A Python wrapper for the UniProt Mapping RESTful API.
Project description
UniProtMapper 
Easily retrieve UniProt data and map protein identifiers using this Python package for UniProt's Retrieve & ID Mapping RESTful APIs. Read the full documentation.
📚 Table of Contents
⛏️ Features
UniProtMapper is a tool for bioinformatics and proteomics research that supports:
- Mapping any UniProt cross-referenced IDs to other identifiers & vice-versa;
- Programmatically retrieving any of the supported return and cross-reference fields from both UniProt-SwissProt and UniProt-TrEMBL (unreviewed) databases. For a full table containing all the supported resources, refer to the supported fields in the docs;
- Querying UniProtKB entries using complex field-based queries with boolean operators
~(NOT),|(OR),&(AND).
For the first two functionalities, check the examples Mapping IDs and Retrieving Information below. The third, see Field-based Querying.
The ID mapping API can also be accessed through the CLI. For more information, check CLI.
📦 Installation
From PyPI (recommended):
python -m pip install uniprot-id-mapper
Directly from GitHub:
python -m pip install git+https://github.com/David-Araripe/UniProtMapper.git
From source:
git clone https://github.com/David-Araripe/UniProtMapper
cd UniProtMapper
python -m pip install .
🛠️ Usage
Mapping IDs
Use UniProtMapper to easily map between different protein identifiers:
from UniProtMapper import ProtMapper
mapper = ProtMapper()
result, failed = mapper.get(
ids=["P30542", "Q16678", "Q02880"], from_db="UniProtKB_AC-ID", to_db="Ensembl"
)
The result is a pandas DataFrame containing the mapped IDs (see below), while failed is a list of identifiers that couldn't be mapped.
| UniProtKB_AC-ID | Ensembl | |
|---|---|---|
| 0 | P30542 | ENSG00000163485.17 |
| 1 | Q16678 | ENSG00000138061.12 |
| 2 | Q02880 | ENSG00000077097.17 |
Retrieving Information
A DataFrame with the supported return fields is accessible through the attribute ProtMapper.fields_table:
from UniProtMapper import ProtMapper
mapper = ProtMapper()
df = mapper.fields_table
df.head()
| label | returned_field | field_type | has_full_version | type | |
|---|---|---|---|---|---|
| 0 | Entry | accession | Names & Taxonomy | - | uniprot_field |
| 1 | Entry Name | id | Names & Taxonomy | - | uniprot_field |
| 2 | Gene Names | gene_names | Names & Taxonomy | - | uniprot_field |
| 3 | Gene Names (primary) | gene_primary | Names & Taxonomy | - | uniprot_field |
| 4 | Gene Names (synonym) | gene_synonym | Names & Taxonomy | - | uniprot_field |
From the DataFrame, all return_field entries can be used to access UniProt data programmatically:
# To retrieve the default fields:
result, failed = mapper.get(["Q02880"])
>>> Fetched: 1 / 1
# Retrieve custom fields:
fields = ["accession", "organism_name", "structure_3d"]
result, failed = mapper.get(["Q02880"], fields=fields)
>>> Fetched: 1 / 1
Further, for the cross-referenced fields that have has_full_version set to yes, returning the same field with extra information is supported by passing <field_name>_full, such as xref_pdb_full.
All available return fields are also accessible through the attribute ProtMapper.supported_return_fields:
from UniProtMapper import ProtMapper
mapper = ProtMapper()
print(mapper.supported_return_fields)
>>> ['accession',
>>> 'id',
>>> 'gene_names',
>>> ...
>>> 'xref_smart_full',
>>> 'xref_supfam_full']
Field-based Querying
UniProtMapper supports complex field-based protein queries using boolean operators (AND, OR, NOT) through the uniprotkb_fields module. This allows you to create sophisticated searches combining multiple criteria. For example:
from UniProtMapper import ProtKB
from UniProtMapper.uniprotkb_fields import (
organism_name,
length,
reviewed,
date_modified
)
# Find reviewed human proteins with length between 100-200 amino acids
# that were modified after January 1st, 2024
query = (
organism_name("human") &
reviewed(True) &
length(100, 200) &
date_modified("2024-01-01", "*")
)
protkb = ProtKB()
result = protkb.get(query)
For a list of all fields and their descriptions, check the API reference for the uniprotkb_fields module reference.
