ZnDraw - Interactive visualization for atomistic simulations
Project description
ZnDraw
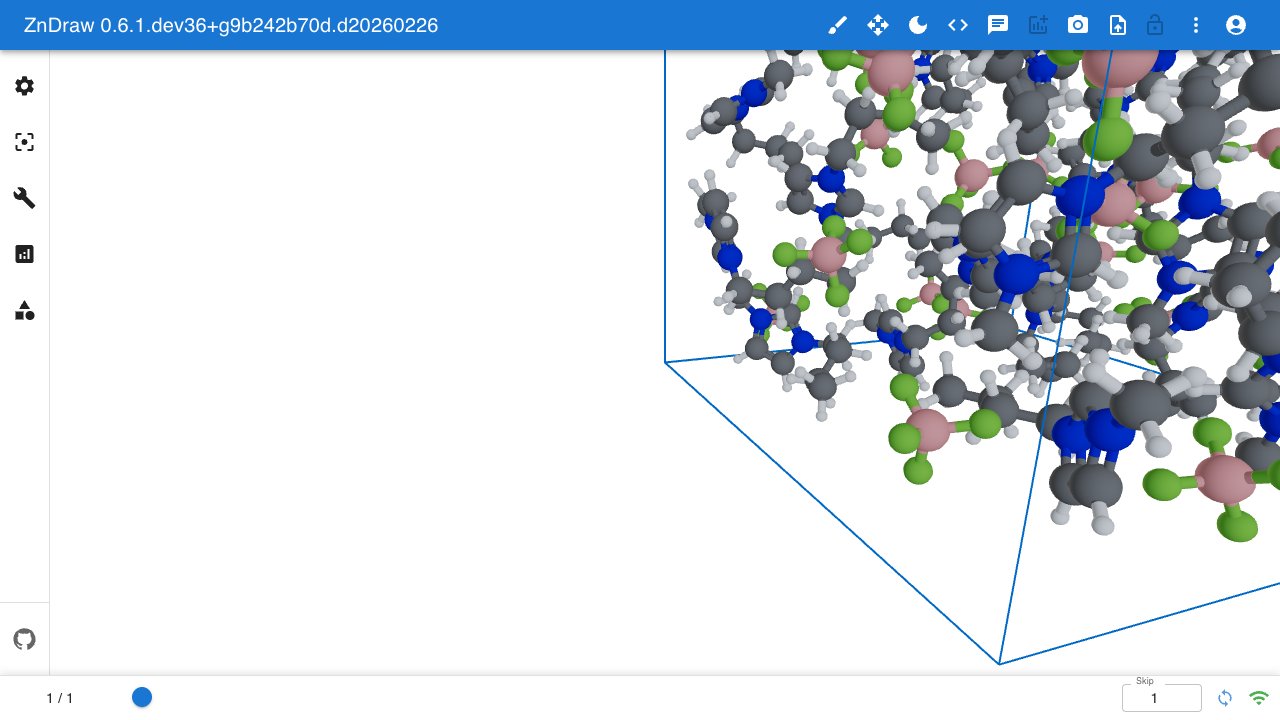
A Python-first visualization and editing tool for atomic structures with real-time collaboration.
Installation
You can install zndraw into your Python environment via pip:
pip install zndraw
or set it up as uv tool to run anywhere:
uvx zndraw <file>
Quick Start
Visualize your trajectories with a single command:
zndraw <file>
[!NOTE] ZnDraw's webapp-based approach allows you to use port forwarding to work with trajectories on remote systems.
Python API
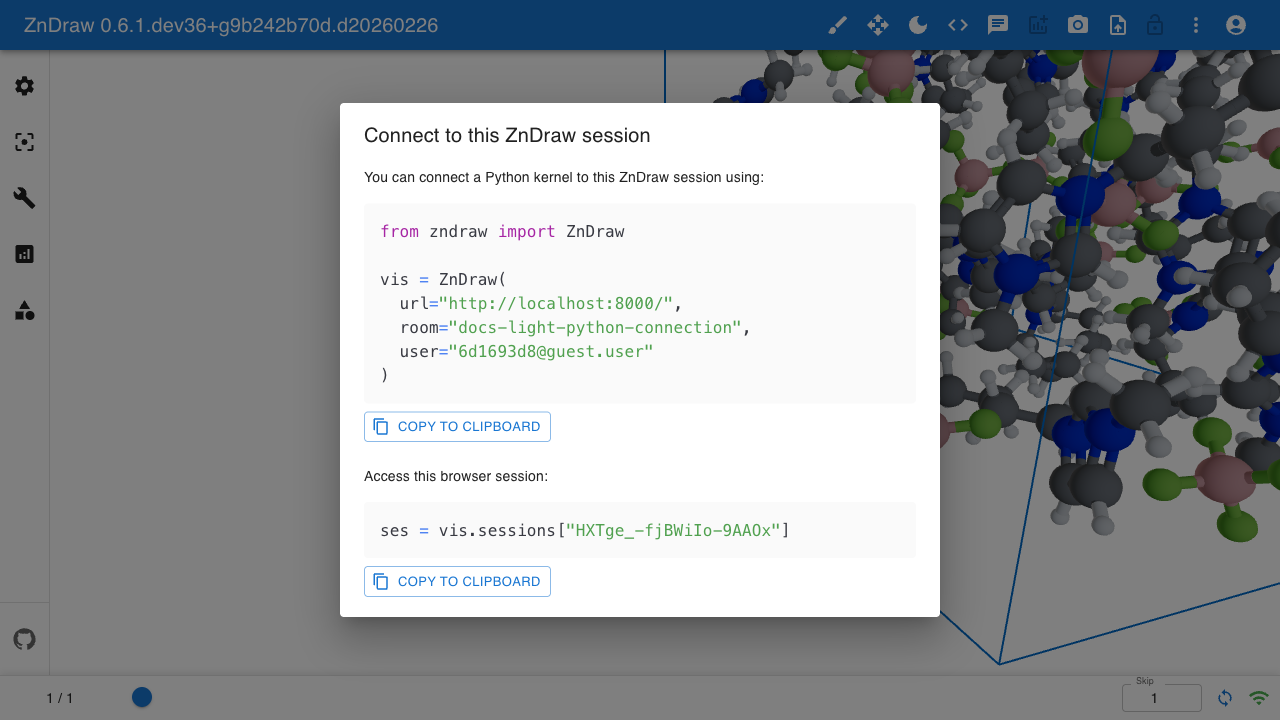
ZnDraw supports multiple clients connecting to the same visualization. Each visualization session is identified by a room name visible in the URL.
[!TIP] Server settings can be configured via environment variables (
ZNDRAW_SERVER_PORT,ZNDRAW_SERVER_HOST, etc.) or in yourpyproject.toml:[tool.zndraw.server] port = 8000 storage = "/data/frames.lmdb"
from zndraw import ZnDraw
vis = ZnDraw(url="http://localhost:1234", room="my-room")
Authentication
For protected deployments, provide credentials:
vis = ZnDraw(
url="http://localhost:1234",
room="my-room",
user="username",
password="password"
)
If no credentials are provided, the server assigns a guest user.
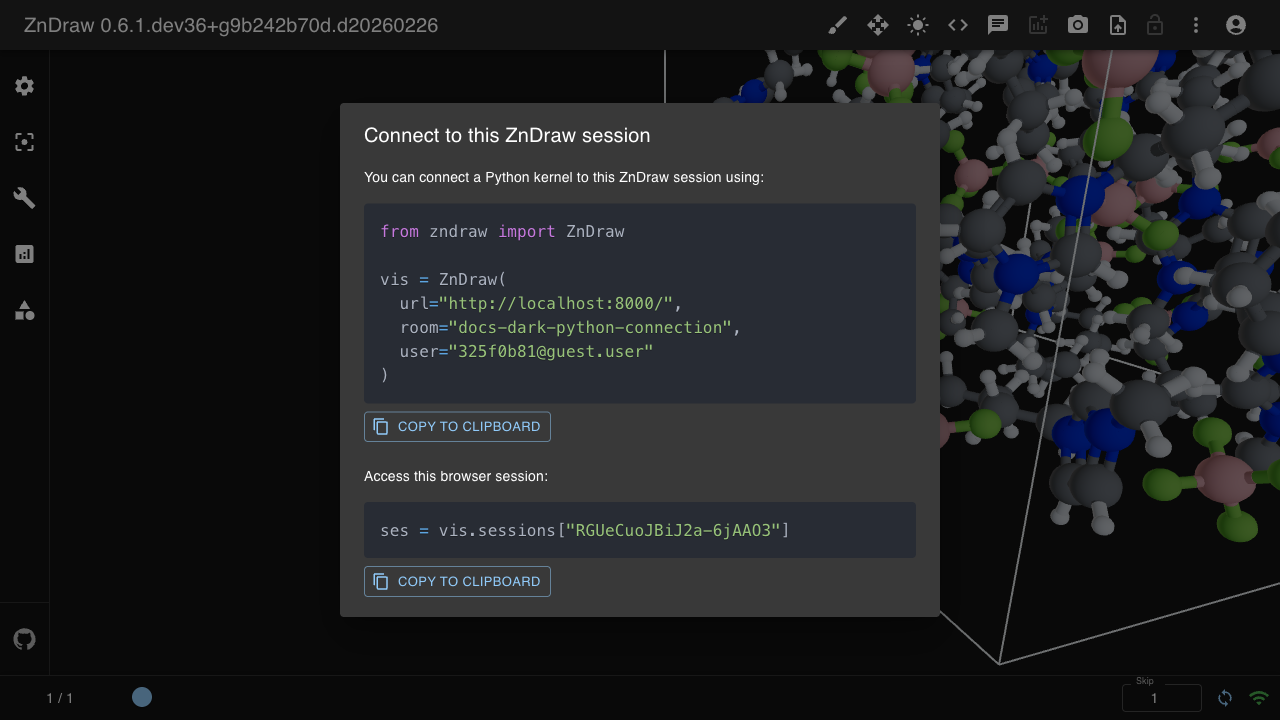
Working with Frames
The vis object behaves like a Python list of ase.Atoms objects. Changes are synchronized in real-time across all connected clients.
