EvoScientist: Towards Self-Evolving AI Scientists for End-to-End Scientific Discovery
Project description

English | 简体中文
EvoScientist aims to harness vibe research by enabling self-evolving AI scientists that autonomously explore, generate insights, and iteratively improve. It is designed to be opinionated and ready to use out of the box, offering a living research system that grows alongside evolving agent skills, toolsets, and memory bases. Moving beyond traditional human-in-the-loop systems, EvoScientist adopts a human-on-the-loop paradigm, where AI acts as a research buddy that co-evolves with human researchers and internalizes scholarly taste and scientific judgment.
🏆 Awards & Recognition

Best Paper & Appraisal Award |

AI-Generated Best Paper |

#1 on DeepResearch Bench II |
⚡ Unified Control, Different Surfaces
🖥️ CLI / TUI |
📱 Mobile |
|---|---|
| View demo video | View mobile demo |
✨ Features
- 🤖 Multi-Agent Team — 6 sub-agents (plan, research, code, debug, analyze, write) working in concert.
- 🧠 Persistent Memory — Context, preferences, and findings survive across sessions.
- 🌐 Multi-Provider — Anthropic, OpenAI, Google, NVIDIA — one config to switch.
- 📱 Multi-Channel — CLI as the hub; Telegram, Slack, Feishu, WeChat, and more — one agent session.
- 🔬 Scientific Workflow — Intake → plan → execute → evaluate → write → verify.
- 🔌 MCP & Skills — Plug in MCP servers or install skills from GitHub on the fly.
[!TIP] Looking for ready-to-use research skills? Check out EvoSkills — powered by EvoScientist's engine and installable skills, the entire end-to-end research lifecycle is covered out of the box. EvoSkills are also compatible with other AI coding agents like Claude Code, Cursor, and OpenClaw.
🔥 News
- [13 Mar 2026] 🚀 EvoScientist officially debuts!
- [11 Mar 2026] ⛳ Technical Report is live! Check it out 👈
- [06 Mar 2026] 🥇 Ranked #1 on DeepResearch Bench II at submission time! Leaderboard 👈
- [24 Nov 2025] 🏆 6/6 accepted at ICAIS 2025 AI Scientist Track — Best Paper & AI Reviewer's Appraisal Award! Details 👈
📖 Table of Contents
- 📦 Installation
- 🔑 Configuration
- ⚡ Quick Start
- 🔌 MCP Integration
- 📱 Channels
- 📚 Acknowledgments
- 🎯 Roadmap
- 🌍 Project Roles
- 🤝 Contributing
- 📝 Citation
📦 Installation
[!TIP] Requires Python 3.11+ (< 3.14). We recommend uv or conda for dependency management and virtual environments.
🪛 Install uv (if you don't have it)
curl -LsSf https://astral.sh/uv/install.sh | sh
Quick Install
uv tool install EvoScientist
Or install into the current environment instead:
uv pip install EvoScientist
Development Install
git clone https://github.com/EvoScientist/EvoScientist.git
cd EvoScientist
uv sync --dev
Using conda
conda create -n EvoSci python=3.11 -y
conda activate EvoSci
pip install -e ".[dev]"
Using PyPi
pip install EvoScientist # quick install
pip install -e ".[dev]" # development install
Optional: Channel dependencies
Messaging channel integrations require extra dependencies. Install only what you need:
uv pip install "EvoScientist[telegram]" # Telegram
uv pip install "EvoScientist[discord]" # Discord
uv pip install "EvoScientist[slack]" # Slack
uv pip install "EvoScientist[wechat]" # WeChat
uv pip install "EvoScientist[qq]" # QQ
uv pip install "EvoScientist[feishu]" # Feishu
uv pip install "EvoScientist[all-channels]" # everything
Upgrade to the latest code base
git pull && uv sync --dev
🔑 Configuration
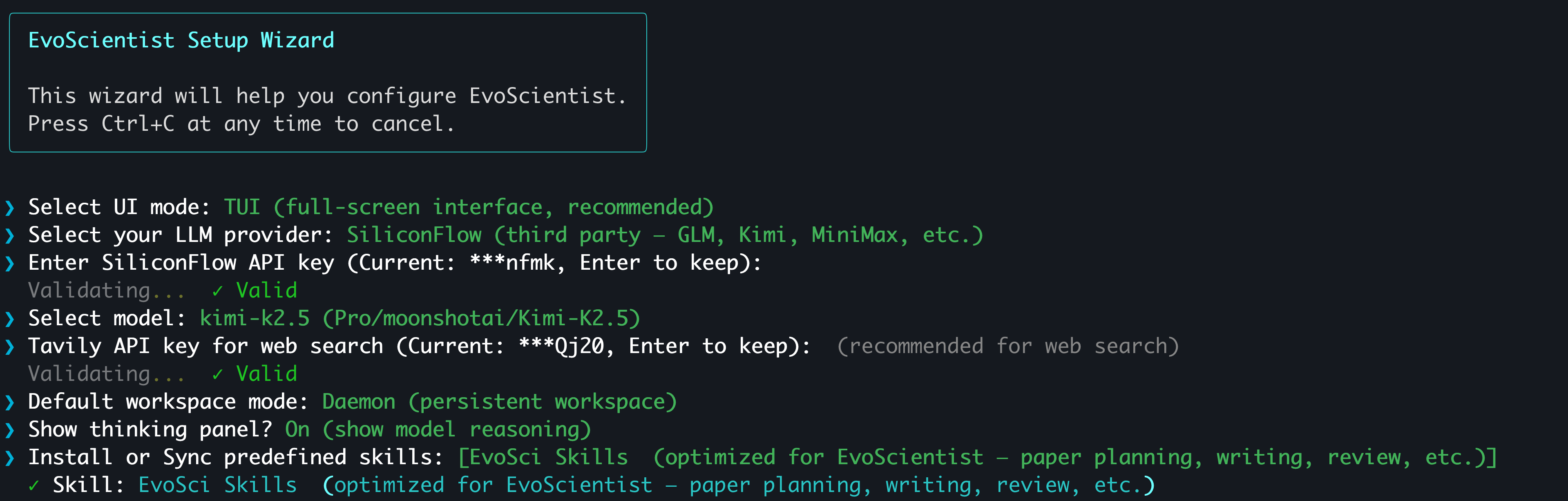
The easiest way to configure API keys is the interactive wizard:
EvoSci onboard
[!TIP] It walks you through provider selection, key validation, model choice, and workspace mode. Supports OAuth sign-in for Claude Code and Codex CLI users — no API key needed.
📟 Manual configuration via environment variables
Set at least one LLM provider key and (optionally) a search key:
# Pick one LLM provider
export ANTHROPIC_API_KEY="sk-..." # Claude — console.anthropic.com
export OPENAI_API_KEY="sk-..." # GPT — platform.openai.com
export GOOGLE_API_KEY="AI..." # Gemini — aistudio.google.com/api-keys
export NVIDIA_API_KEY="nvapi-..." # NIM — build.nvidia.com
# Web search (optional)
export TAVILY_API_KEY="tvly-..." # app.tavily.com
Or use EvoSci config set to persist keys in ~/.config/evoscientist/config.yaml.
Alternatively, copy the example .env file for project-level configuration:
cp .env.example .env # then fill in your keys
⚠️ Never commit
.envfiles with real keys. It is already in.gitignore.
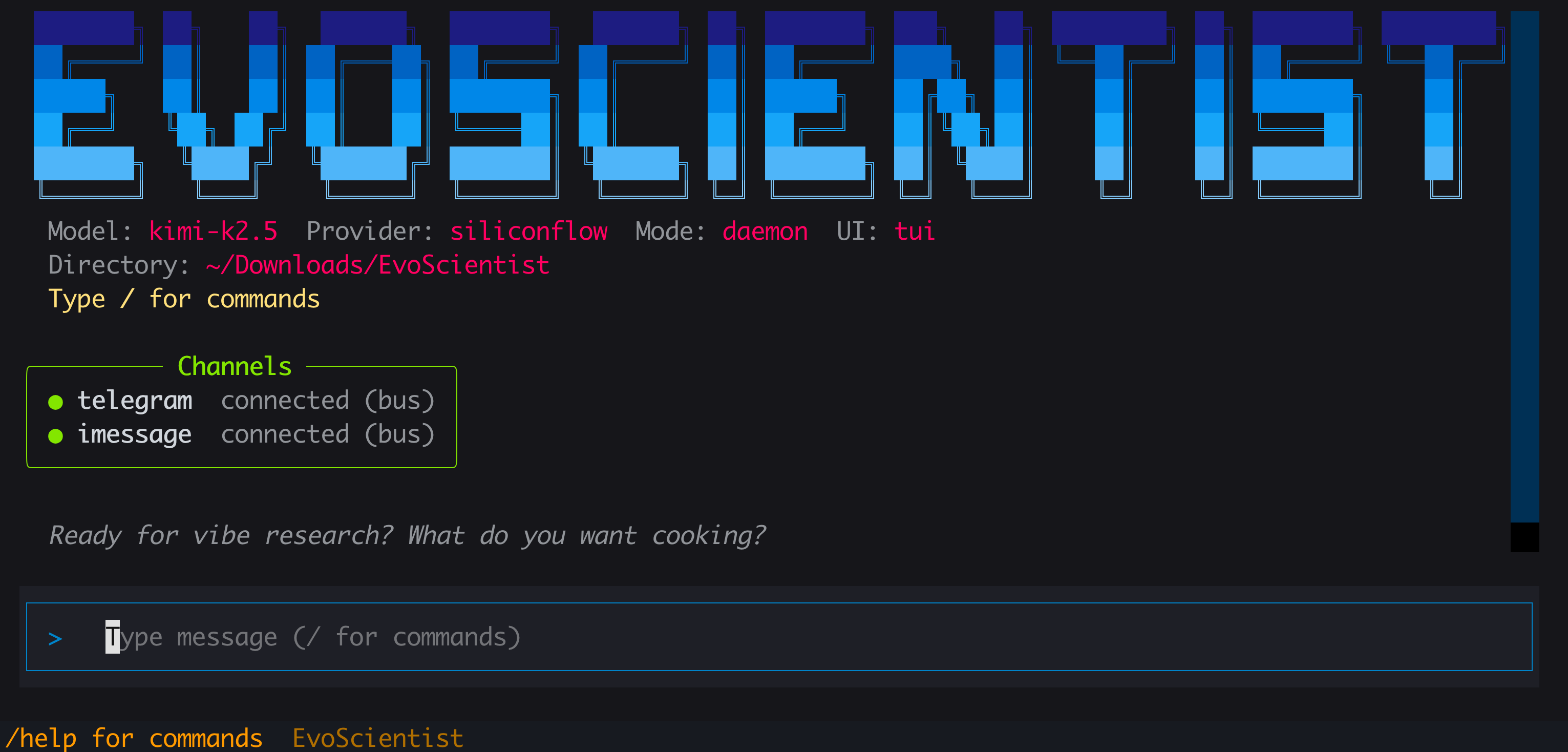
⚡ Quick Start
EvoSci # or EvoScientist — interactive mode (TUI by default)
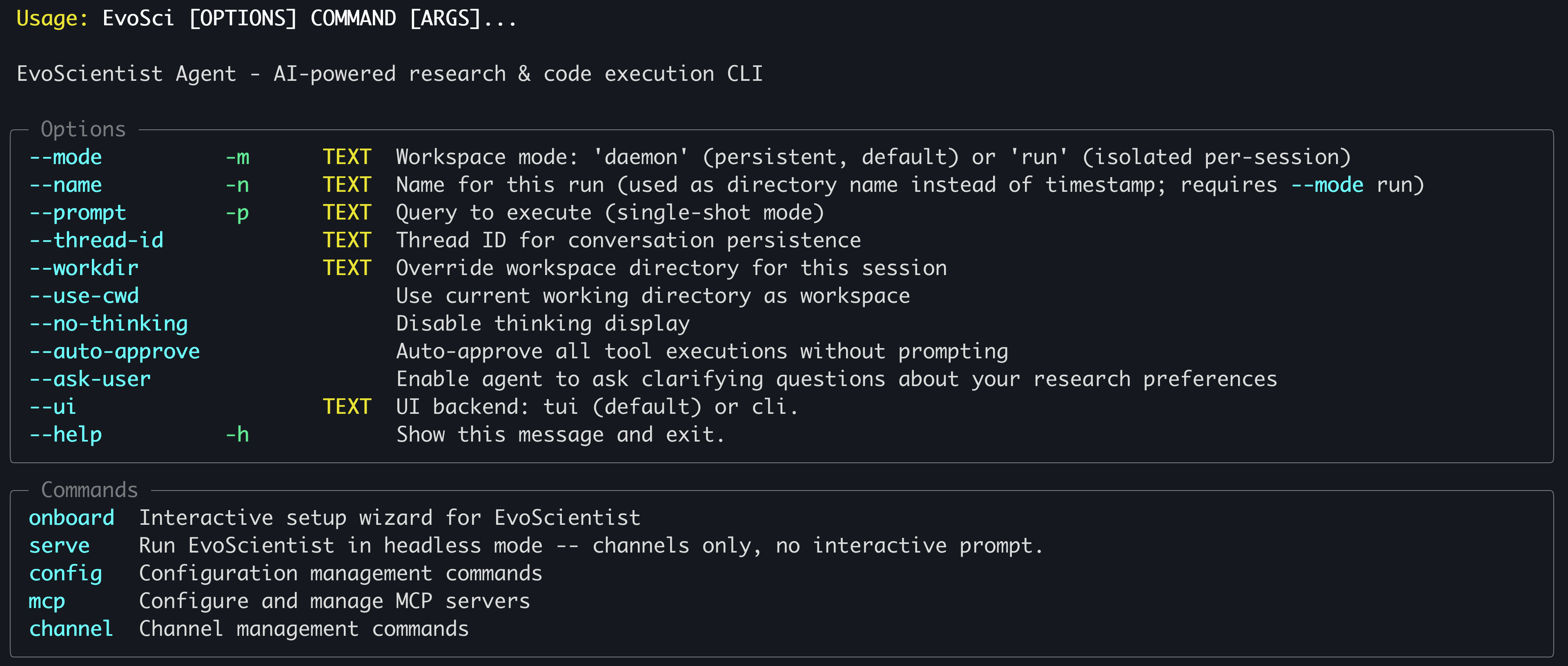
Run
EvoSci -hfor all CLI options.
Common examples
EvoSci # interactive mode (TUI by default)
EvoSci -p "your question" # single-shot mode
EvoSci --workdir /path/to/project # open in a specific directory
EvoSci -m run # isolated per-session workspace
EvoSci --ui cli # classic CLI (lightweight)
EvoSci serve # headless mode — channels only, no interactive prompt
Action Approval
By default, shell commands (execute tool) require human approval before running. To skip approval prompts:
# Per-session: auto-approve via CLI flag
EvoSci --auto-approve
EvoSci -p "query" --auto-approve
# Persistent: set in config (applies to all future sessions)
EvoSci config set auto_approve true
# Or allow only specific command prefixes
EvoSci config set shell_allow_list "python,pip,pytest,ruff,git"
During a session you can also reply 3 (Approve all) at any approval prompt to auto-approve for the rest of that session.
Agent Questions
The agent can proactively ask you questions when it needs clarification (e.g., dataset choice, experiment direction). This is enabled by default. To disable:
# Persistent: set in config
EvoSci config set enable_ask_user false
# Re-enable
EvoSci config set enable_ask_user true
In-session commands
| Command | Description |
|---|---|
/current |
Show current session info |
/threads |
List recent sessions |
/resume |
Resume a previous session |
/delete |
Delete a saved session |
/new |
Start a new session |
/clear |
Clear chat history |
/skills |
List installed skills |
/install-skill <src> |
Add a skill from path or GitHub |
/uninstall-skill <name> |
Remove an installed skill |
/mcp |
Manage MCP servers |
/channel |
Configure messaging channels |
/help |
Show available commands |
/exit |
Quit |
Script Inference
from EvoScientist import EvoScientist_agent
