Automatic multiple sclerosis lesion segmentation in MRI using YOLO11-seg
Project description

🧠💻 YOLO-MSLesSeg: Automatic Multiple Sclerosis Lesion Segmentation with YOLO11-seg
yolo_mslesseg is a Python package that provides a complete pipeline for the automatic segmentation and evaluation of multiple sclerosis lesions in MRI images, using YOLO11-seg models. The system is based on the MSLesSeg Competition (ICPR 2024) dataset, an international benchmark for the validation of automatic methods for multiple sclerosis lesion segmentation.
The package introduces an approach that combines deep learning models with different image enhancement techniques as a preprocessing stage, with the goal of quantifying lesions consistently and reducing the variability associated with manual segmentation.
Table of Contents
- Visual examples
- Pipeline overview
- Installation
- Repository structure
- Running the pipeline
- References
- License
- Citation
Visual Examples
Below are representative examples of the outputs generated by the pipeline. These visualisations allow a direct appreciation of the type of segmentations produced by the model and their anatomical consistency across the different viewing planes. A GIF is also included that traverses all slices of a patient, showing the consistency of predictions across the entire volume.
Segmentation across the three anatomical planes
The following example corresponds to a reference patient (P1) and shows the overlay of the automatic segmentation on the FLAIR image in the axial, coronal, and sagittal planes.
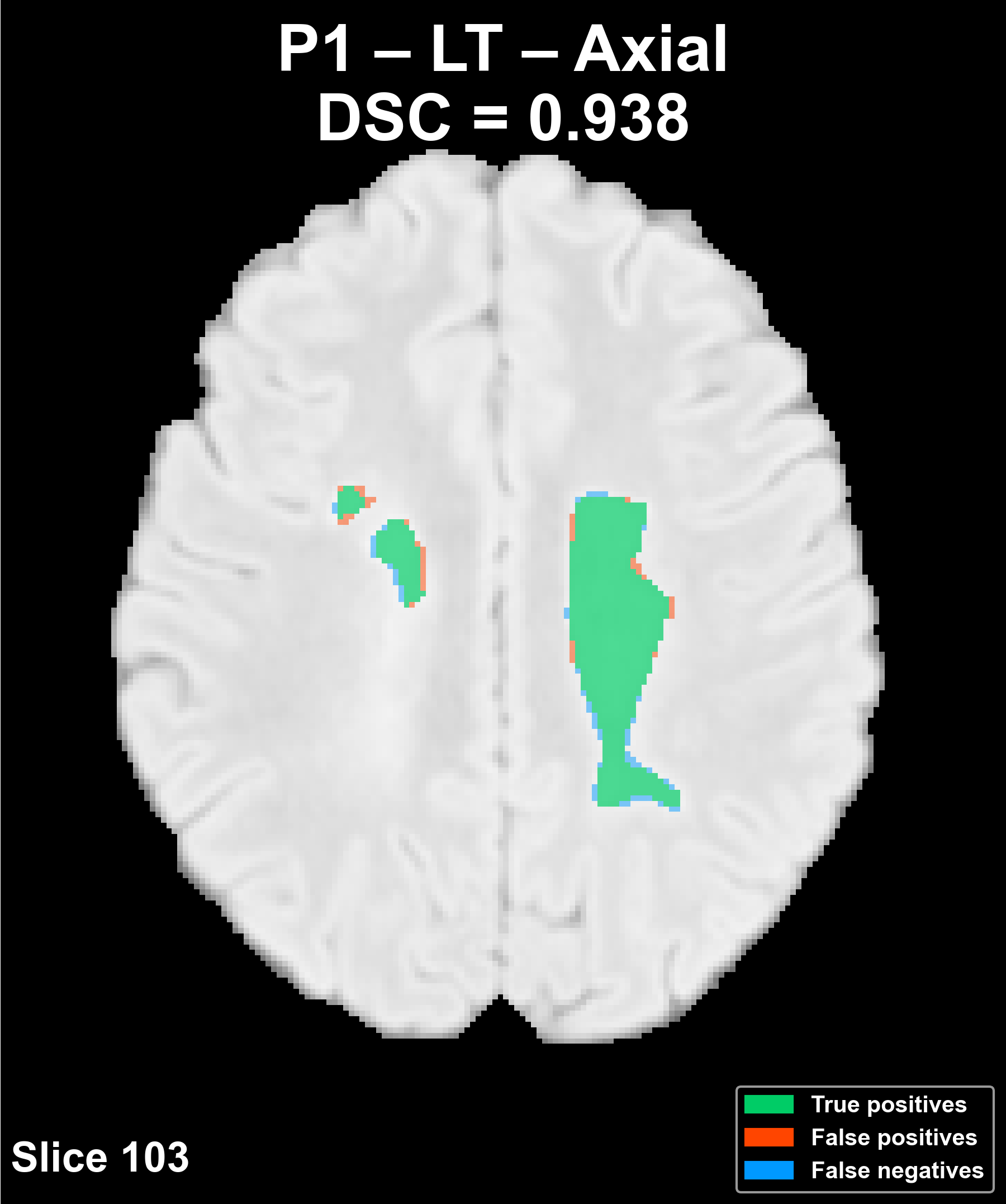
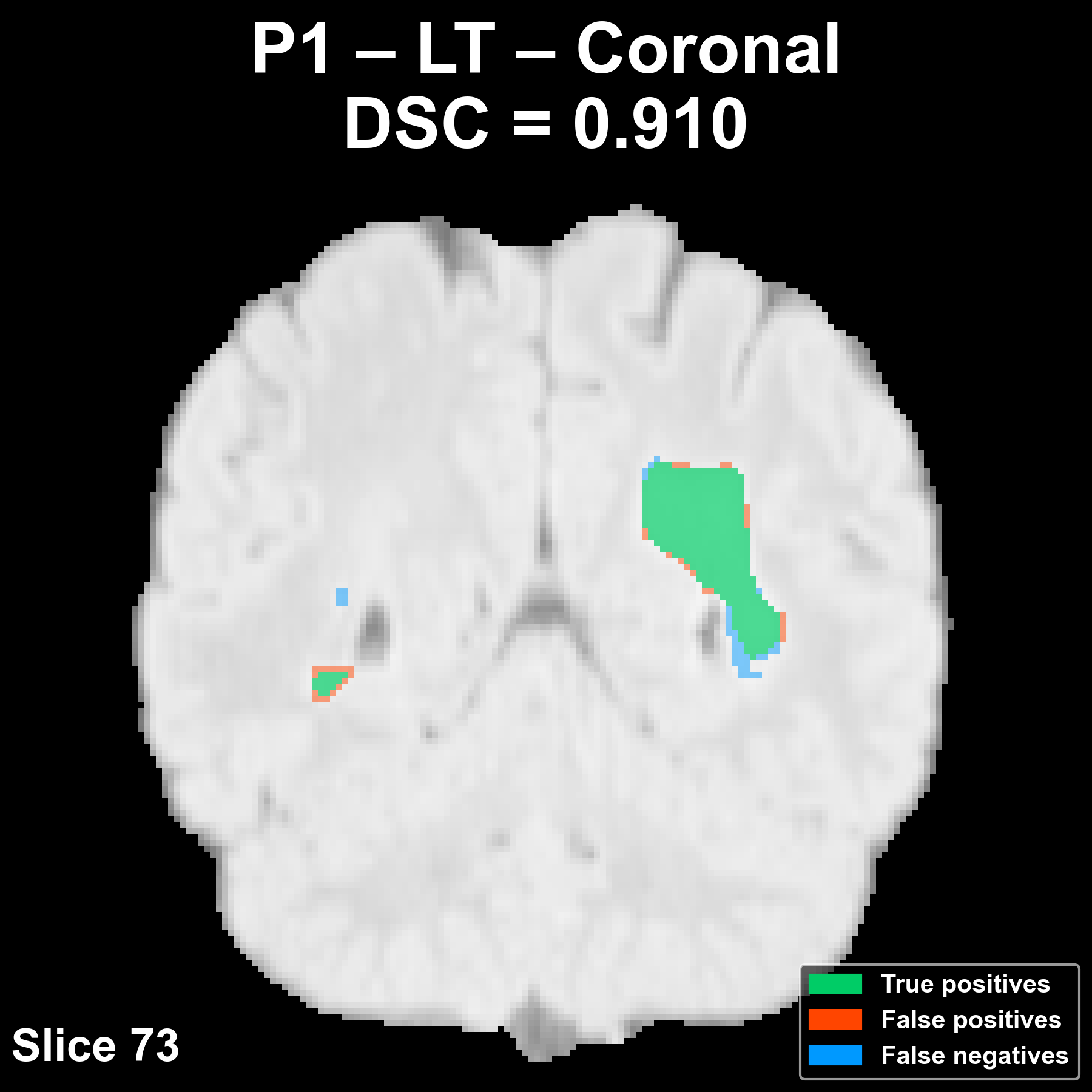
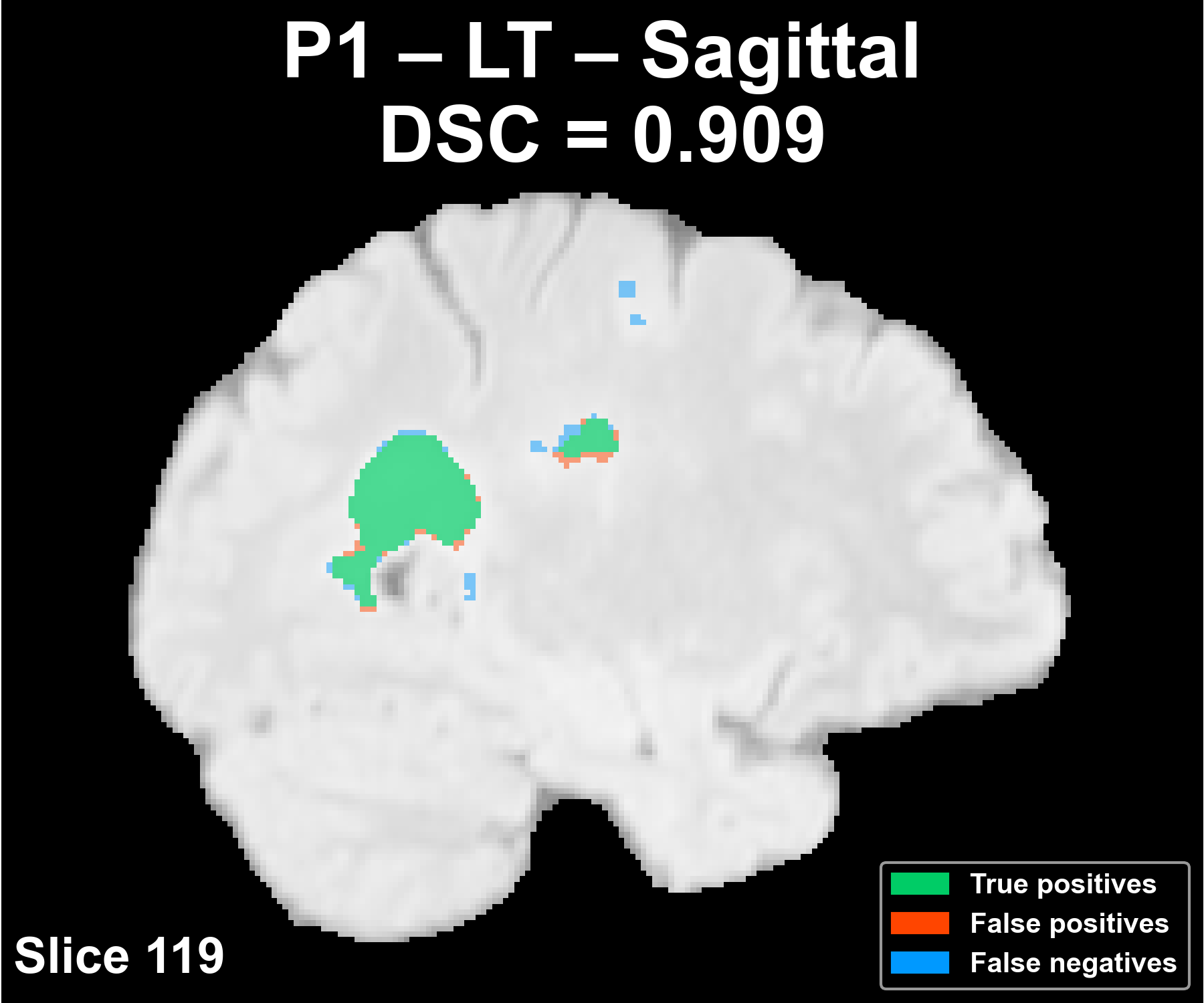
Complete patient sequence
The following animation shows the segmentation generated for another reference patient (P42) across all slices of the volume in the axial plane.
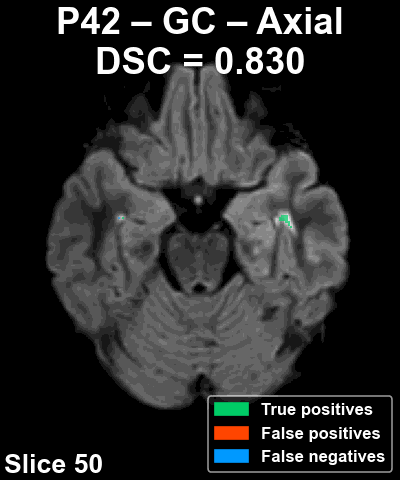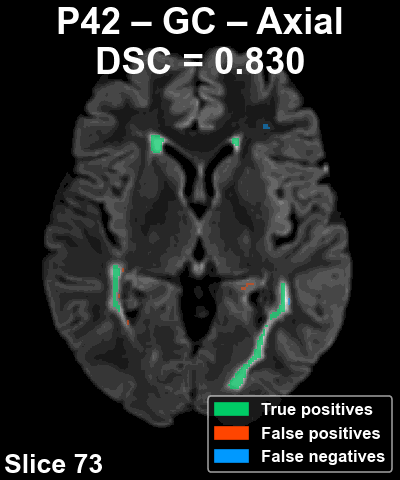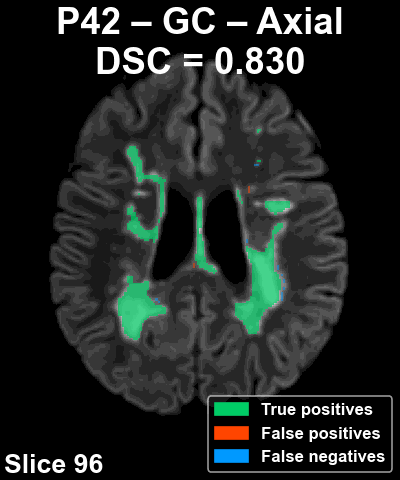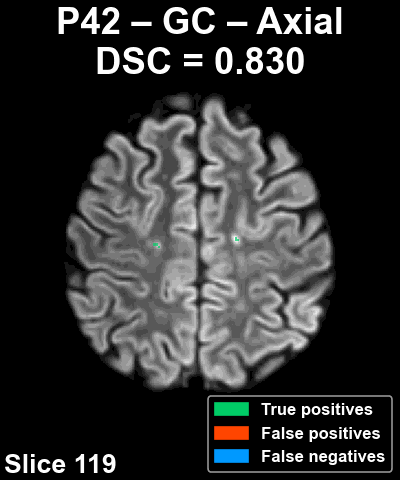
Pipeline Overview
The complete process consists of eight sequential stages,
automated via the run_pipeline.py script:
- Download and preparation of the official MSLesSeg dataset.
- Preprocessing and slice extraction in a format compatible with the YOLO model.
- Training of the YOLO11-seg model (optional).
- Generation of two-dimensional predictions.
- Reconstruction of three-dimensional volumes from predicted slices.
- Combination of volumes predicted across different planes (consensus).
- Quantitative evaluation using performance metrics.
- Computation of global experiment results.
Each module can be executed independently or through the global pipeline, ensuring flexibility for debugging or experimentation.
Installation
Dependencies
- Python 3.10+
- numpy, opencv-python, nibabel, matplotlib, ultralytics, requests, tqdm, pyyaml, scipy, scikit-learn, pandas, polars, psutil
Note: PyTorch is required but must be installed separately to allow GPU/CPU variant selection. Follow the official instructions.
User installation
pip install yolo-mslesseg
Repository Structure
The repository is organised as follows:
📦 yolo_mslesseg/
│
├── run_pipeline.py # Entry point for the full pipeline
├── 📁 configs/ # Per-stage configuration classes
├── 📁 scripts/ # Pipeline stage scripts
├── 📁 utils/ # Utilities and enhancement algorithms
└── 📁 extras/ # Additional utility scripts
Running the Pipeline
Once installed, run the full pipeline with:
yolo-mslesseg --plane axial --modality FLAIR --num_slices P50 --enhancement CLAHE --k_folds 5 --epochs 50 --full
Note:
yolo-mslessegis a CLI-first package. The recommended way to run experiments is through the command line as shown above. Programmatic use viaimportis supported for advanced users building custom workflows.
For the full list of arguments and stage-by-stage execution, see the GitHub repository.
References
- Ultralytics (2025). YOLO11 documentation.
- Guarnera, F., Rondinella, A., Crispino, E., et al. (2025). MSLesSeg: Baseline and benchmarking of a new Multiple Sclerosis lesion segmentation dataset. Scientific Data, 12, 920. https://doi.org/10.1038/s41597-025-05250-y.
License
This project is licensed under the MIT License.
Citation
If you use this work, please cite it once a reference is available. A citable reference will be added upon publication.
@article{rozenblum2026yolomslesseg,
author = {Jiménez-Partinen, Ariadna and Rozenblum, Sebastián and
Pascual-González, Mario and Ordóñez-Walkowiak, María Paulina and
Guirado-Osorio, Víctor and Molina-Cabello, Miguel A.},
title = {YOLO-MSLesSeg: Automated Multiple Sclerosis Lesion Segmentation
in MRI with Image Enhancement Techniques},
journal = {},
year = {2026},
doi = {}
}
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file yolo_mslesseg-0.1.3.tar.gz.
File metadata
- Download URL: yolo_mslesseg-0.1.3.tar.gz
- Upload date:
- Size: 85.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8ce1e814f6ba1a37e10b31b5a99ed5906d6c55855ce3981585f36ed6cddf4313
|
|
| MD5 |
76ee8708a0a97216cb251ea81223c1c8
|
|
| BLAKE2b-256 |
457de8649686999e6958b8203eafbe4a4f2895c495b28dcda06c39f8fdb9e509
|
File details
Details for the file yolo_mslesseg-0.1.3-py3-none-any.whl.
File metadata
- Download URL: yolo_mslesseg-0.1.3-py3-none-any.whl
- Upload date:
- Size: 117.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
893d0dd1f0cd36fac9f4fe1c28e9f6449034192ce1c5ba29c7dea3be304acd4a
|
|
| MD5 |
2186483eee1e036f266cb57b220a8b77
|
|
| BLAKE2b-256 |
27a6d69d95752d17327fbe0cbe8b626fa3eb59fe517de3ccdc77f7458985da1f
|












