VitalPy: A Vital Signal Analysis Package
Project description
VitalPy: A Vital Signal Analysis Package
Currently, the package supports PPG preprocessing and extraction of more than 400 features. The PPG pipeline was originally implemented for analysis of the AuroraBP database.
It provides:
- PPG preprocessing: Singal quality metrics, baseline extraction, etc.
- PPG feature extraction: time-domain, frequency-domain, statistical features ( >400 features)
- Compatibility with PPG recorded from 128 Hz to 500 Hz: tested with local devices and large datasets.
Use
VitalPy is written in Python (3.9+). Navigate to the Python repository and install the required packages:
pip install -r requirements.txt
Import:
from src.ppg.PPGSignal import PPGSignal
Check the signal (make sure that the file waveform_df is in dataframe format and contains the columns 't' for time and 'ppg' for the signal values):
signal = PPGSignal(waveform_df, verbose=1)
signal.check_keypoints()
Get features:
signal = PPGSignal(waveform_df, verbose=0)
features = signal.extract_features()
Example plots for a AuroraBP signal
Used file: measurements_oscillometric/o001/o001.initial.Sitting_arm_down.tsv
Mean signal
The following figure shows the mean template computed from all templates within the signal given as input.
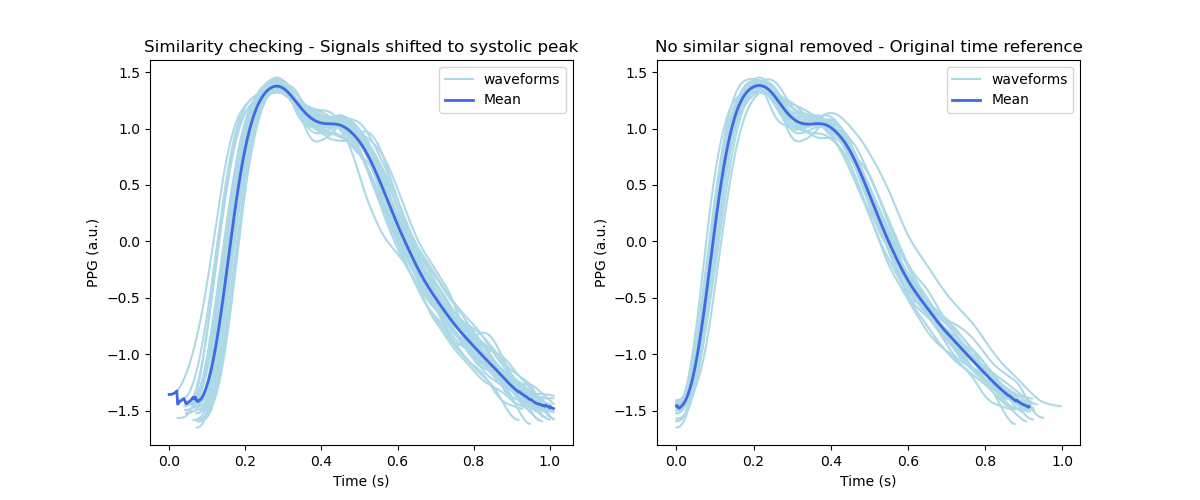
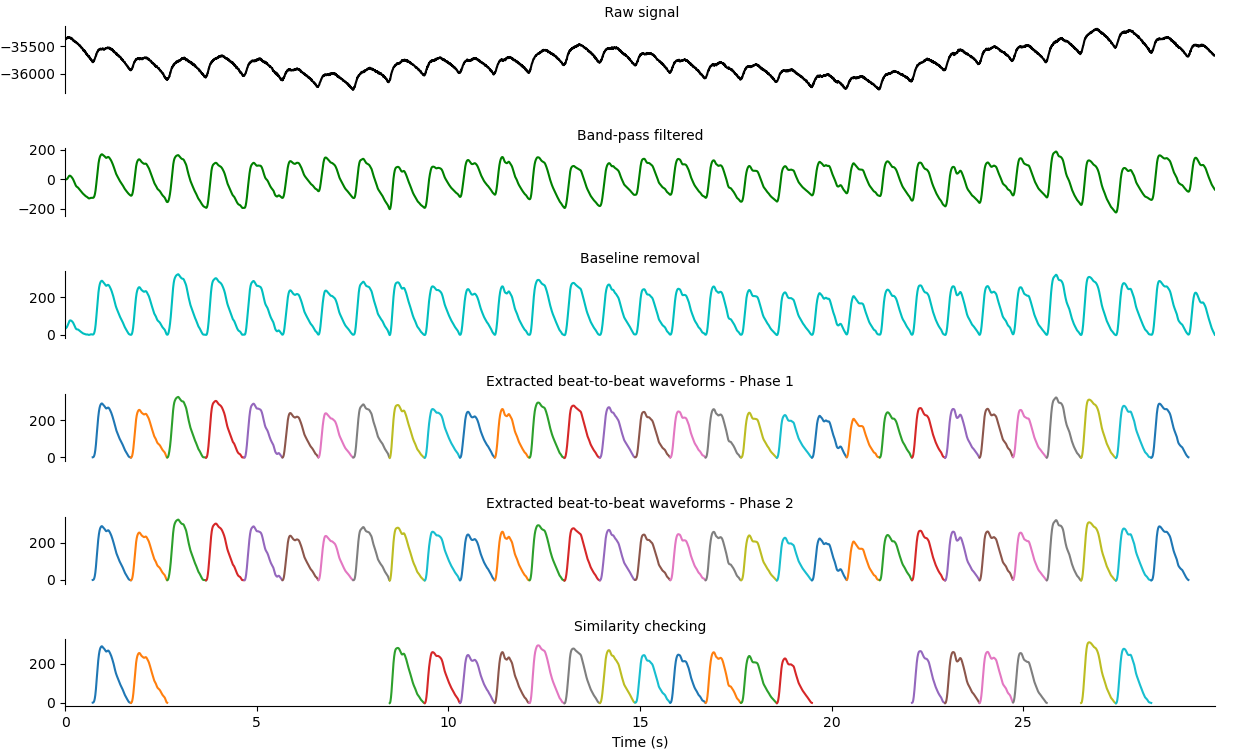
Preprocessing steps
All preprocessing steps are depicted. The final result should have filtered out all low quality waveforms.
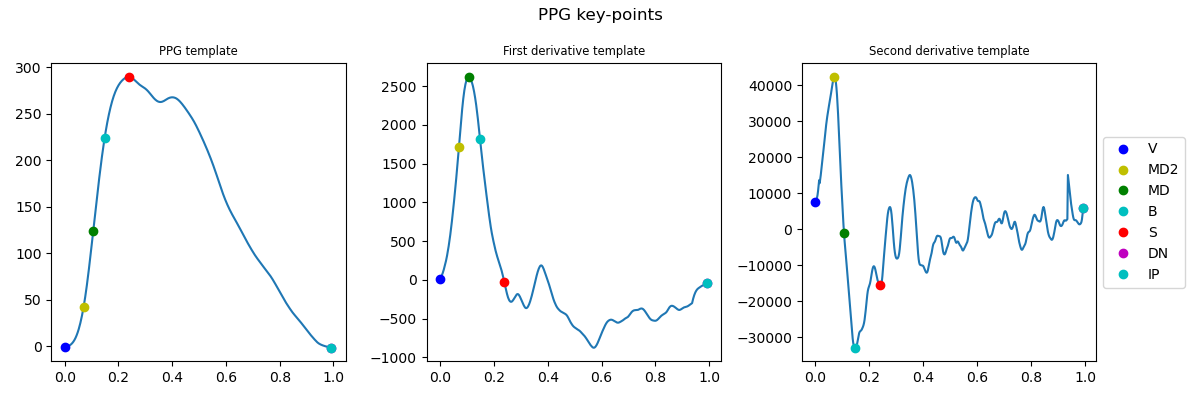
PPG Keypoints
Exemplary PPG keypoint extraction.
License
VitalPy is available under the General Public License v3.0.
Citation
If you use this repository or any of its components and/or our paper as part of your research, please cite the publication as follows:
A. Cisnalet al. "Robust Feature Selection for Continuous BP Estimation in Multiple Populations: Towards Cuffless Ambulatory BP Monitoring," IEEE J Biomed Health Inform, Under Review (2023).
@unpublished{vitalpy,
title={Robust Feature Selection for Continuous BP Estimation in Multiple Populations: Towards Cuffless Ambulatory BP Monitoring},
author={Cisnal, Ana and Li, Yanke and Fuchs, Bertram and Ejtehadi, Mehdi and Riener, Robert and Paez-Granados, Diego},
journal={IEEE Journal of Biomedical and Health Informatics},
year={2023},
note = "Under review"
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file vitalpython-0.1.0.tar.gz.
File metadata
- Download URL: vitalpython-0.1.0.tar.gz
- Upload date:
- Size: 193.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.9.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a1e1fd5951954691af253d13f9d9ea2eac67f13bda52f641fadae0285c801de5
|
|
| MD5 |
c394b33ad52893e0dc6fa0c9b553f0ec
|
|
| BLAKE2b-256 |
e93c88938ea582db7a18ccbc25d9f72fabae23c02af41bb83f82f718bd995fe3
|
File details
Details for the file vitalpython-0.1.0-py3-none-any.whl.
File metadata
- Download URL: vitalpython-0.1.0-py3-none-any.whl
- Upload date:
- Size: 45.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.9.18
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cd71cc3e967cd98c399b8c4ca3a3e5d63a2d70587b66aaa81dce9656fb59050f
|
|
| MD5 |
ac704da72f3ef8f068d34e265747b9a2
|
|
| BLAKE2b-256 |
f97904d4483d69059b72240ef433b14c257f60434467a63207b332b256b9d7ce
|