from langchain_core.messages import HumanMessage
from EvoScientist.utils import format_messages
thread = {"configurable": {"thread_id": "1"}}
last_len = 0
for state in EvoScientist_agent.stream(
{"messages": [HumanMessage(content="Hi?")]},
config=thread,
stream_mode="values",
):
msgs = state["messages"]
if len(msgs) > last_len:
format_messages(msgs[last_len:])
last_len = len(msgs)
🔌 MCP Integration
Add external tools via MCP servers with a single command:
# Usage
EvoSci mcp add <name> <command> [-- args...]
# Example
EvoSci mcp add sequential-thinking npx -- -y @modelcontextprotocol/server-sequential-thinking
[!TIP] For command options, config fields, tool routing, wildcard filtering, and troubleshooting, see the MCP Integration Guide.
📱 Channels
Connect messaging platforms so they share the same agent session as the CLI:
# Usage
EvoSci channel setup <channel>
# Example
EvoSci channel setup telegram
Multiple channels can run concurrently — comma-separate names in the config:
channel_enabled: "telegram,slack,feishu,qq"
The channel can also be started interactively with /channel in the CLI session.
[!TIP] For per-channel setup guides, capability matrix, architecture details, and troubleshooting, see the Channel Integration Guide.
📚 Acknowledgments
This project builds upon the following outstanding open-source works:
- LangChain — A framework for building agents and LLM-powered applications.
- DeepAgents — The batteries-included agent harness.
We thank the authors for their valuable contributions to the open-source community.
🎯 ᯓ➤ Roadmap
Coming soon:
- 🖥️ Full-screen TUI and classic CLI interfaces
- 📻 EvoMemory v1.0 shipped
- ⚒️ 200+ predefined skills built in
- 🧩 Built-in research-lifecycle skills shipped
- 👋 Human-in-the-loop action approval
- 🦾 Agent-initiated human clarification
- 📑 Technical report on the way
- 🔐 OAuth sign-in (Anthropic, OpenAI, etc.)
- 📺 Web app with workspace UI
- 📹 Demo and tutorial in the works
- 📊 Benchmark suite to be released
- ⏰ Scheduled tasks for the core system planned
Stay tuned — more features are on the way!
🌍 Project Roles
Core Contributors

