Fast Linear Algebra for Scalable Hybrid Deconvolution of Spatial Transcriptomics
Project description
FlashDeconv
Fast Linear Algebra for Scalable Hybrid Deconvolution
Unlocking atlas-scale spatial biology with randomized numerical linear algebra.
FlashDeconv is a high-performance spatial transcriptomics deconvolution method designed for atlas-scale and subcellular-resolution platforms (Visium HD, Stereo-seq, Xenium). It leverages structure-preserving randomized sketching to estimate cell type proportions with linear scalability—processing 1 million spots in ~3 minutes on commodity hardware.
Paper: Chen Yang, Jun Chen, Xianyang Zhang. FlashDeconv enables atlas-scale, multi-resolution spatial deconvolution via structure-preserving sketching. bioRxiv, 2025. DOI: 10.64898/2025.12.22.696108
Reproducibility: To reproduce figures and benchmarks from the paper, visit the flashdeconv-reproducibility repository.
Key Features
- Ultra-fast & Scalable: Deconvolve 1 million spots in ~3 minutes. Time and memory scale linearly O(N) with dataset size.
- Hardware Friendly: No GPU required. Runs efficiently on laptops (e.g., 32GB RAM handles 1M spots).
- Rare Cell Detection: Uses leverage-score sampling to preserve transcriptomically distinct but low-abundance cell types (e.g., Tuft cells, endothelial cells) that variance-based methods systematically miss.
- Spatially Aware: Sparse graph Laplacian regularization ensures spatial coherence without the O(N²) cost of dense kernel methods.
- Visium HD Ready: Specifically optimized for the extreme sparsity and scale of subcellular resolution technologies (2µm–16µm bin sizes).
- Statistically Rigorous: Log-CPM normalization with leverage-weighted gene selection preserves both common and rare cell populations.
Installation
# From PyPI (recommended)
pip install flashdeconv
# With scanpy/anndata integration
pip install flashdeconv[io]
For development:
# From source
git clone https://github.com/cafferychen777/flashdeconv.git
cd flashdeconv
pip install -e ".[dev]"
Requirements: Python ≥ 3.9, numpy, scipy, numba. Optional: scanpy, anndata for AnnData workflow.
Quick Start
With Scanpy/AnnData
import scanpy as sc
import flashdeconv as fd
adata_st = sc.read_h5ad("visium_hd.h5ad")
adata_ref = sc.read_h5ad("reference.h5ad")
fd.tl.deconvolve(adata_st, adata_ref, cell_type_key="cell_type")
adata_st.obsm["flashdeconv"] # Cell type proportions
sc.pl.spatial(adata_st, color="flashdeconv_Hepatocyte")
With NumPy
from flashdeconv import FlashDeconv
model = FlashDeconv()
proportions = model.fit_transform(Y, X, coords) # (n_spots, n_cell_types)
Algorithm Under the Hood
FlashDeconv reformulates spatial deconvolution as Graph-Regularized Non-Negative Least Squares, solved in a compressed "sketch" space via randomized numerical linear algebra (RandNLA):
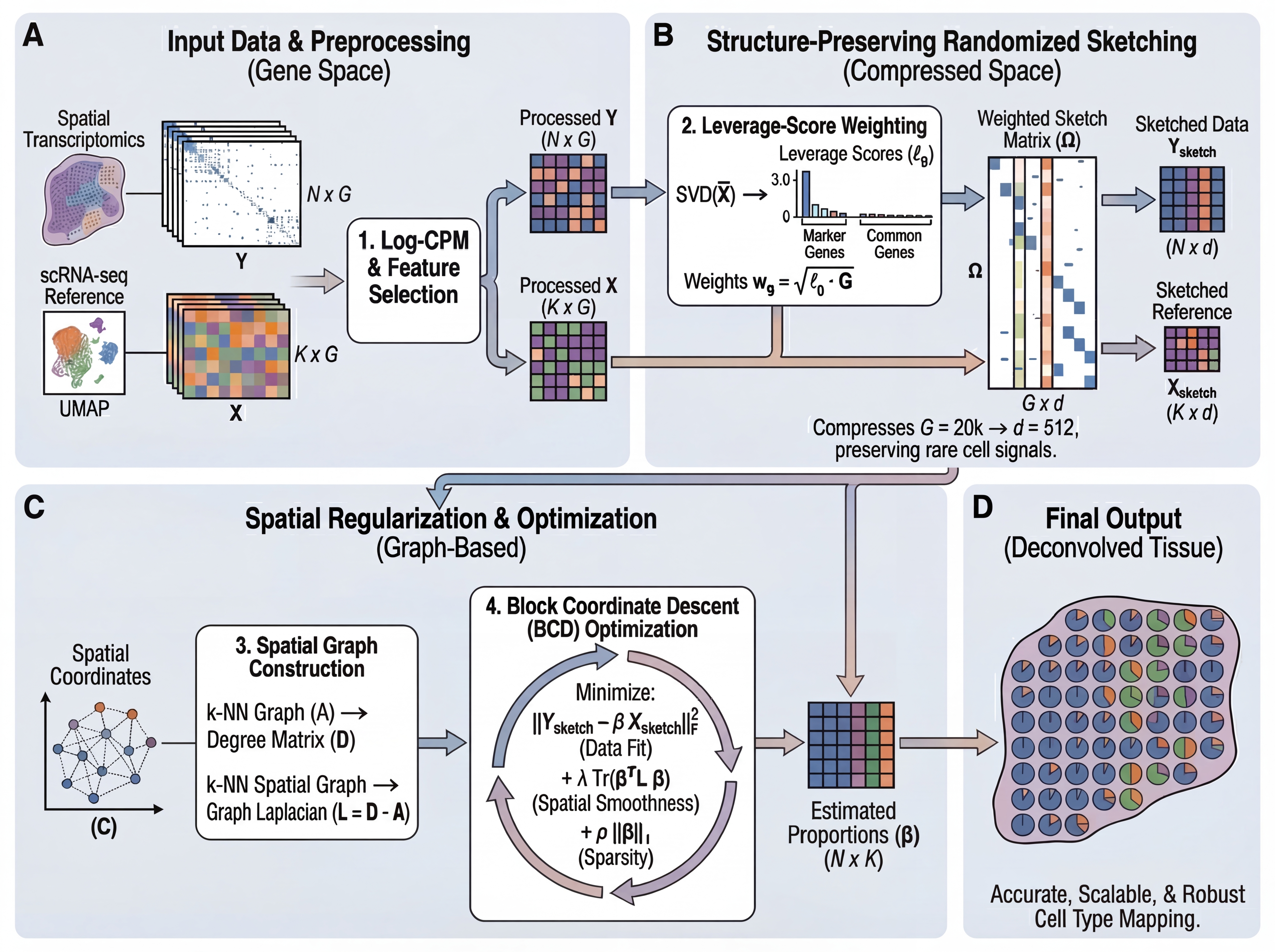
Three-Stage Framework
-
Preprocessing & Gene Selection
- Log-CPM normalization: Stabilizes variance and prevents high-expression genes from dominating the sketch
- Leverage-weighted gene selection: Combines highly variable genes (HVGs) with cell-type-specific markers, weighted by statistical leverage scores. Unlike variance (which conflates abundance with informativeness), leverage scores identify genes that define transcriptomically distinct directions, preserving rare cell type markers.
-
Structure-Preserving Sketching
- Randomized projection: Compress gene space (~20,000 genes → 512 dimensions) using CountSketch with leverage-score importance sampling
- Johnson-Lindenstrauss guarantee: Preserves Euclidean distances between cell type signatures with high probability
- Key innovation: Leverage-weighted sampling amplifies rare cell type markers relative to housekeeping genes, preventing signal loss during hash collisions
-
Spatial Graph Regularization
- Sparse graph Laplacian: Constructs k-NN spatial graph (O(N) memory vs. O(N²) for dense kernels like CARD)
- Numba-accelerated Block Coordinate Descent (BCD): Fast closed-form updates with non-negativity constraints
- Linear scalability: Spatial term complexity O(N·k) enables million-spot analysis
Why This Works
- Log-CPM bounds dynamic range while preserving sparsity (log1p(0) = 0)
- Leverage scores decouple biological identity from population abundance—markers of rare cell types (0.1% frequency) receive equal weight to abundant types (30% frequency)
- Sparse graph Laplacian encodes spatial autocorrelation as a Gaussian Markov Random Field (GMRF) without dense matrix operations
Benchmarks
FlashDeconv exhibits linear O(N) scaling for both time and memory:
| Dataset Size | Runtime | Memory | Hardware |
|---|---|---|---|
| 10K spots | < 1 sec | < 1 GB | MacBook Pro M2 Max |
| 100K spots | ~4 sec | ~2 GB | (32GB unified memory) |
| 1M spots | ~3 min | ~21 GB | No GPU required |
Accuracy on Synthetic Benchmarks (Spotless suite):
- Pearson correlation: 0.944 (mean across 56 datasets spanning 6 tissues)
- RMSE: 0.065 (median)
- Rare cell detection (AUPR): 0.960 ± 0.036 (standard deviation)
Real-world validation:
- Mouse liver (Visium): JSD = 0.056, ranking 3rd among 13 methods
- Melanoma tumor (Visium): JSD = 0.027, ranking 5th among 13 methods
- Reference stability: Ranked 1st for robustness to different scRNA-seq protocols
FlashDeconv matches top-tier Bayesian methods (Cell2Location, RCTD) on accuracy while accelerating inference by orders of magnitude.
API Reference
fd.tl.deconvolve
fd.tl.deconvolve(
adata_st, # Spatial AnnData
adata_ref, # Reference AnnData
cell_type_key="cell_type", # Column in adata_ref.obs
key_added="flashdeconv", # Key for results in adata_st
random_state=0, # Random seed for reproducibility
copy=False, # If True, return copy instead of inplace
)
Results stored in adata_st:
.obsm["flashdeconv"]— Cell type proportions (DataFrame).obs["flashdeconv_dominant"]— Dominant cell type per spot.uns["flashdeconv_params"]— Parameters used
FlashDeconv Class
class FlashDeconv:
def __init__(
self,
sketch_dim=512, # Sketch space dimension
lambda_spatial="auto", # Spatial regularization (auto-tuned)
rho_sparsity=0.01, # L1 sparsity penalty
n_hvg=2000, # Number of highly variable genes
n_markers_per_type=50, # Marker genes per cell type
spatial_method="knn", # "knn", "radius", or "grid"
k_neighbors=6, # k for k-NN graph
max_iter=100, # BCD max iterations
tol=1e-4, # Convergence tolerance
preprocess="log_cpm", # "log_cpm", "pearson", or "raw"
random_state=0, # Random seed for reproducibility
verbose=False,
): ...
def fit(self, Y, X, coords, gene_names=None, cell_type_names=None) -> self
def fit_transform(self, Y, X, coords, **kwargs) -> np.ndarray
def get_cell_type_proportions(self) -> np.ndarray
def get_abundances(self) -> np.ndarray
def get_dominant_cell_type(self) -> np.ndarray
def summary(self) -> dict
Parameters
| Parameter | Type | Default | Description |
|---|---|---|---|
sketch_dim |
int | 512 | Dimension of sketch space (higher = more info, slower) |
lambda_spatial |
float or "auto" | "auto" | Spatial regularization strength (auto-tuned to data scale) |
rho_sparsity |
float | 0.01 | L1 sparsity penalty |
n_hvg |
int | 2000 | Number of highly variable genes to select |
n_markers_per_type |
int | 50 | Top markers per cell type |
k_neighbors |
int | 6 | Neighbors for spatial graph |
max_iter |
int | 100 | Maximum BCD iterations |
tol |
float | 1e-4 | Convergence tolerance |
preprocess |
str | "log_cpm" | Preprocessing: "log_cpm" (recommended), "pearson", or "raw" |
random_state |
int | 0 | Random seed for reproducibility (scanpy convention) |
Attributes (After Fitting)
| Attribute | Shape | Description |
|---|---|---|
beta_ |
(n_spots, n_cell_types) | Raw cell type abundances |
proportions_ |
(n_spots, n_cell_types) | Normalized proportions (sum to 1) |
gene_idx_ |
(n_selected,) | Indices of genes used |
lambda_used_ |
float | Actual λ value used |
info_ |
dict | Optimization info (converged, n_iterations, final_objective) |
cell_type_names_ |
array | Cell type names (if provided) |
Input Data Formats
FlashDeconv accepts multiple input formats:
Spatial Data (Y)
- NumPy array: Dense (n_spots, n_genes)
- SciPy sparse matrix: CSR/CSC format (recommended for Visium HD to reduce memory usage)
- AnnData:
.Xor specified layer (e.g.,adata.layers["counts"])
Reference (X)
- NumPy array: Dense (n_cell_types, n_genes) signature matrix
- AnnData: Automatically aggregated from single-cell data via
prepare_data()using mean expression per cell type
Coordinates
- NumPy array: (n_spots, 2) for 2D spatial coordinates, or (n_spots, 3) for 3D (e.g., z-stacked sections)
- From AnnData: Automatically extracted from
.obsm["spatial"],.obsm["X_spatial"], or.obs[["x", "y"]]
Reference Data Quality
The algorithm is only as good as the reference data it's given.
Deconvolution accuracy depends critically on reference quality. Before running FlashDeconv, ensure your reference data meets these criteria:
| Requirement | Threshold | Why It Matters |
|---|---|---|
| Cells per type | ≥500 | Fewer cells → unstable signatures |
| Marker expression | ≥80% cells | Low expression → identity not captured |
| Marker fold-change | ≥5× | Low FC → cannot distinguish from others |
| Signature correlation | <0.95 | High correlation → algorithm cannot separate |
Common failure modes:
- Rare cell types with <200 cells produce noisy signatures
- Cell types annotated only by positive markers may include contaminants
- Similar subtypes (e.g., T cell subsets) often have correlation >0.98
📖 For detailed guidance: See Building High-Quality Reference Data — a comprehensive guide covering dual-marker annotation, QC pipelines, and troubleshooting.
Citation
If you use FlashDeconv in your research, please cite:
Plain text:
Yang, C., Chen, J. & Zhang, X. FlashDeconv enables atlas-scale, multi-resolution spatial deconvolution via structure-preserving sketching. bioRxiv (2025). https://doi.org/10.64898/2025.12.22.696108
BibTeX:
@article{yang2025flashdeconv,
title={FlashDeconv enables atlas-scale, multi-resolution spatial deconvolution via structure-preserving sketching},
author={Yang, Chen and Chen, Jun and Zhang, Xianyang},
journal={bioRxiv},
year={2025},
doi={10.64898/2025.12.22.696108},
url={https://doi.org/10.64898/2025.12.22.696108}
}
Contributing
Contributions are welcome! Please see CONTRIBUTING.md for guidelines.
License
This project is licensed under the BSD-3-Clause License.
Related Resources
- Paper Reproducibility: flashdeconv-reproducibility — Complete code to reproduce all figures and benchmarks
- Documentation: ReadTheDocs (coming soon)
- Issues & Support: GitHub Issues
Acknowledgments
We thank the developers of Spotless, Cell2Location, and RCTD for their benchmarking frameworks and methodological contributions to the spatial transcriptomics field.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file flashdeconv-0.1.4.tar.gz.
File metadata
- Download URL: flashdeconv-0.1.4.tar.gz
- Upload date:
- Size: 44.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
76b38aab6bee96f74b52fbc083b1b0d1c57d4fb978d6d76e65bc2a60b76ac254
|
|
| MD5 |
5c8dc4e08e68a98f2d17465555c60d7b
|
|
| BLAKE2b-256 |
1c7226d49a3a5f9e40b3ea8be5dbae636012dde6f083126d5701c47fd8165465
|
File details
Details for the file flashdeconv-0.1.4-py3-none-any.whl.
File metadata
- Download URL: flashdeconv-0.1.4-py3-none-any.whl
- Upload date:
- Size: 36.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.13.5
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
72c0d1bf821aa0b46e045103b480ae84924fd7efe7a28d20194ff4fbaed7e7ed
|
|
| MD5 |
46bee029db784b4bc93ae420c484b4b0
|
|
| BLAKE2b-256 |
ce874db20fd4c885c54a0c983c3ecde633304bb991a27b71ab8f6c7887ec775c
|