📖 Documentation
- Stable Branch Documentation (master branch)
- Development Documentation (dev branch)
💻 Command Line Interface (CLI)
UniProtMapper provides a CLI for the ID Mapping class, ProtMapper, for easy access to lookups and data retrieval. Here is a list of the available arguments, shown by protmap -h:
usage: UniProtMapper [-h] -i [IDS ...] [-r [RETURN_FIELDS ...]] [--default-fields] [-o OUTPUT]
[-from FROM_DB] [-to TO_DB] [-over] [-pf]
Retrieve data from UniProt using UniProt's RESTful API. For a list of all available fields, see: https://www.uniprot.org/help/return_fields
Alternatively, use the --print-fields argument to print the available fields and exit the program.
optional arguments:
-h, --help show this help message and exit
-i [IDS ...], --ids [IDS ...]
List of UniProt IDs to retrieve information from. Values must be
separated by spaces.
-r [RETURN_FIELDS ...], --return-fields [RETURN_FIELDS ...]
If not defined, will pass `None`, returning all available fields.
Else, values should be fields to be returned separated by spaces. See
--print-fields for available options.
--default-fields, -def
This option will override the --return-fields option. Returns only the
default fields stored in: <pkg_path>/resources/cli_return_fields.txt
-o OUTPUT, --output OUTPUT
Path to the output file to write the returned fields. If not provided,
will write to stdout.
-from FROM_DB, --from-db FROM_DB
The database from which the IDs are. For the available cross
references, see: <pkg_path>/resources/uniprot_mapping_dbs.json
-to TO_DB, --to-db TO_DB
The database to which the IDs will be mapped. For the available cross
references, see: <pkg_path>/resources/uniprot_mapping_dbs.json
-over, --overwrite If desired to overwrite an existing file when using -o/--output
-pf, --print-fields Prints the available return fields and exits the program.
Usage example, retrieving default fields from <pkg_path>/resources/cli_return_fields.txt:
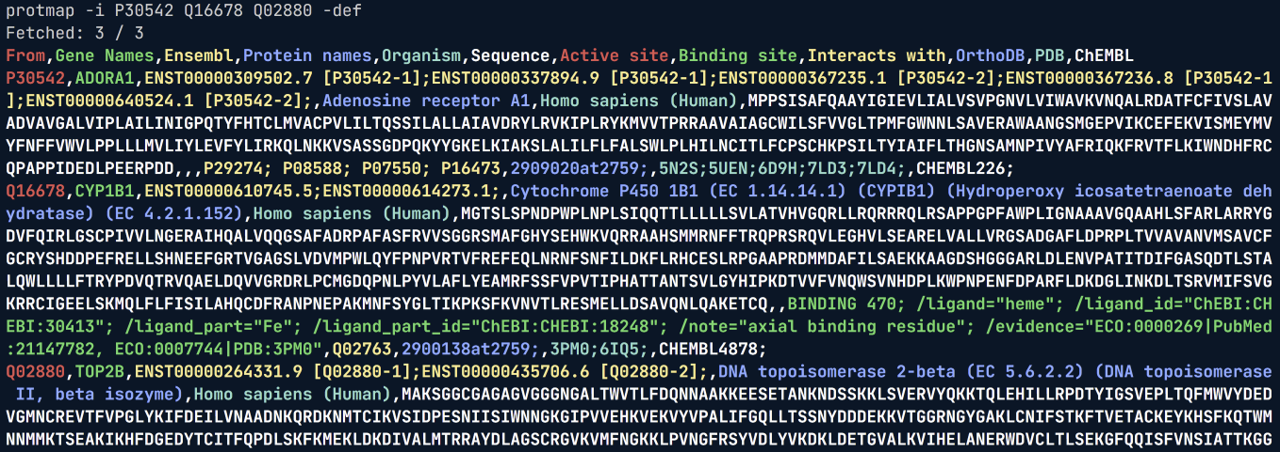
👏🏼 Credits
- UniProt for providing the API and the amazing database;
- Andrew White and the University of Rochester for the protein emoji;
For issues, feature requests, or questions, please open an issue on the GitHub repository.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file uniprot_id_mapper-1.1.5-py3-none-any.whl.
File metadata
- Download URL: uniprot_id_mapper-1.1.5-py3-none-any.whl
- Upload date:
- Size: 46.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dc8627ea24ec1dfe1a07783dc05195db0751270ab9342ef1a1091dcfe4b04f7f
|
|
| MD5 |
7109f8709317c97059aabc052c372a11
|
|
| BLAKE2b-256 |
11c265600302d40dadd0aca374823ede7bb073dbbd3cb78aa1c283eebe9cbe18
|

