Xi Zhang |

Yougang Lyu |

Dinos Papakostas |

Yuyue Zhao |

Ziheng Zhang |

Xiaohui Yan |
Contributors
Jan Piotrowski, Wiktor Cupiał, Jakub Kaliski, Jakub Filipiuk, Xinhao Yi, Shuyu Guo, Andreas Sauter, Wenxiang Hu, Jacopo Urbani, Zaiqiao Meng, Jun Luo, Lun Zhou
Xiaoyi DeepResearch Team and the wider open-source community contribute to this project.
For any inquiries or collaboration opportunities, please contact: EvoScientist.ai@gmail.com
🤝 Contributing

We welcome contributions from developers, researchers, and AI coding agents at all levels. Our Contributing Guidelines are designed for both humans and AI agents — covering architecture, patterns, extension guides, and code standards to help you contribute safely and effectively.
👥 Community Contributors
⚗️ Join the EvoScientist community to discuss AI-driven research, share experiment results, and help shape the future of automated scientific discovery.
- Discord — Ask questions, share findings, and collaborate with researchers and developers in real-time.
- WeChat — Connect with our Chinese-speaking research community.
Every contribution brings us one step closer to a future where AI accelerates scientific breakthroughs for all of humanity.

📈 Star History
📝 Citation
If you find our paper and code useful in your research and applications, please cite using this BibTeX:
@article{evoscientist2026,
title={EvoScientist: Towards Multi-Agent Evolving AI Scientists for End-to-End Scientific Discovery},
author={Yougang Lyu and Xi Zhang and Xinhao Yi and Yuyue Zhao and Shuyu Guo and Wenxiang Hu and Jan Piotrowski and Jakub Kaliski and Jacopo Urbani and Zaiqiao Meng and Lun Zhou and Xiaohui Yan},
journal={arXiv preprint arXiv:2603.08127},
year={2026}
}
📜 License
This project is licensed under the Apache License 2.0 - see the LICENSE file for details.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file evoscientist-0.0.2.tar.gz.
File metadata
- Download URL: evoscientist-0.0.2.tar.gz
- Upload date:
- Size: 490.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6663e54277ef31c3cb9a99932204fb11d010656dfb85cf372aaaf74fc22a2a2f
|
|
| MD5 |
fab46ad7a02a392eb38feb97671b4a29
|
|
| BLAKE2b-256 |
5bf1ed301691ce8bb5c2a7f2d625f951a342b392bf5a2e8771f67cf1dd54f8ce
|
File details
Details for the file evoscientist-0.0.2-py3-none-any.whl.
File metadata
- Download URL: evoscientist-0.0.2-py3-none-any.whl
- Upload date:
- Size: 477.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.14
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7317586f2ec0300f4d5be4793cafd2c096c28cd9d1fb0523ff8101c917f77039
|
|
| MD5 |
aa063dfea65d4ccbc5ca3d8dac52866e
|
|
| BLAKE2b-256 |
3699b5caae9f73cc13bb3e2645730d2474084c9d63352f7edc0c54e47068883f
|