from ase.collections import s22
# Add structures
vis.extend(list(s22))
# Access current frame
atoms = vis[vis.step]
# Iterate over frames
for atoms in vis:
print(atoms)
# Slice operations
subset = vis[10:20]
Scene Properties
Control various aspects of the visualization:
vis.selection # Currently selected atoms
vis.step # Current frame index
vis.figures # Plotly figures
vis.bookmarks # Saved frame annotations
vis.geometries # 3D geometry overlays (dict-like)
vis.sessions # Session configuration
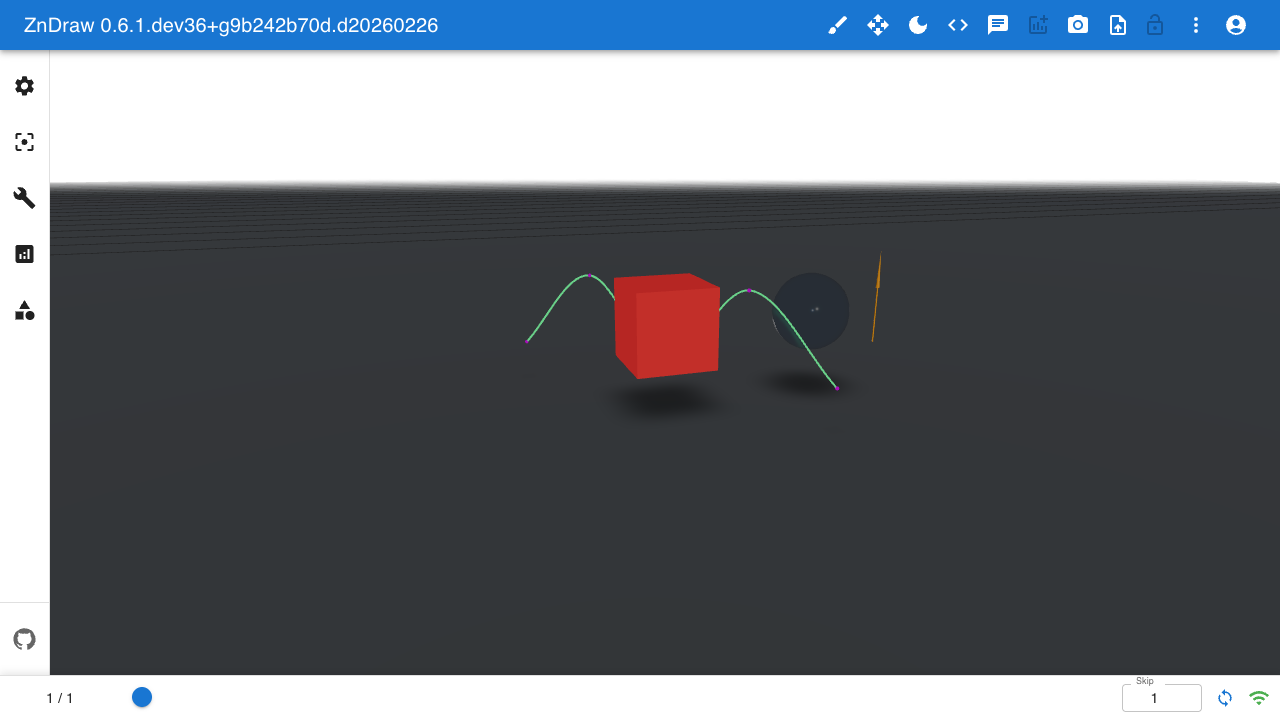
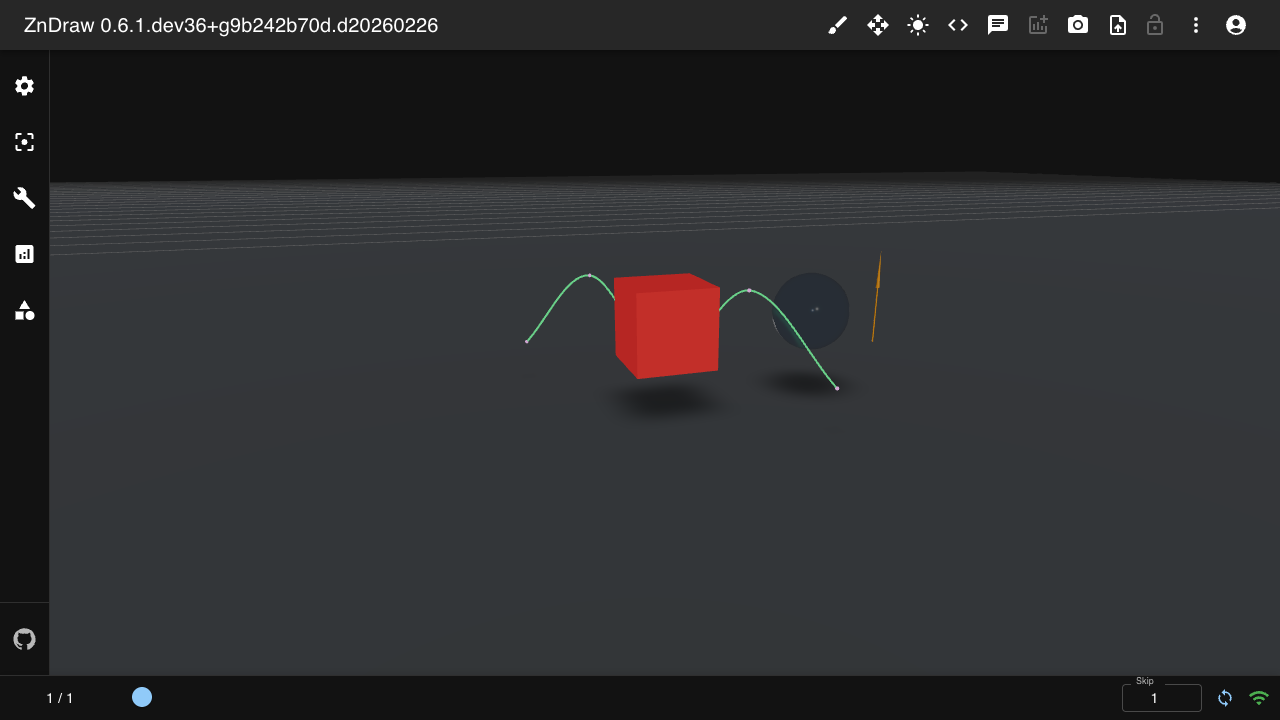
Geometries
Add 3D geometry overlays to your visualization:
