Fast Linear Algebra for Scalable Hybrid Deconvolution of Spatial Transcriptomics
Project description
FlashDeconv
Spatial deconvolution with linear scalability for atlas-scale data.
FlashDeconv estimates cell type proportions from spatial transcriptomics data (Visium, Visium HD, Stereo-seq). It is designed for large-scale analyses where computational efficiency is essential, while maintaining attention to low-abundance cell populations through leverage-score-based feature weighting.
Paper: Yang, C., Zhang, X. & Chen, J. FlashDeconv enables atlas-scale, multi-resolution spatial deconvolution via structure-preserving sketching. bioRxiv (2025). DOI: 10.64898/2025.12.22.696108
Installation
pip install flashdeconv
For development or additional I/O support, see Installation Options.
Quick Start
import scanpy as sc
import flashdeconv as fd
# Load data
adata_st = sc.read_h5ad("spatial.h5ad")
adata_ref = sc.read_h5ad("reference.h5ad")
# Deconvolve
fd.tl.deconvolve(adata_st, adata_ref, cell_type_key="cell_type")
# Results stored in adata_st.obsm["flashdeconv"]
sc.pl.spatial(adata_st, color="flashdeconv_Hepatocyte")
Overview
Spatial deconvolution methods offer different trade-offs. Probabilistic approaches like Cell2Location and RCTD provide rigorous uncertainty quantification; methods like CARD incorporate spatial structure through dense kernel matrices. FlashDeconv takes a complementary approach, prioritizing computational efficiency for million-scale datasets.
Design Principles
-
Linear complexity — O(N) time and memory through randomized sketching and sparse graph regularization.
-
Leverage-based feature weighting — Variance-based selection (PCA, HVG) can underweight markers of low-abundance populations. We use leverage scores from the reference SVD to identify genes that define distinct transcriptomic directions, regardless of expression magnitude.
-
Sparse spatial regularization — Graph Laplacian smoothing with O(N) complexity, avoiding the O(N²) cost of dense kernel methods.
Performance
Scalability
| Spots | Time | Memory |
|---|---|---|
| 10,000 | < 1 sec | < 1 GB |
| 100,000 | ~4 sec | ~2 GB |
| 1,000,000 | ~3 min | ~21 GB |
Benchmarked on MacBook Pro M2 Max (32GB unified memory), CPU-only.
Accuracy
On the Spotless benchmark:
| Metric | FlashDeconv | RCTD | Cell2Location |
|---|---|---|---|
| Pearson (56 datasets) | 0.944 | 0.905 | 0.895 |
Performance varies by tissue type and experimental conditions. We recommend evaluating on data similar to your use case.
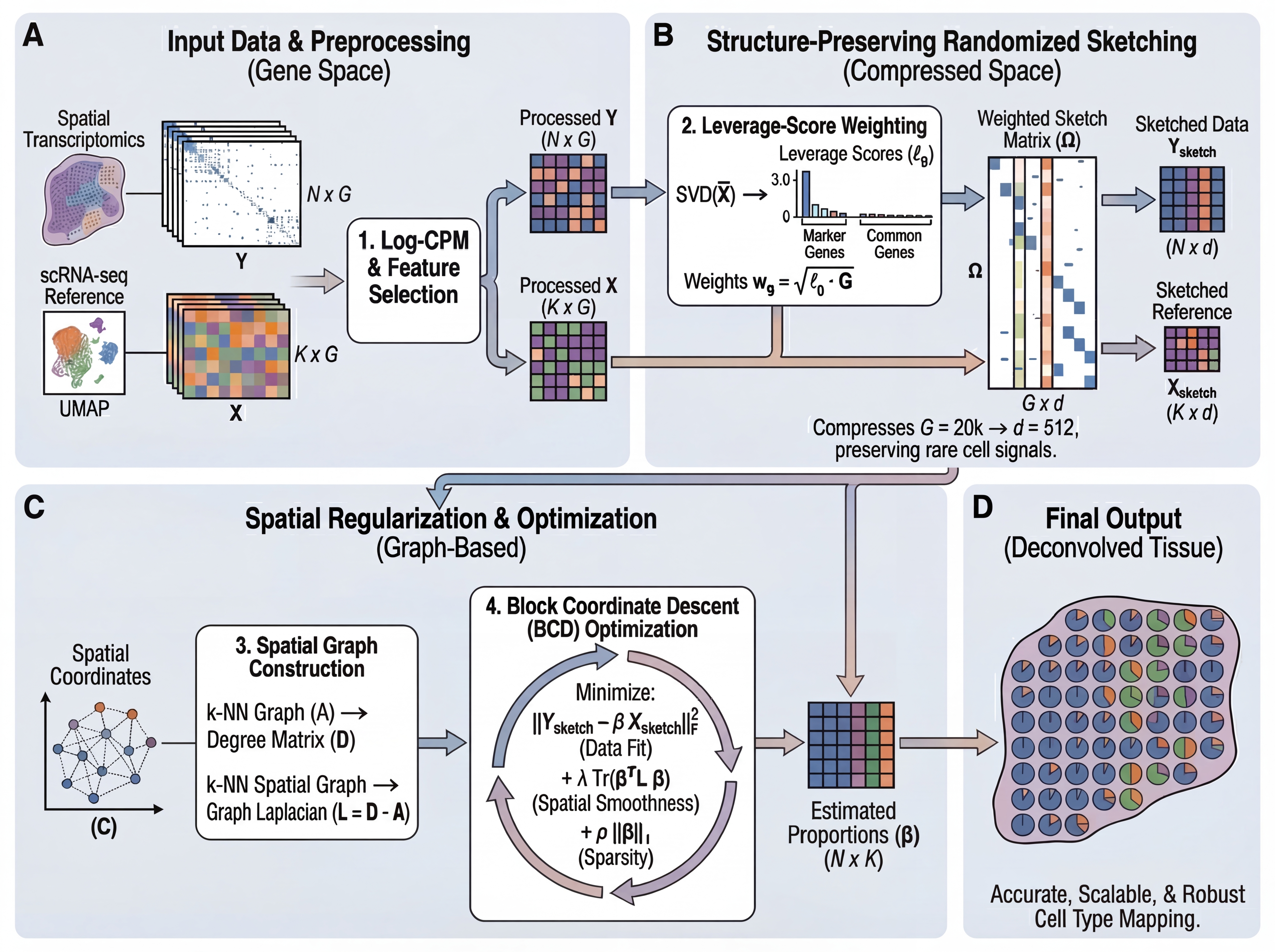
Algorithm
FlashDeconv solves a graph-regularized non-negative least squares problem:
minimize ½‖Y - βX‖²_F + λ·Tr(βᵀLβ) + ρ‖β‖₁, subject to β ≥ 0
where Y is spatial expression, X is reference signatures, L is the graph Laplacian, and β represents cell type abundances.
Pipeline:
- Select informative genes (HVG ∪ markers) and compute leverage scores
- Compress gene space via weighted CountSketch (G → 512 dimensions)
- Construct sparse k-NN spatial graph
- Solve via block coordinate descent with spatial smoothing
API
Scanpy-style
fd.tl.deconvolve(
adata_st, # Spatial AnnData
adata_ref, # Reference AnnData
cell_type_key="cell_type", # Column in adata_ref.obs
key_added="flashdeconv", # Key for results
)
NumPy
from flashdeconv import FlashDeconv
model = FlashDeconv(
sketch_dim=512,
lambda_spatial="auto",
n_hvg=2000,
k_neighbors=6,
random_state=0,
)
proportions = model.fit_transform(Y, X, coords)
Parameters
| Parameter | Default | Description |
|---|---|---|
sketch_dim |
512 | Sketch dimension |
lambda_spatial |
"auto" | Spatial regularization (auto-tuned) |
n_hvg |
2000 | Highly variable genes |
k_neighbors |
6 | Spatial graph neighbors |
preprocess |
"log_cpm" | Normalization: "log_cpm", "pearson", or "raw" |
random_state |
0 | Random seed for reproducibility |
Output
| Attribute | Description |
|---|---|
proportions_ |
Cell type proportions (N × K), sum to 1 |
beta_ |
Raw abundances (N × K) |
info_ |
Convergence statistics |
Input Formats
- Spatial data: AnnData, NumPy array (N × G), or SciPy sparse matrix
- Reference: AnnData (aggregated by cell type) or NumPy array (K × G)
- Coordinates: Extracted from
adata.obsm["spatial"]or NumPy array (N × 2)
Reference Quality
Deconvolution accuracy depends on reference quality:
| Requirement | Guideline |
|---|---|
| Cells per type | ≥ 500 recommended |
| Marker fold-change | ≥ 5× for distinguishability |
| Signature correlation | < 0.95 between types |
| No Unknown cells | Filter before deconvolution |
Critical: Always remove cells labeled "Unknown", "Unassigned", or similar. These cells act as universal signatures that absorb proportions from specific types—a fundamental property of regression-based deconvolution, not a FlashDeconv limitation.
See Reference Data Guide for details.
Installation Options
# Standard
pip install flashdeconv
# With AnnData support
pip install flashdeconv[io]
# Development
git clone https://github.com/cafferychen777/flashdeconv.git
cd flashdeconv && pip install -e ".[dev]"
Requirements: Python ≥ 3.9, numpy, scipy, numba. Optional: scanpy, anndata.
Citation
If you use FlashDeconv in your research, please cite:
Yang, C., Zhang, X. & Chen, J. FlashDeconv enables atlas-scale, multi-resolution spatial deconvolution via structure-preserving sketching. bioRxiv (2025). DOI: 10.64898/2025.12.22.696108
@article{yang2025flashdeconv,
title={FlashDeconv enables atlas-scale, multi-resolution spatial deconvolution
via structure-preserving sketching},
author={Yang, Chen and Zhang, Xianyang and Chen, Jun},
journal={bioRxiv},
year={2025},
doi={10.64898/2025.12.22.696108}
}
Resources
- Paper reproducibility code
- Reference data guide — Building quality reference signatures
- Stereo-seq guide — Platform-specific considerations
- GitHub Issues
- BSD-3-Clause License
Acknowledgments
We thank the developers of Spotless, Cell2Location, RCTD, CARD, and other deconvolution methods whose work contributed to this field.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file flashdeconv-0.1.6.tar.gz.
File metadata
- Download URL: flashdeconv-0.1.6.tar.gz
- Upload date:
- Size: 38.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9d36c0b236ff76bb62a196505142865d374670b480da7efb4418a2361828538e
|
|
| MD5 |
7e1c8a6fde421b2a14b85778274057b1
|
|
| BLAKE2b-256 |
3f79cba63c1f636234868d061a7791c65aca3dbdb74508e6ab7adc29fc157149
|
File details
Details for the file flashdeconv-0.1.6-py3-none-any.whl.
File metadata
- Download URL: flashdeconv-0.1.6-py3-none-any.whl
- Upload date:
- Size: 35.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.14.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
152294b84a1334960a28859d60639f9b271df34800ebbe00ec37aeccc089b791
|
|
| MD5 |
6e16694b3ed8f8f109dbe169e0a2a3b1
|
|
| BLAKE2b-256 |
f2a6c7e5ac938c0dacbb2b46c245f6528292ec052aeb95404682c216fda540c2
|