A tool for rapid anatomical structure detection and analysis.
Project description
TotalSegmentator 2D: A Tool for Rapid Anatomical Structure Analysis
⚡ Latest Updates
- 2025-08-26: v1.1.0 released → includes models pretrained on the TotalSegmentator v2.0.1
- 2025-07-31: v1.0.0 released → includes [MIUA2025a] TS2D models pretrained on the TotalSegmentator v1.0 dataset
- 🚧 Coming soon: TSXR — X-Ray segmentation models, see [MIUA2025b]
What is TS2D?
TotalSegmentator 2D (TS2D) is a fast and lightweight tool for anatomical structure segmentation and analysis.
It adapts TotalSegmentator (3D) by projecting CT scans into 2D views, enabling:
- ⚡ Rapid inference (results in less than a second, compared to several minutes for 3D methods)
- 💻 Low GPU/CPU requirements
- 🧠 Accurate segmentation of 117 anatomical structures (see
ts2d-v2models) - 🩻 Segmentation of native 2D X-ray scans (see the upcoming
tsxrmodels)
Use cases include:
- Anatomical structure segmentation and analysis in CT images.
- Body-region segmentation and detection in CT scans.
- X-ray analysis (coming soon)
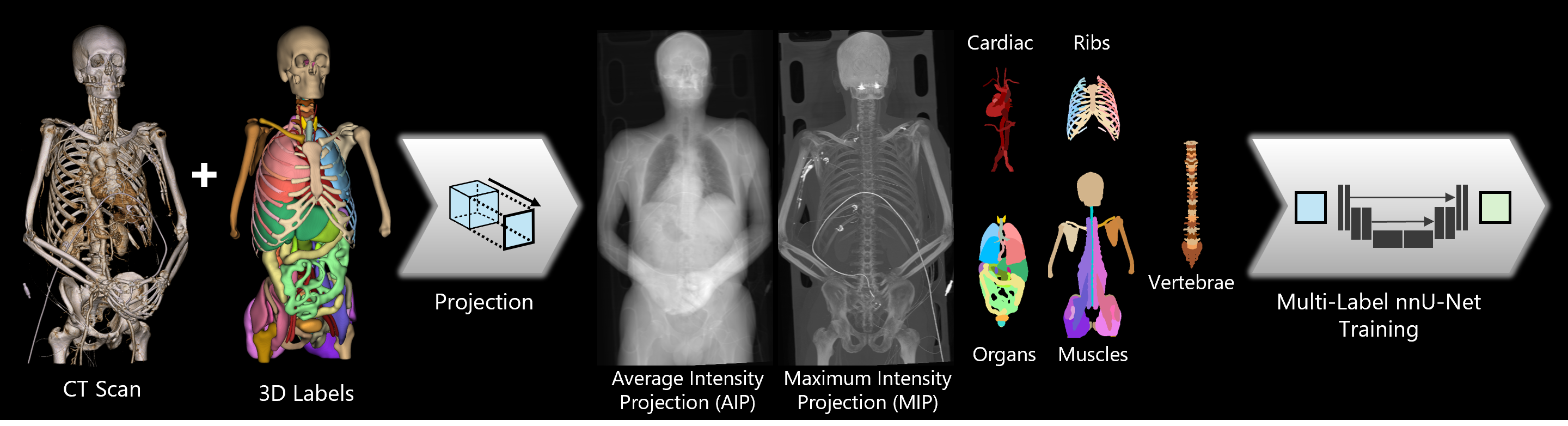
Figure 1: Standard CT workflow. Volumetric scans and ground-truth labels are projected onto the coronal plane to train five specialized 2D U-Net models. These models enable fast and efficient inference of 2D anatomical labels for any projected CT scan.
How It Works
TS2D uses coronal projection images generated through maximum and average intensity projection for the segmentation of anatomical structures. The two-channel input is processed by a 2D U-Net, implemented using our adapted nnU-Net framework to support multi-label output and thus correctly handle overlapping structures (see Figure 1). Pretrained models for both version 1 and version 2 of TotalSegmentator are available. Version 2 supports segmentation of 117 anatomical labels. The segmentation task was distributed across five specialized models, each focused on a distinct group of anatomical structures (see Figure 2).
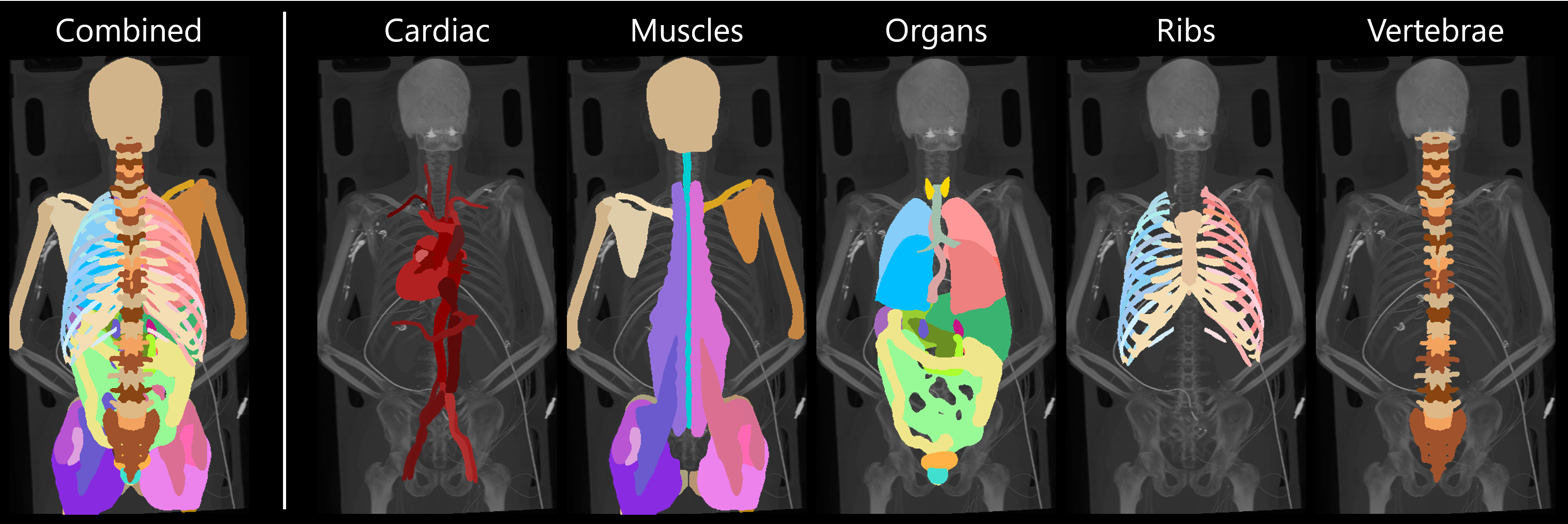
Figure 2: Segmentation results for the five anatomical group models used with the default TS2D configuration (ts2d-v2-ep4000b2), along with the combined output (Patient s0616).
TS2D was evaluated using projected ground-truth labels and its performance was compared to the original TotalSegmentator tool (TS3D), with both methods' inference results projected to 2D for consistency. A comprehensive comparison can be found in our publication [MIUA2025a].
| Method | Overall | Bone Structures | Soft-Tissue Structures | Inference Time (Nvidia RTX 4090) |
|---|---|---|---|---|
| TS2D (Ours) | 0.86 | 0.90 | 0.81 | 0.5-0.9 secs |
| TS3D | 0.97 | 0.97 | 0.97 | 43–146 secs |
Note: The table shows results for the TotalSegmentator v1 dataset to ensure comparability with the original TS3D publication.
Usage
Setup
TS2D has been tested with Python 3.12 and Pytorch 2.7.1 (CUDA 11.8) on a Windows 11 system and on Ubuntu (tested with CPU only).
👉 Install PyTorch before installing TS2D (see PyTorch setup).
Then install TS2D via:
- from PyPI:
pip install ts2d - from a local clone:
pip install . - from GitHub:
pip install git+https://github.com/risc-mi/totalsegmentator2D.git.
Get Started
You can run TS2D using the Command line interface (CLI):
ts2d -i <input_image> -o <output_directory>
or alternatively, you can use the API to run TS2D in your Python scripts:
from ts2d import TS2D
with TS2D() as model:
result = model.predict('<input_image>')
result.save(dest='<output_directory>')
TS2D will project the input image, run the segmentation models and save a multilabel segmentation file to the output directory. The segmentation labels can be parsed from the metadata, to view the segmentation use e.g. 3D Slicer to view the results. For more information, refer to the CLI help or the API documentation.
Default Models
By default, TS2D uses the model key ts2d-v2, which refers to models trained on the TotalSegmentator v2.0.1 dataset.
ts2d-v2→ pretrained on v2.0.1 dataset (117 labels)ts2d-v1→ pretrained on v1.0 dataset (104 labels)
➡️ See Available Models for a full list and instructions on selecting specific sub-models.
Performance Evaluation
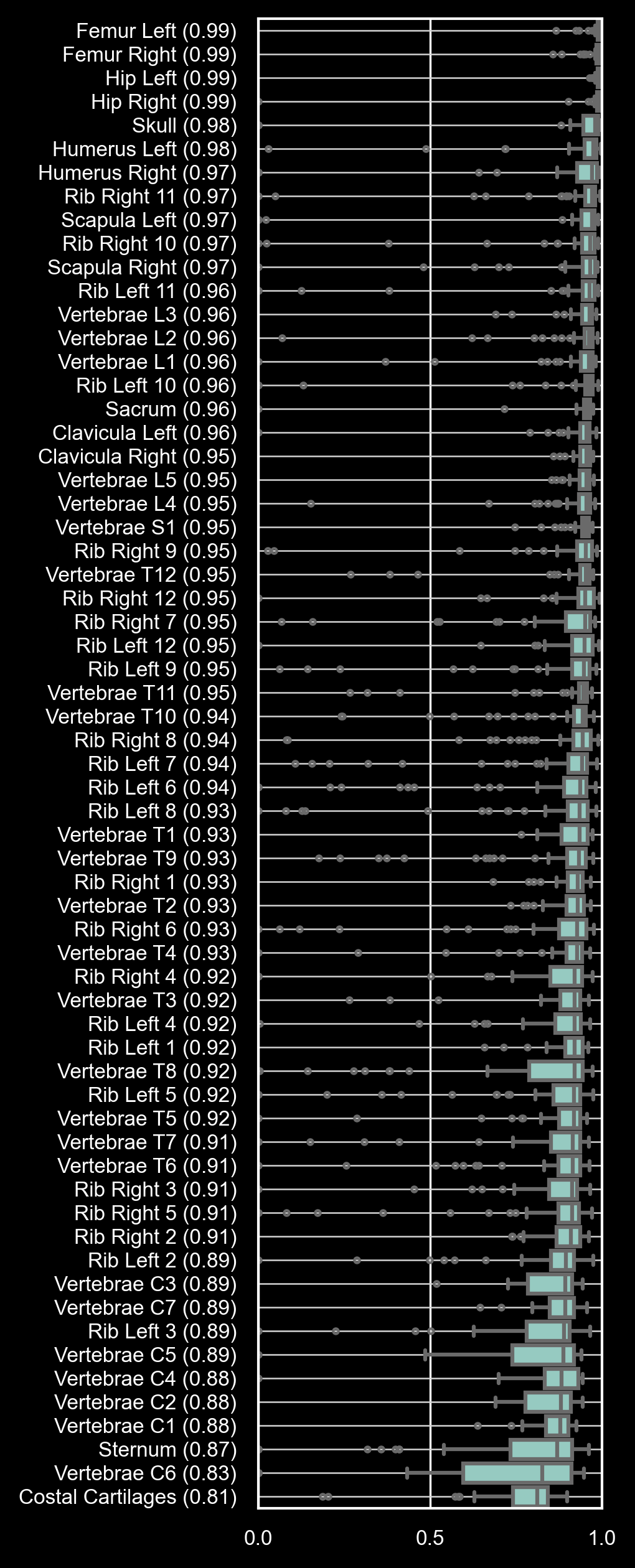
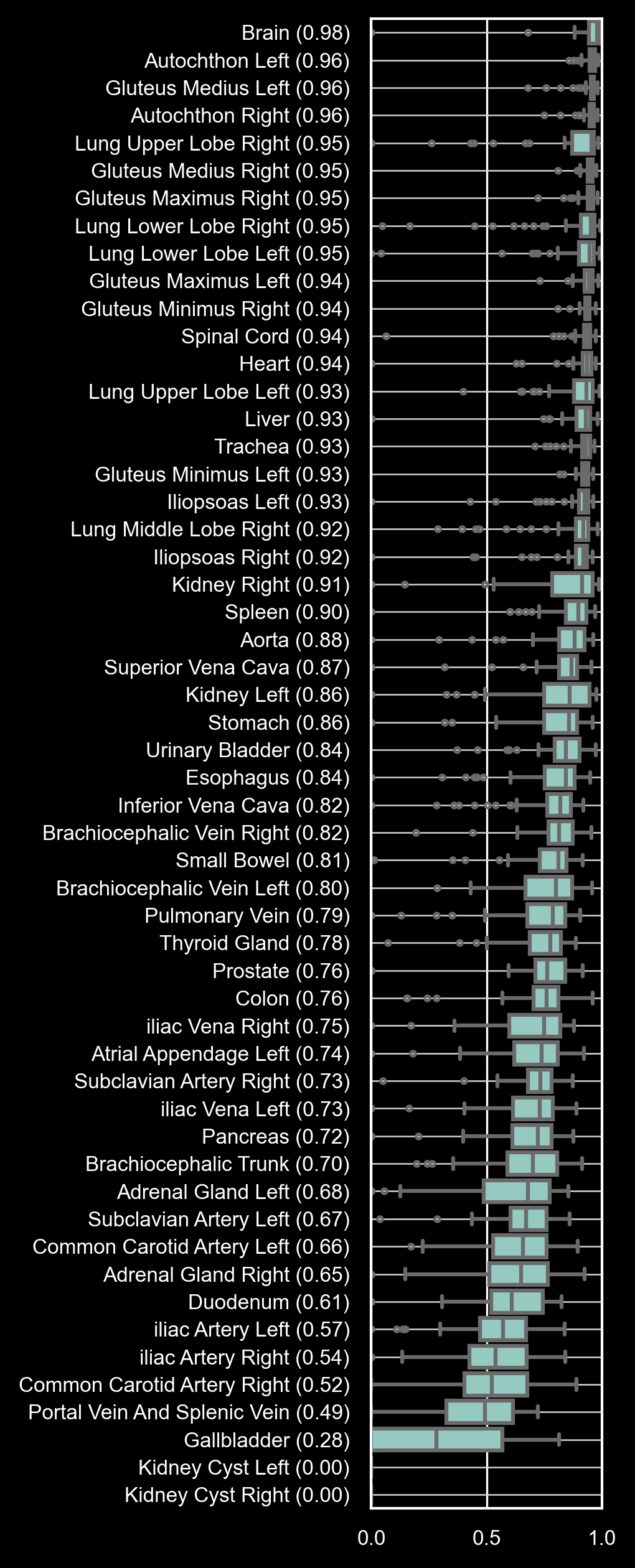
The images below illustrate segmentation results for bone tissue (left) and soft tissue (right) using the ts2d-v2 models.
Dice Similarity Coefficients (DSC) are reported for all 117 anatomical structures, sorted from highest to lowest median DSC.
TS2D is recommended for bone tissue and most soft tissue structures; however, caution is advised for smaller soft tissue structures with lower DSC values.
Publications
Our following publications are related to the development and application of TS2D:
-
Original publication introducing TS2D:
- [MIUA2025a] TotalSegmentator 2D: A Tool for Rapid Anatomical Structure Analysis
Presented at Medical Image Understanding and Analysis (MIUA) Conference 2025
Full Reference:Sabrowsky-Hirsch, B., Alshenoudy, A., Thumfart, S., Giretzlehner, M. (2025). TotalSegmentator 2D (TS2D): A Tool for Rapid Anatomical Structure Analysis. Medical Image Understanding and Analysis 2025 (MIUA 2025). Springer Nature.
- [MIUA2025a] TotalSegmentator 2D: A Tool for Rapid Anatomical Structure Analysis
-
TS2D extended to the segmentation of X-Ray images:
- [MIUA2025b] Leveraging Synthetic Data for Whole-Body Segmentation in X-ray Images
Presented at Medical Image Understanding and Analysis (MIUA) Conference 2025
Full Reference:Alshenoudy, A., Sabrowsky-Hirsch, B., Thumfart, S., Giretzlehner, M. (2025). Leveraging Synthetic Data for Whole-Body Segmentation in X-Ray Images. Medical Image Understanding and Analysis 2025.
- [MIUA2025b] Leveraging Synthetic Data for Whole-Body Segmentation in X-ray Images
-
Our earlier work on body-region segmentation for an industrial usecase:
- [AIROV2025] Efficient Automatic Detection of Scanned Body Regions in CT Scans
Presented at Austrian Symposium on AI, Robotics, and Vision (AIRoV) Conference 2025
Full Reference:Sabrowsky-Hirsch, Bertram, et al. “Efficient Automatic Detection of Scanned Body Regions in CT Scans.” In Proceedings of the Joint Austrian Computer Vision and Robotics Workshop 2025. Verlag der TU Graz (2025).
- [AIROV2025] Efficient Automatic Detection of Scanned Body Regions in CT Scans
References
TotalSegmentator 2D builds upon two key works in the field of medical image segmentation:
-
Isensee et al. (2021): nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation. Nature Methods, 18, 203–211. https://doi.org/10.1038/s41592-020-01008-z
-
Wasserthal et al. (2023): TotalSegmentator: Robust Segmentation of 104 Anatomic Structures in CT Images. Radiology: Artificial Intelligence. https://doi.org/10.1148/ryai.230024
Contact
If you have any inquiries, please open a GitHub issue.
Links
GitHub · PyPI · Zenodo · RISC Software GmbH
Acknowledgements


This project is financed by research subsidies granted by the government of Upper Austria. RISC Software GmbH is Member of UAR (Upper Austrian Research) Innovation Network.
Versions
- v1.0.0: first release of TS2D including the [MIUA2025a] models.
- v1.1.0: added models trained on the TotalSegmentator v2 dataset
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file ts2d-1.1.3.tar.gz.
File metadata
- Download URL: ts2d-1.1.3.tar.gz
- Upload date:
- Size: 55.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cdb996961cbc2a344735d9f468f0026f6153bb55902eb326345d0e211dd00915
|
|
| MD5 |
5e3a14e49ac1afe238febf018f0b0872
|
|
| BLAKE2b-256 |
cd12fd091c61ab83180c128c33f257491dead3433d0e383149fa3403c7ae0a5c
|
File details
Details for the file ts2d-1.1.3-py3-none-any.whl.
File metadata
- Download URL: ts2d-1.1.3-py3-none-any.whl
- Upload date:
- Size: 57.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.11
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8fe528dbfdb86f6e1838aa8c72757fe83011b1703635148f9131cb7f8c15da0c
|
|
| MD5 |
e9f41170cfd48b59493936cc7b8b1ffa
|
|
| BLAKE2b-256 |
2021e72c648e820d089bd48a4ad5466116fbad3ce6c3798969259451d4d39b5a
|