from zndraw.geometries import Box, Sphere, Arrow, Camera, Curve
vis.geometries["box"] = Box(position=(0, 1, 2))
Available geometry types: Sphere, Arrow, Bond, Curve, Cell, Floor, Box, Plane, Shape, Camera, Isosurface.
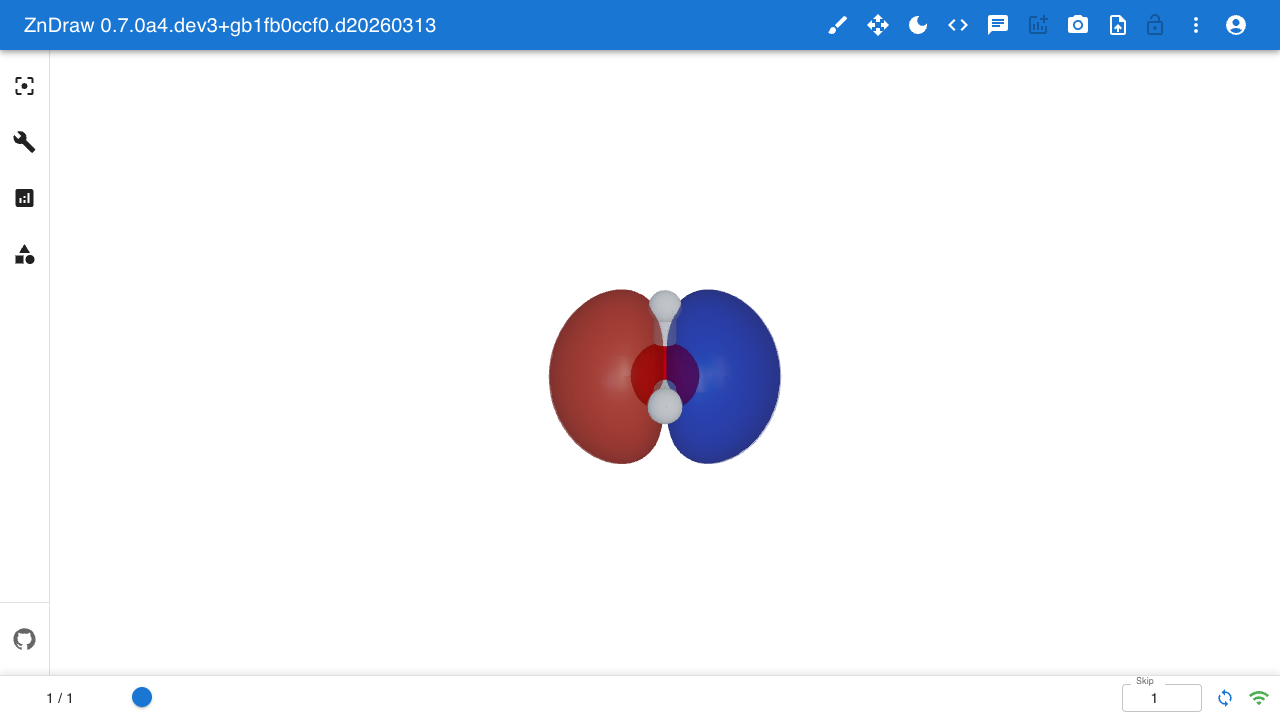
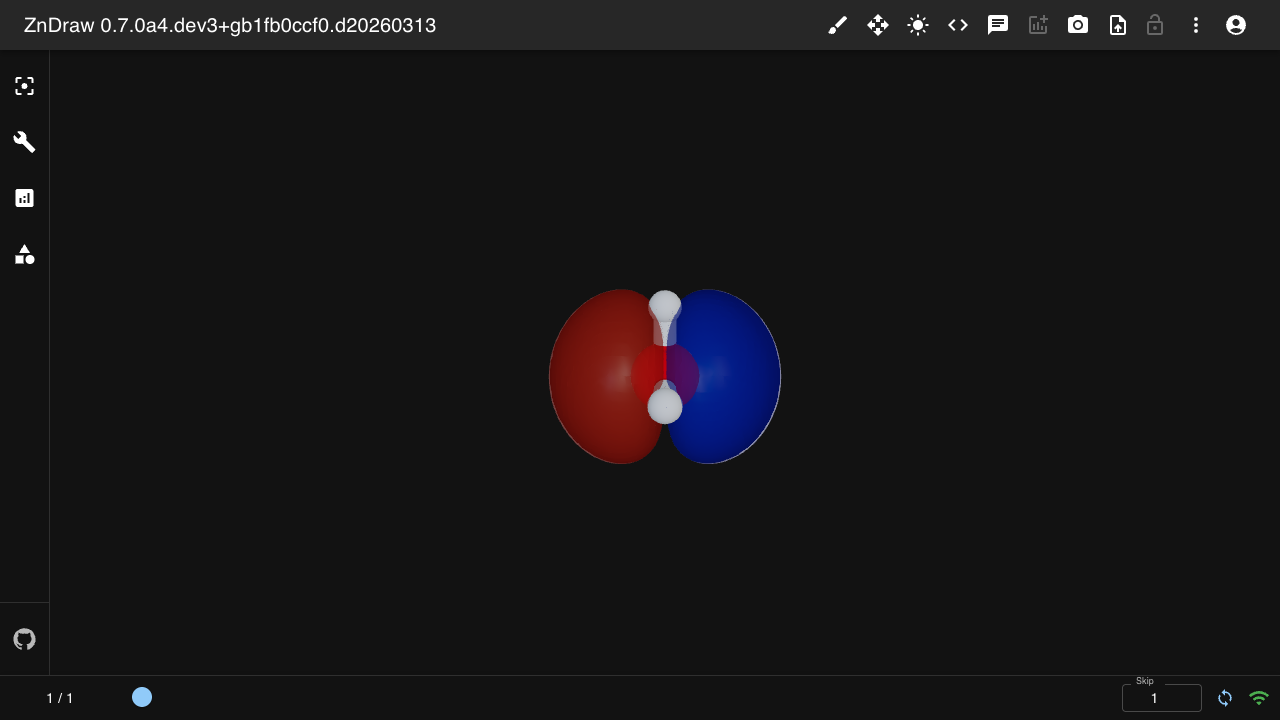
Isosurface
