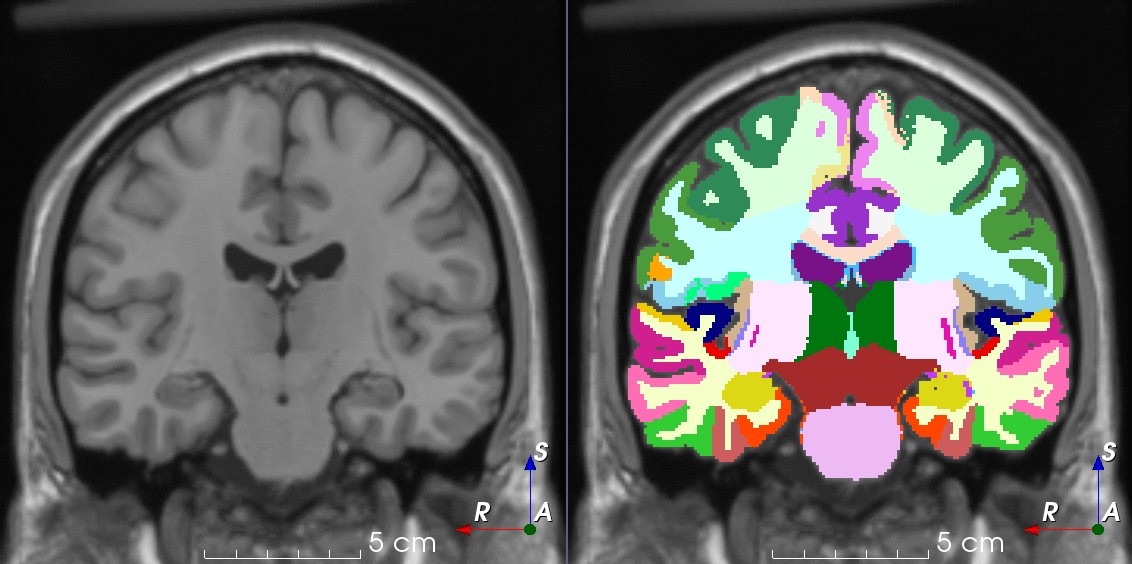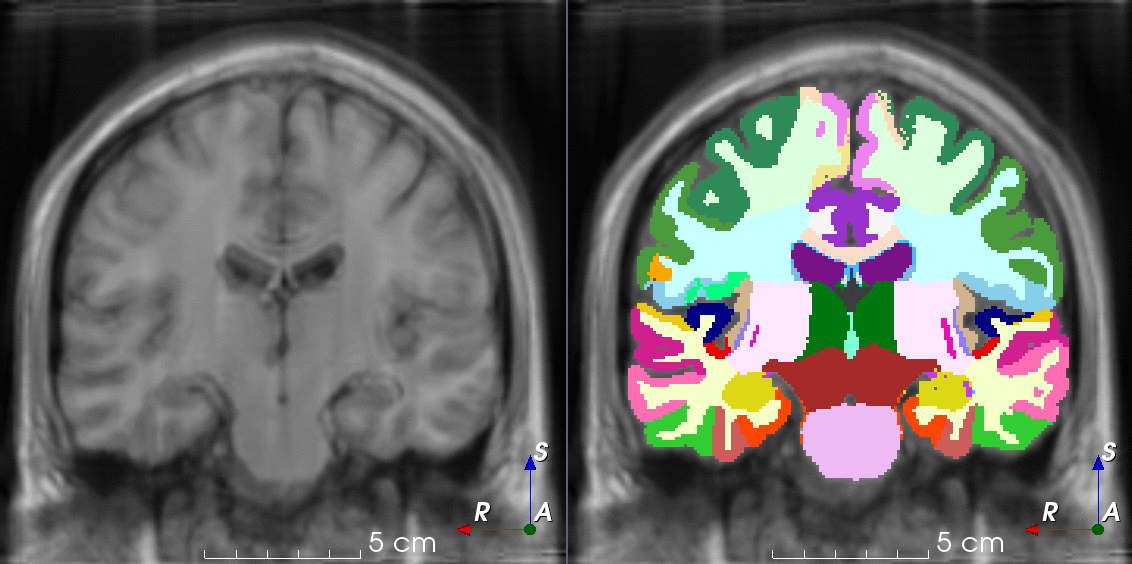
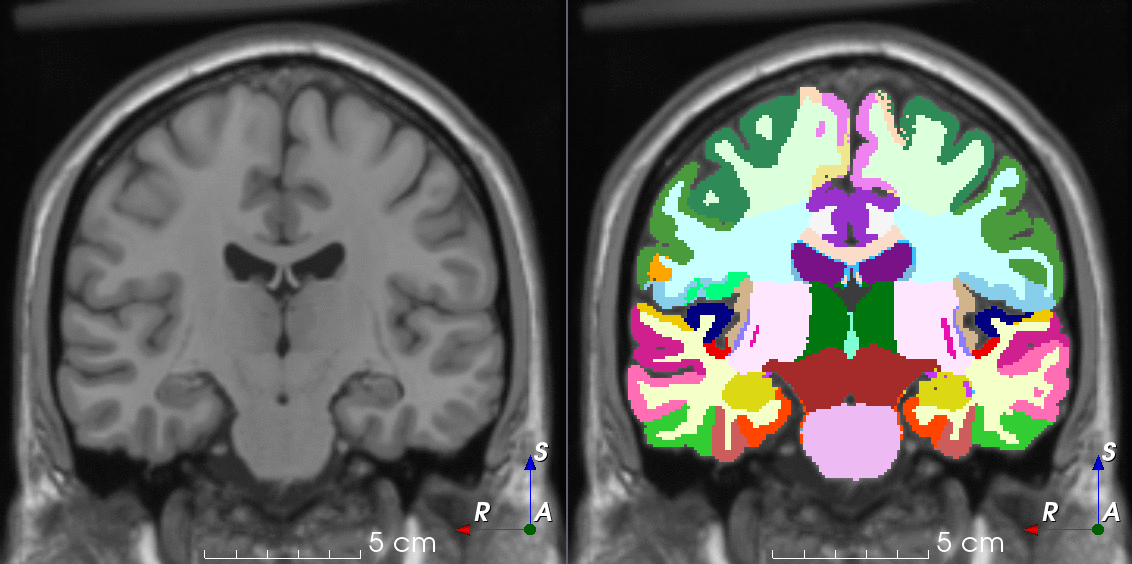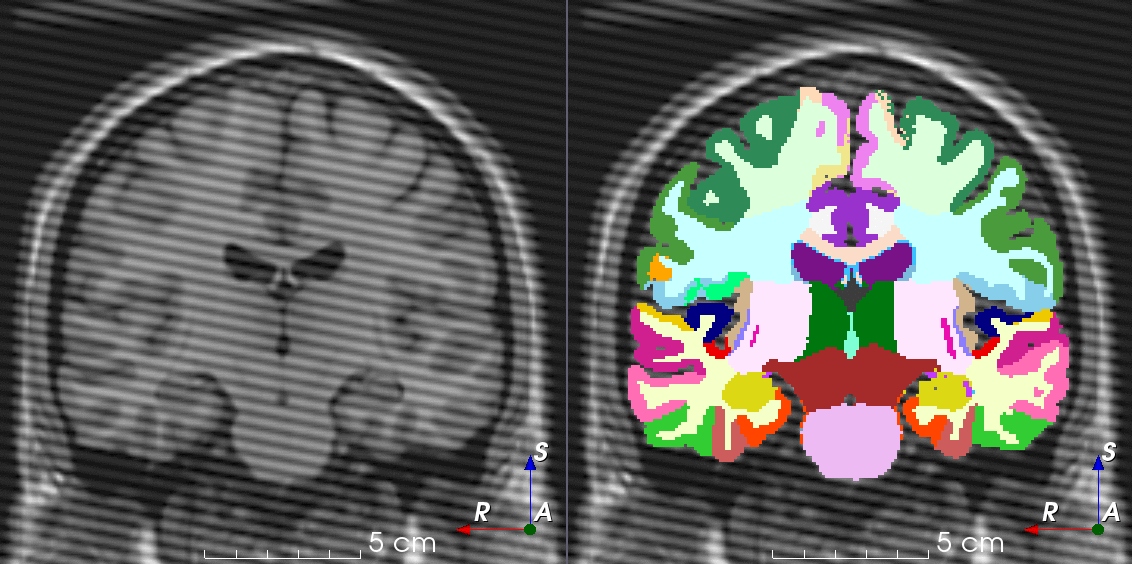
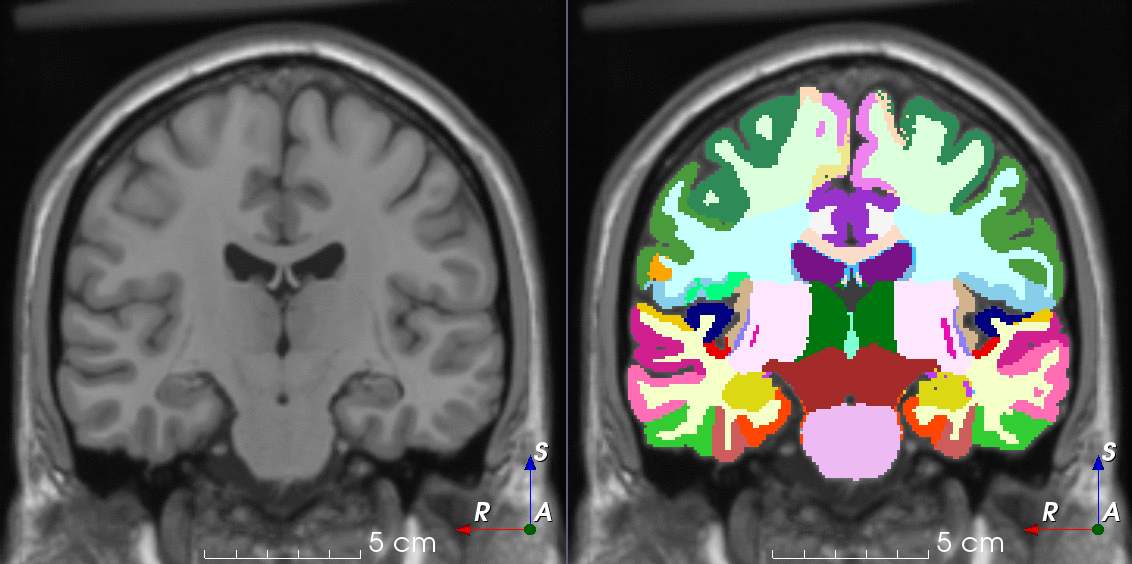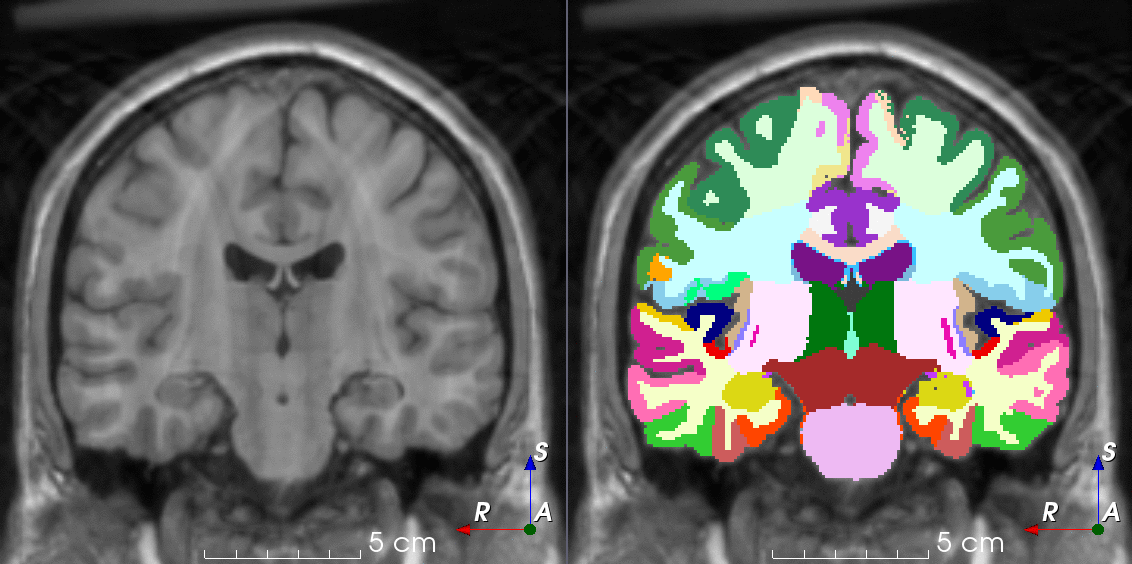
TorchIO is a Python package containing a set of tools to efficiently read, preprocess, sample, augment, and write 3D medical images in deep learning applications written in PyTorch, including intensity and spatial transforms for data augmentation and preprocessing. Transforms include typical computer vision operations such as random affine transformations and also domain-specific ones such as simulation of intensity artifacts due to MRI magnetic field inhomogeneity or k-space motion artifacts.
This package has been greatly inspired by NiftyNet, which is not actively maintained anymore.
Credits
If you like this repository, please click on Star!
If you use this package for your research, please cite our paper:
BibTeX entry:
@article{perez-garcia_torchio_2021,
title = {{TorchIO}: a {Python} library for efficient loading, preprocessing, augmentation and patch-based sampling of medical images in deep learning},
journal = {Computer Methods and Programs in Biomedicine},
pages = {106236},
year = {2021},
issn = {0169-2607},
doi = {https://doi.org/10.1016/j.cmpb.2021.106236},
url = {https://www.sciencedirect.com/science/article/pii/S0169260721003102},
author = {P{\'e}rez-Garc{\'i}a, Fernando and Sparks, Rachel and Ourselin, S{\'e}bastien},
}
This project is supported by the following institutions:
- Engineering and Physical Sciences Research Council (EPSRC) & UK Research and Innovation (UKRI)
- EPSRC Centre for Doctoral Training in Intelligent, Integrated Imaging In Healthcare (i4health) (University College London)
- Wellcome / EPSRC Centre for Interventional and Surgical Sciences (WEISS) (University College London)
- School of Biomedical Engineering & Imaging Sciences (BMEIS) (King's College London)
Getting started
See Getting started for installation instructions and a Hello, World! example.
Longer usage examples can be found in the tutorials.
Read the documentation for more information.
Please create an issue if you think something is missing.
Contributors
Thanks goes to all these people (emoji key):
This project follows the all-contributors specification. Contributions of any kind welcome!
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file torchio-0.21.3.tar.gz.
File metadata
- Download URL: torchio-0.21.3.tar.gz
- Upload date:
- Size: 145.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: uv/0.10.0 {"installer":{"name":"uv","version":"0.10.0","subcommand":["publish"]},"python":null,"implementation":{"name":null,"version":null},"distro":{"name":"Ubuntu","version":"24.04","id":"noble","libc":null},"system":{"name":null,"release":null},"cpu":null,"openssl_version":null,"setuptools_version":null,"rustc_version":null,"ci":true}
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
fb19d70c34e6378b5e44ef613832ac818bf145cdb65e82ad00fa8bc97219839b
|
|
| MD5 |
022fccfc15379b10b2ac626865e312dc
|
|
| BLAKE2b-256 |
dfec01eead3fc5ec42c852448a75c35ad4ee5c208a9bb9516cba7b7c118e8b04
|
Provenance
The following attestation bundles were made for torchio-0.21.3.tar.gz:
Publisher:
publish.yml on TorchIO-project/torchio
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
torchio-0.21.3.tar.gz -
Subject digest:
fb19d70c34e6378b5e44ef613832ac818bf145cdb65e82ad00fa8bc97219839b - Sigstore transparency entry: 936985444
- Sigstore integration time:
-
Permalink:
TorchIO-project/torchio@3752b4e84f6a30a4c26eb6df99705ac1c0e98afc -
Branch / Tag:
refs/tags/v0.21.3 - Owner: https://github.com/TorchIO-project
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@3752b4e84f6a30a4c26eb6df99705ac1c0e98afc -
Trigger Event:
push
-
Statement type:
File details
Details for the file torchio-0.21.3-py3-none-any.whl.
File metadata
- Download URL: torchio-0.21.3-py3-none-any.whl
- Upload date:
- Size: 194.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: uv/0.10.0 {"installer":{"name":"uv","version":"0.10.0","subcommand":["publish"]},"python":null,"implementation":{"name":null,"version":null},"distro":{"name":"Ubuntu","version":"24.04","id":"noble","libc":null},"system":{"name":null,"release":null},"cpu":null,"openssl_version":null,"setuptools_version":null,"rustc_version":null,"ci":true}
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1c08864de2dcea3e493ceb3424b3987b6f4b369d4e41ecfd2a969903412054c2
|
|
| MD5 |
6245d7c0cd077a63bff511f511c66eb8
|
|
| BLAKE2b-256 |
2f880c52bde45aa2c08634dd4a51f8dfb0ec8e6fb9a6af7a099d0ca672c32252
|
Provenance
The following attestation bundles were made for torchio-0.21.3-py3-none-any.whl:
Publisher:
publish.yml on TorchIO-project/torchio
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
torchio-0.21.3-py3-none-any.whl -
Subject digest:
1c08864de2dcea3e493ceb3424b3987b6f4b369d4e41ecfd2a969903412054c2 - Sigstore transparency entry: 936985450
- Sigstore integration time:
-
Permalink:
TorchIO-project/torchio@3752b4e84f6a30a4c26eb6df99705ac1c0e98afc -
Branch / Tag:
refs/tags/v0.21.3 - Owner: https://github.com/TorchIO-project
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@3752b4e84f6a30a4c26eb6df99705ac1c0e98afc -
Trigger Event:
push
-
Statement type: