Package for parsing, writing, and modifying molecular structure files
Project description


A python package for parsing, modifying, and analysis of molecular structure files.
Installation
Easiest way to install is from PyPI:
pip install xchem-molparse
Usage
Python Module
MolParse is primarily a python module which can be used interactively, or within (batch) scripts:
Use pydoc to see help on the molparse module, or its methods & classes. E.g. from a shell:
pydoc molparse
pydoc molparse.System
Binaries and command-line programs
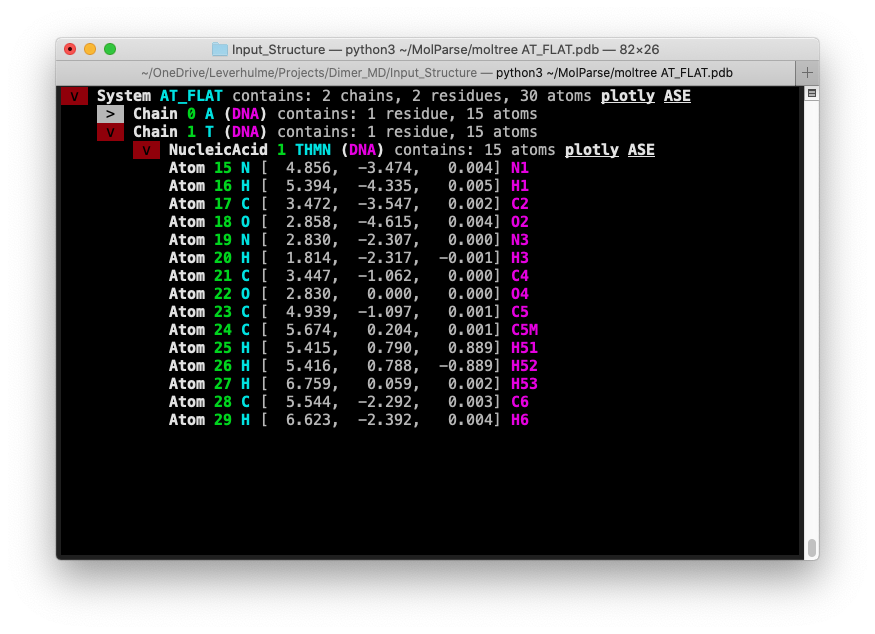
moltree
In addition to the python module, an interactive command-line interface is available with the binary moltree. Pass a
PDB or GRO file as follows:
moltree <FILE>
Use the mouse to interact with buttons and CTR-C to exit.
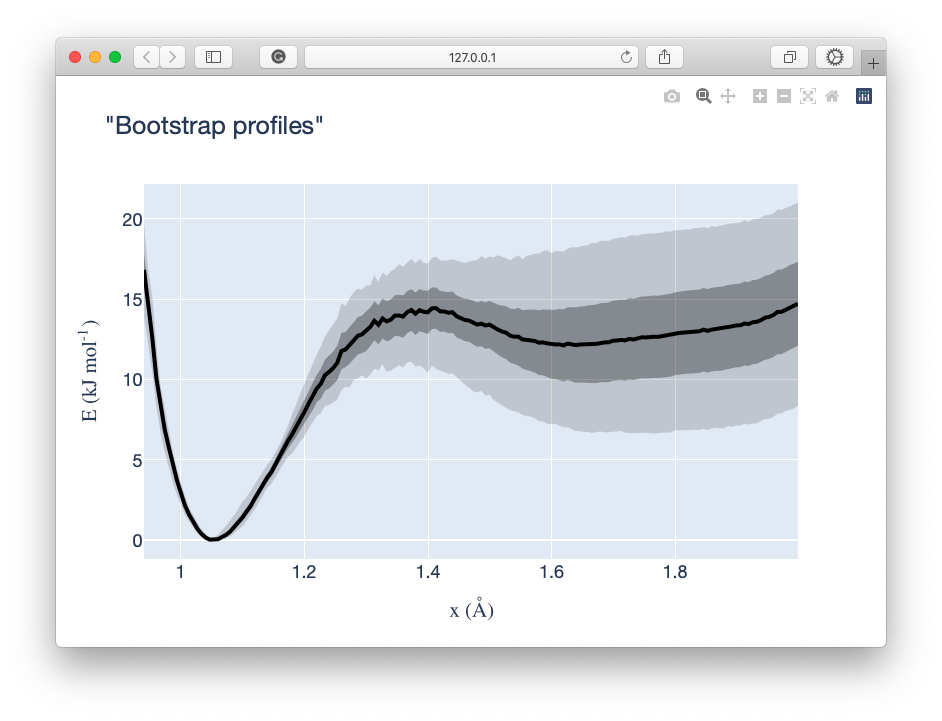
molxvg
Gromacs produces data files in XVG format by default, these can be parsed using the molparse.xvg.parseXVG method from
within a python environment, alternatively a binary exists to access its basic functionality from the command line. Run
the following to open an interactive plotly graph of an xvg:
molxvg [FILE.xvg] -s
Other options can be found by running molxvg --help.
Installation from source
Requirements
Installing ASE
pip install --upgrade --user aseexport PATH=$PATH:~/.local/binto your.bash_profileexport PYTHONPATH=$PYTHONPATH:~/.local/lib/python3.X/site-packagesto your.bash_profileWhere X is your python version.
MPyTools
git clone https://github.com/mwinokan/MPyTools.git- Add
export MPYTOOLS=/path/to/directoryto your.bash_profile - Add
export PYTHONPATH=$PYTHONPATH:$MPYTOOLSto your.bash_profile
MolParse
git clone https://github.com/xchem/MolParse.git- Add
export MOLPARSE=/path/to/directoryto your.bash_profile - Add
export PYTHONPATH=$PYTHONPATH:$MOLPARSEto your.bash_profile
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file xchem_molparse-1.0.0.tar.gz.
File metadata
- Download URL: xchem_molparse-1.0.0.tar.gz
- Upload date:
- Size: 145.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d692a4f4ae59789044693269449b7b81875fc62e23111c7c71666e0d29c32473
|
|
| MD5 |
d0db2edab6694aa9c07044c664a56ac7
|
|
| BLAKE2b-256 |
ce4a0860f7ff079a7595ebcd38d27188b8718bddb22cec0cdb673e4dfb774098
|
Provenance
The following attestation bundles were made for xchem_molparse-1.0.0.tar.gz:
Publisher:
release.yaml on xchem/MolParse
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
xchem_molparse-1.0.0.tar.gz -
Subject digest:
d692a4f4ae59789044693269449b7b81875fc62e23111c7c71666e0d29c32473 - Sigstore transparency entry: 1051739521
- Sigstore integration time:
-
Permalink:
xchem/MolParse@b9bbde95dfbaae9dc2e46fe14ccc37b95e00b1b6 -
Branch / Tag:
refs/tags/1.0.0 - Owner: https://github.com/xchem
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yaml@b9bbde95dfbaae9dc2e46fe14ccc37b95e00b1b6 -
Trigger Event:
release
-
Statement type:
File details
Details for the file xchem_molparse-1.0.0-py3-none-any.whl.
File metadata
- Download URL: xchem_molparse-1.0.0-py3-none-any.whl
- Upload date:
- Size: 176.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d15bb40acd4d2330e3ce21b6925db35ccf7443a27a55ecd5f06384890cb2a48c
|
|
| MD5 |
eccc50fce1bbab8c155898f58e512d93
|
|
| BLAKE2b-256 |
76d27542e0c145337d8c494f2b4ebdaa0cba83592d20c3b525ca460163094af4
|
Provenance
The following attestation bundles were made for xchem_molparse-1.0.0-py3-none-any.whl:
Publisher:
release.yaml on xchem/MolParse
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
xchem_molparse-1.0.0-py3-none-any.whl -
Subject digest:
d15bb40acd4d2330e3ce21b6925db35ccf7443a27a55ecd5f06384890cb2a48c - Sigstore transparency entry: 1051739527
- Sigstore integration time:
-
Permalink:
xchem/MolParse@b9bbde95dfbaae9dc2e46fe14ccc37b95e00b1b6 -
Branch / Tag:
refs/tags/1.0.0 - Owner: https://github.com/xchem
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
release.yaml@b9bbde95dfbaae9dc2e46fe14ccc37b95e00b1b6 -
Trigger Event:
release
-
Statement type: