Efficient variant screening of RNA-seq and DNA-seq data using k-mer-based alignment against a reference of known variants.
Project description
varseek
varseek is a free, open-source command-line tool and Python package that provides variant screening of RNA-seq and DNA-seq data using k-mer-based alignment against a reference of known variants. The name comes from "seeking variants" or, alternatively, "seeing k-variants" (where a "k-variant" is defined as a k-mer containing a variant).
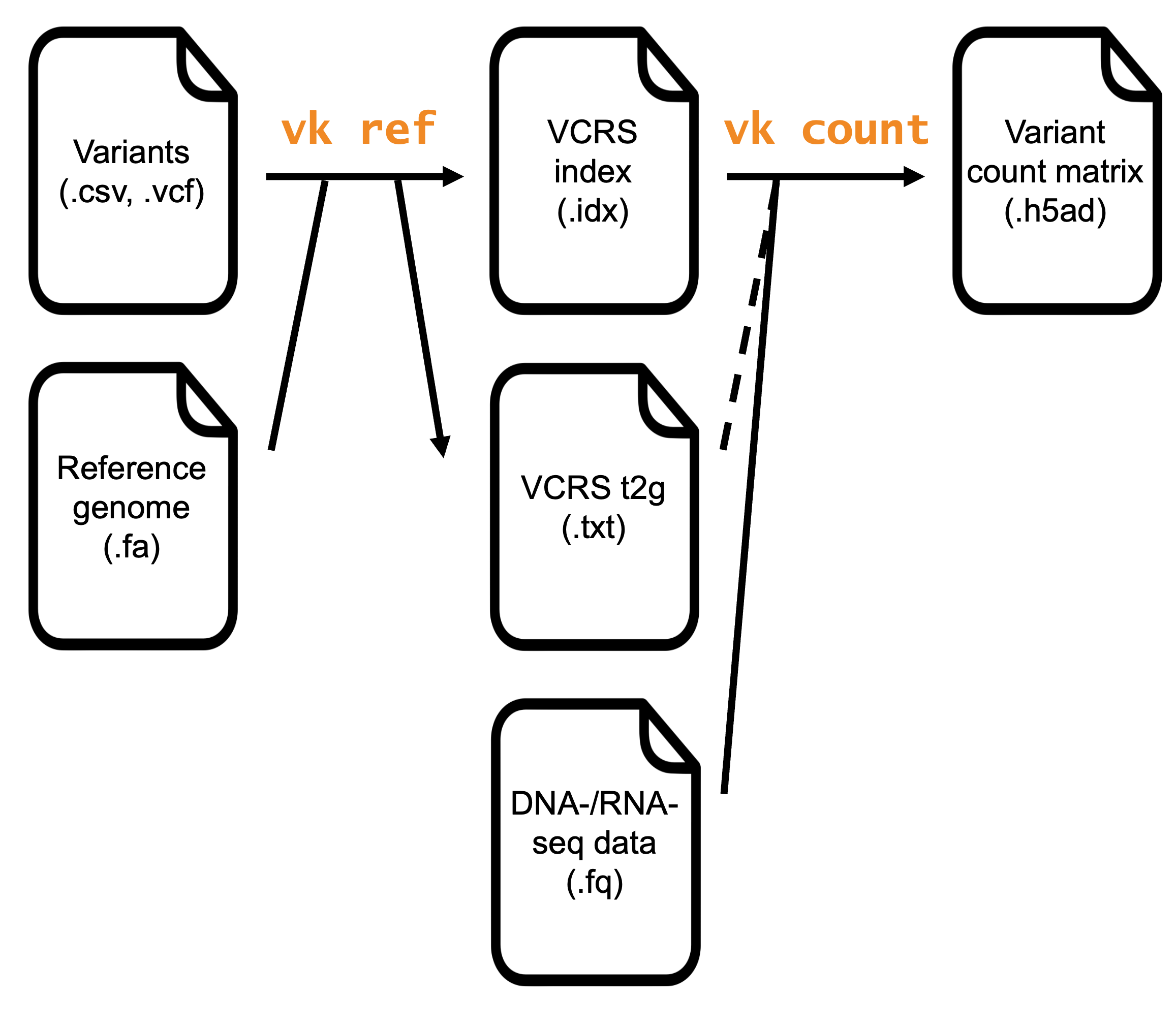
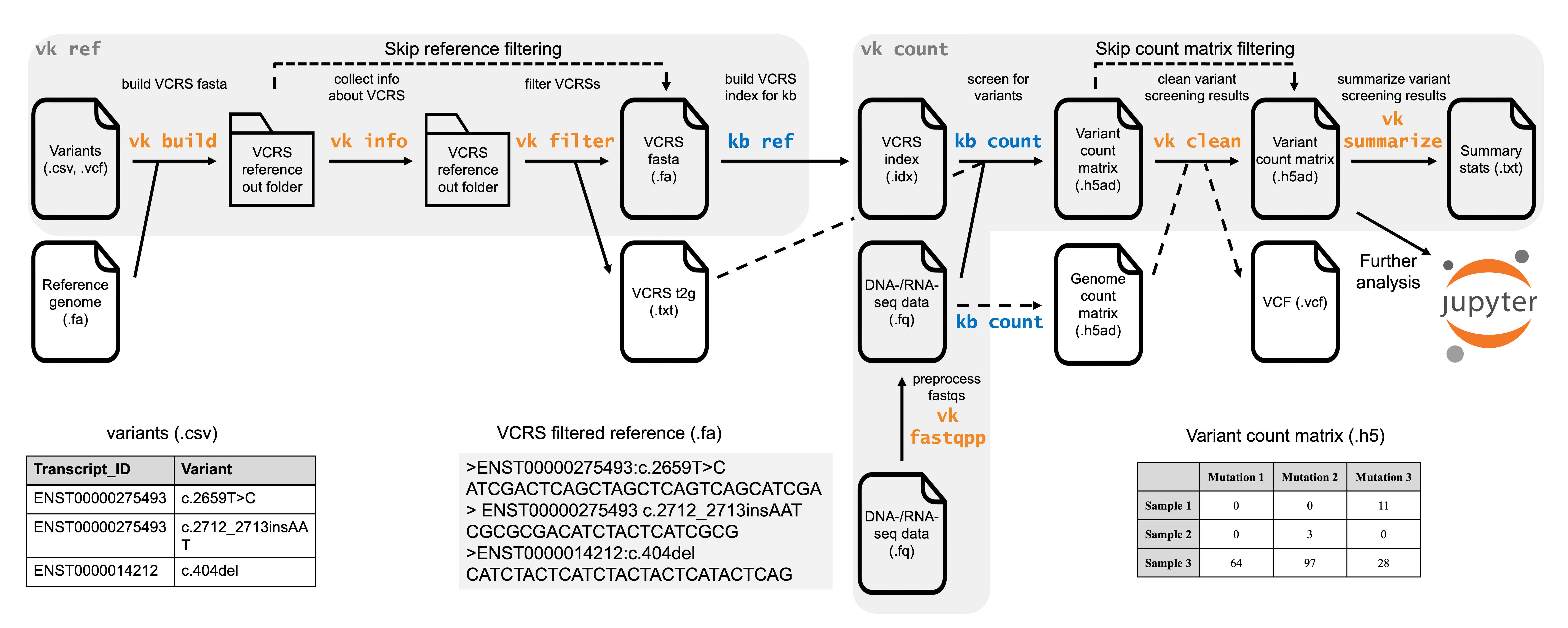
The two commands used in a standard workflow are varseek ref and varseek count. varseek ref generates a variant-containing reference sequence (VCRS) index that serves as the basis for variant calling. varseek count pseudoaligns RNA-seq or DNA-seq reads against the VCRS index and generates a variant count matrix. The variant count matrix can be used for downstream analysis. Each step wraps around other steps within the varseek package and the kb-python package, as described below.
The functions of varseek are described in the table below.
| Description | Bash | Python (with import varseek as vk) |
|---|---|---|
| Build a variant-containing reference sequence (VCRS) fasta file | vk build ... |
vk.build(...) |
| Describe the VCRS reference in a dataframe for filtering | vk info ... |
vk.info(...) |
| Filter the VCRS file based on the CSV generated from varseek info | vk filter ... |
vk.filter(...) |
| Preprocess the FASTQ files before pseudoalignment | vk fastqpp ... |
vk.fastqpp(...) |
| Process the variant count matrix | vk clean ... |
vk.clean(...) |
| Analyze the variant count matrix results | vk summarize ... |
vk.summarize(...) |
| Wrap vk build, vk info, vk filter, and kb ref | vk ref ... |
vk.ref(...) |
| Wrap vk fastqpp, kb count, vk clean, and vk summarize | vk count ... |
vk.count(...) |
| Create synthetic RNA-seq dataset with variant-containing reads | vk sim ... |
vk.sim(...) |
After aligning and generating a variant count matrix with varseek, you can explore the data using our pre-built notebooks. The notebooks are described in the table below.
| Description | Notebook |
|---|---|
| Preprocessing the variant count matrix | 3_matrix_preprocessing.ipynb |
| Sequence visualization of variants | 4_1_variant_analysis_sequence_visualization.ipynb |
| Heatmap visualization of variant patterns | 4_2_variant_analysis_heatmaps.ipynb |
| Protein-level variant analysis | 4_3_variant_analysis_protein_variant.ipynb |
| Heatmap analysis of gene expression | 5_1_gene_analysis_heatmaps.ipynb |
| Drug-target analysis for genes | 5_2_gene_analysis_drugs.ipynb |
| Pathway analysis using Enrichr | 6_1_pathway_analysis_enrichr.ipynb |
| Gene Ontology enrichment analysis (GOEA) | 6_2_pathway_analysis_goea.ipynb |
You can find more examples of how to use varseek in the GitHub repository for our preprint GitHub - pachterlab/RLSRWP_2025.
If you use varseek in a publication, please cite the following study:
PAPER CITATION
Read the article here: PAPER DOI
Installation
pip install varseek
🪄 Quick start guide
1. Acquire a Reference
Follow one of the below options:
a. Download a Pre-built Reference
- (optional) View all downloadable references:
vk ref --list_downloadable_references vk ref --download --variants VARIANTS --sequences SEQUENCES
b. Make custom reference – screen for user-defined variants
vk ref --variants VARIANTS --sequences SEQUENCES ...
c. Customize reference building process – customize the VCRS filtering process (e.g., add additional information by which to filter, add custom filtering logic, tune filtering parameters based on the results of intermediate steps, etc.)
vk build --variants VARIANTS --sequences SEQUENCES ...- (optional)
vk info --input_dir INPUT_DIR ... - (optional)
vk filter --input_dir INPUT_DIR ... kb ref --workflow custom --index INDEX ...
2. Screen for variants
Follow one of the below options:
a. Standard workflow
- (optional) fastq quality control
vk count --index INDEX --t2g T2G ... --fastqs FASTQ1 FASTQ2...
b. Customize variant screening process - additional fastq preprocessing, custom count matrix processing
- (optional) fastq quality control
- (optional)
vk fastqpp ... --fastqs FASTQ1 FASTQ2... kb count --index INDEX --t2g T2G ... --fastqs FASTQ1 FASTQ2...- (optional)
kb count --index REFERENCE_INDEX --t2g REFERENCE_T2G ... --fastqs FASTQ1 FASTQ2... - (optional)
vk clean --adata ADATA ... - (optional)
vk summarize --adata ADATA ...
Examples for getting started: GitHub - pachterlab/varseek Manuscript: ... Repository for manuscript figures: GitHub - pachterlab/RLSRP_2025
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file varseek-0.1.1.tar.gz.
File metadata
- Download URL: varseek-0.1.1.tar.gz
- Upload date:
- Size: 237.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d6765f9fda272be7d1855106d5a52acf43500795b0002480d2b0f7600dbf011a
|
|
| MD5 |
79bbfa0214253557c805b2299e2aa686
|
|
| BLAKE2b-256 |
7d518aacda62660f88b7283865a76ac330574d64a2f83d6aeab3bc62d858248f
|
File details
Details for the file varseek-0.1.1-py3-none-any.whl.
File metadata
- Download URL: varseek-0.1.1-py3-none-any.whl
- Upload date:
- Size: 234.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
85be51268e0fd2359d8adb9431f31dac41e47c83e44b1e9e10212820469c02d8
|
|
| MD5 |
9aff5806f4f96e4b36be66c401e7c3a1
|
|
| BLAKE2b-256 |
963fda5133a35fc41ab16a7ae9e8f7dd6c4606cfdd71864a618e30a4cd050ec8
|