Visualize volumetric data (e.g. molecular orbitals) as 3D isosurfaces:
from zndraw.geometries import Isosurface
# atoms.info["orbital_homo"] must contain:
# {"grid": np.ndarray (Nx,Ny,Nz), "origin": np.ndarray (3,), "cell": np.ndarray (3,3)}
vis.geometries["homo"] = Isosurface(cube_key="info.orbital_homo", isovalue=0.02)
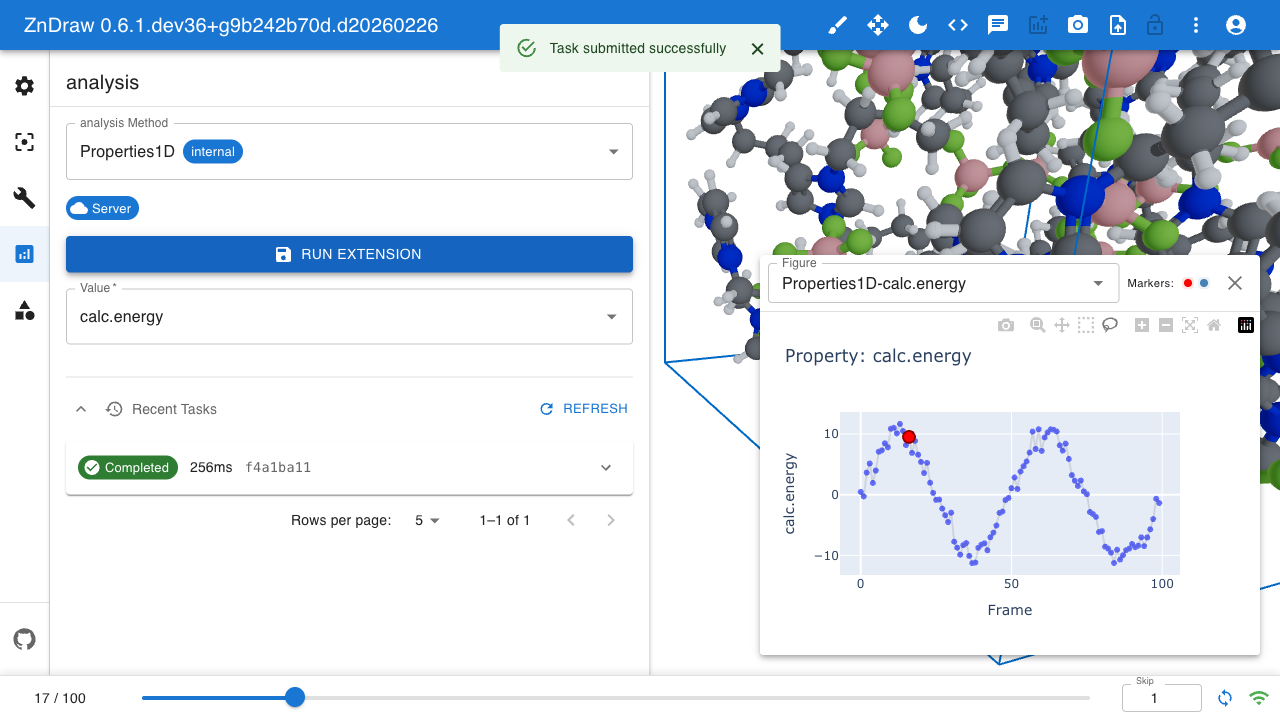
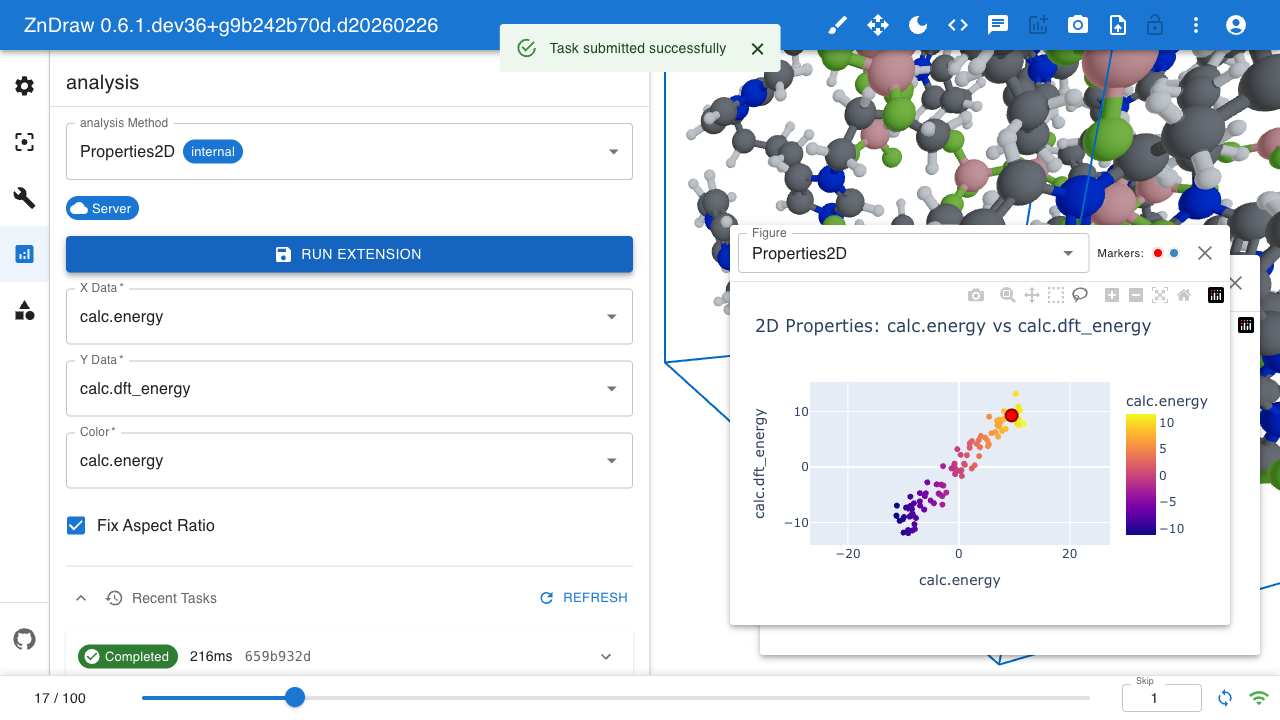
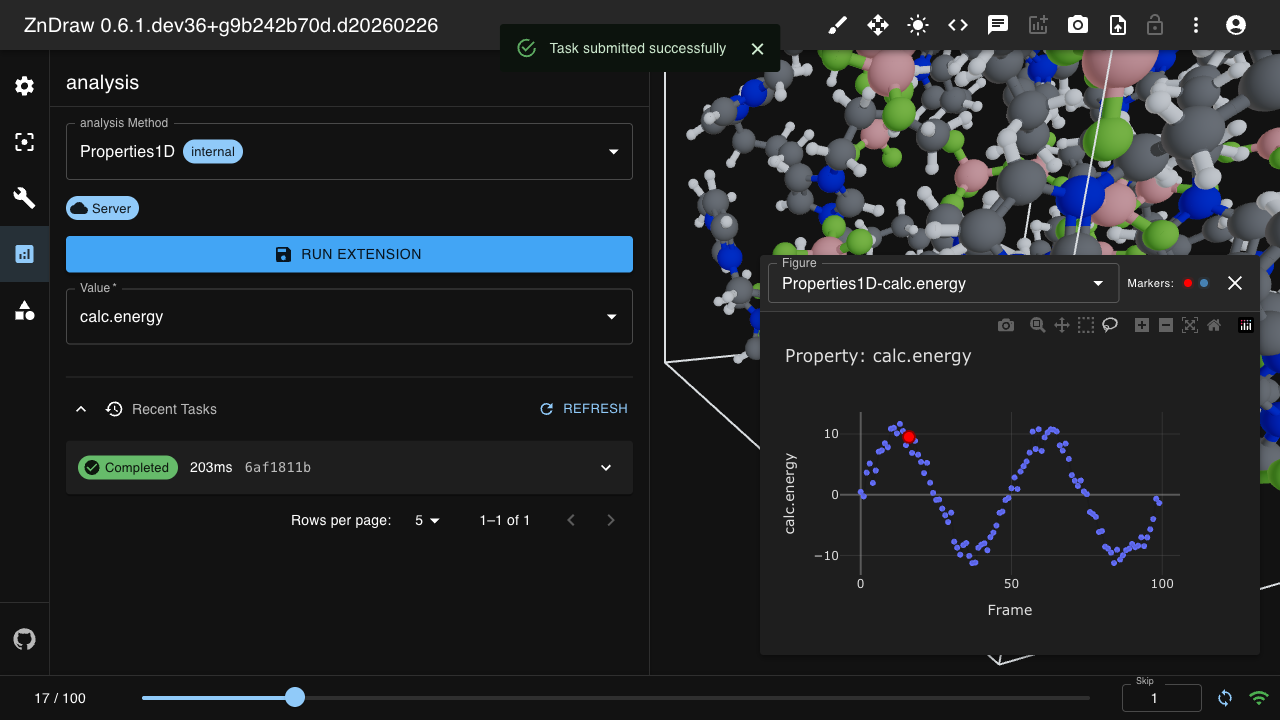
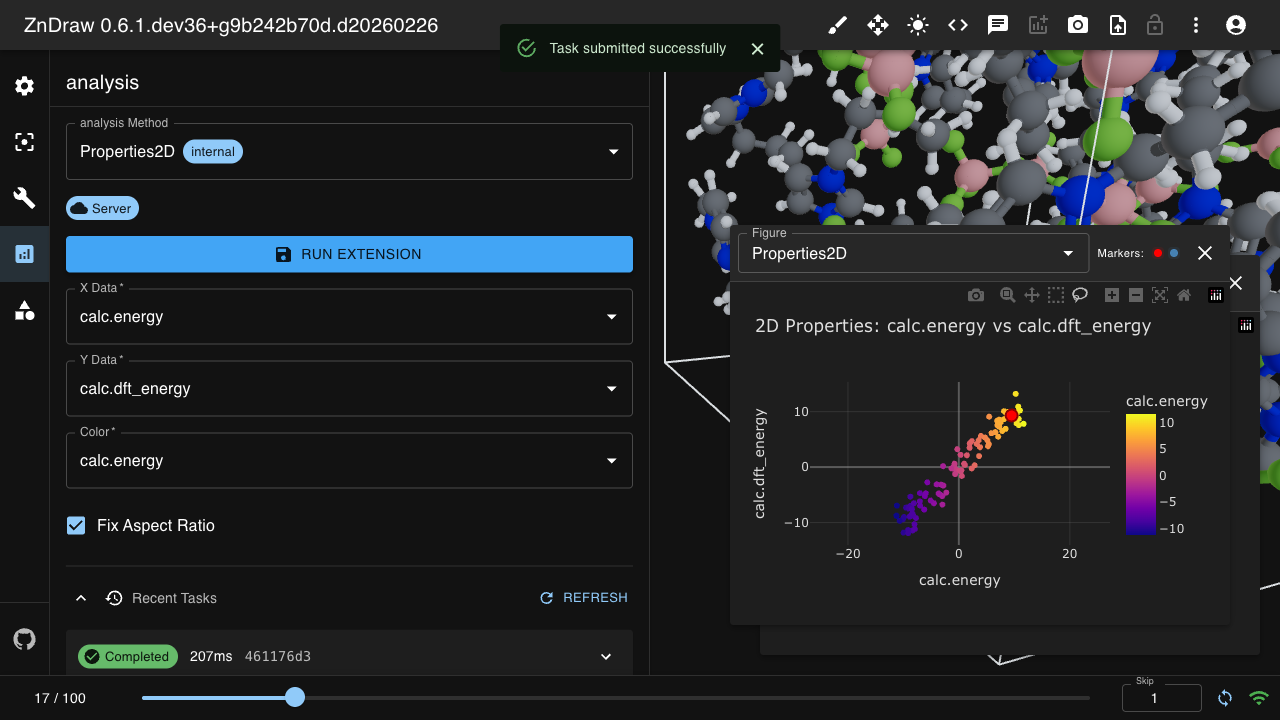
Analysis
ZnDraw integrates with Plotly for interactive data visualization. It automatically detects available properties and provides selection menus.
Extensions
Create custom tools accessible via the ZnDraw UI:
from molify import smiles2atoms
from zndraw import ZnDraw, Extension, Category
class AddMolecule(Extension):
category = Category.MODIFIER
smiles: str
def run(self, vis, **kwargs) -> None:
vis.append(smiles2atoms(self.smiles))
vis.step = len(vis) - 1
vis = ZnDraw()
vis.register_job(AddMolecule) # room-scoped (default)
vis.wait()
Extensions can be registered as room-scoped (default, visible only in the current room) or global (vis.register_job(cls, room="@global"), admin-only, visible in all rooms).
Pass heavy, pre-loaded objects (e.g. ML models) via run_kwargs — they stay in worker memory and are forwarded to run() on every task:
model = torch.load("model.pt")
vis.register_job(Predict, run_kwargs={"model": model})
Register filesystem providers so users can load files from the UI:
import fsspec
vis.register_fs(fsspec.filesystem("file"), name="local")
Hosted Version
A hosted version is available at https://zndraw.icp.uni-stuttgart.de
zndraw <file> --url https://zndraw.icp.uni-stuttgart.de
Self-Hosting
ZnDraw can be deployed using Docker:
See the docker/ directory for complete deployment configurations:
- Standalone - Simple single-instance deployment for personal use or small teams
- Production - Horizontal scaling with nginx load balancer for high load
References
If you use ZnDraw in your research, please cite:
@article{elijosius2025zero,
title = {Zero-shot molecular generation via similarity kernels},
author = {Elijo{\v s}ius, Rokas and Zills, Fabian and Batatia, Ilyes and Norwood, Sam Walton and Kov{\'a}cs, D{\'a}vid P{\'e}ter and Holm, Christian and Cs{\'a}nyi, G{\'a}bor},
journal = {Nature Communications},
volume = {16},
pages = {5479},
year = {2025},
doi = {10.1038/s41467-025-60963-3},
url = {https://doi.org/10.1038/s41467-025-60963-3},
}
Acknowledgements
The creation of ZnDraw was supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) in the framework of the priority program SPP 2363, "Utilization and Development of Machine Learning for Molecular Applications - Molecular Machine Learning" Project No. 497249646. Further funding through the DFG under Germany's Excellence Strategy - EXC 2075 - 390740016 and the Stuttgart Center for Simulation Science (SimTech) was provided.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file zndraw-0.7.0a8.tar.gz.
File metadata
- Download URL: zndraw-0.7.0a8.tar.gz
- Upload date:
- Size: 10.9 MB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
aedca30f5a630cbedabac9f48e2f674230c362587d26a2b999c848fffbcb90ea
|
|
| MD5 |
85462d51bba9c4bfe937745d9fa0c919
|
|
| BLAKE2b-256 |
a9081fc8988d3c3c4b11f5931c70a2c39434db532f638dc87cb5c76a24dff8c4
|
Provenance
The following attestation bundles were made for zndraw-0.7.0a8.tar.gz:
Publisher:
publish.yaml on zincware/ZnDraw
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
zndraw-0.7.0a8.tar.gz -
Subject digest:
aedca30f5a630cbedabac9f48e2f674230c362587d26a2b999c848fffbcb90ea - Sigstore transparency entry: 1205491740
- Sigstore integration time:
-
Permalink:
zincware/ZnDraw@42b7c36d0deba8bd6799f6f6696022aef72b02d4 -
Branch / Tag:
refs/tags/v0.7.0a8 - Owner: https://github.com/zincware
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yaml@42b7c36d0deba8bd6799f6f6696022aef72b02d4 -
Trigger Event:
release
-
Statement type:
File details
Details for the file zndraw-0.7.0a8-py3-none-any.whl.
File metadata
- Download URL: zndraw-0.7.0a8-py3-none-any.whl
- Upload date:
- Size: 11.4 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
22df714648f78ef3e618c59bd18cc0950ac5a229d84c1b47552a38d42a22ddbf
|
|
| MD5 |
4d0e35c330a3680e95f7730223dbc247
|
|
| BLAKE2b-256 |
51862961f5afe1676b518c1838122f1239bc6c1b2c0861202a34386ccc7b8533
|
Provenance
The following attestation bundles were made for zndraw-0.7.0a8-py3-none-any.whl:
Publisher:
publish.yaml on zincware/ZnDraw
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
zndraw-0.7.0a8-py3-none-any.whl -
Subject digest:
22df714648f78ef3e618c59bd18cc0950ac5a229d84c1b47552a38d42a22ddbf - Sigstore transparency entry: 1205491762
- Sigstore integration time:
-
Permalink:
zincware/ZnDraw@42b7c36d0deba8bd6799f6f6696022aef72b02d4 -
Branch / Tag:
refs/tags/v0.7.0a8 - Owner: https://github.com/zincware
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yaml@42b7c36d0deba8bd6799f6f6696022aef72b02d4 -
Trigger Event:
release
-
Statement type: