preprocess microbiome data
Project description
Preprocessing for 16S values.
The input file for the preprocessing should contain detailed unnormalized OTU/Feature values as a biom table, the appropriate taxonomy as a tsv file, and a possible tag file, with the class of each sample. The tag file is not used for the preprocessing, but is used to provide some statistics on the relation between the features and the class. You can also run the preprocessing without a tag file.
input
Here is an example of how the input OTU file should look like : (file example)
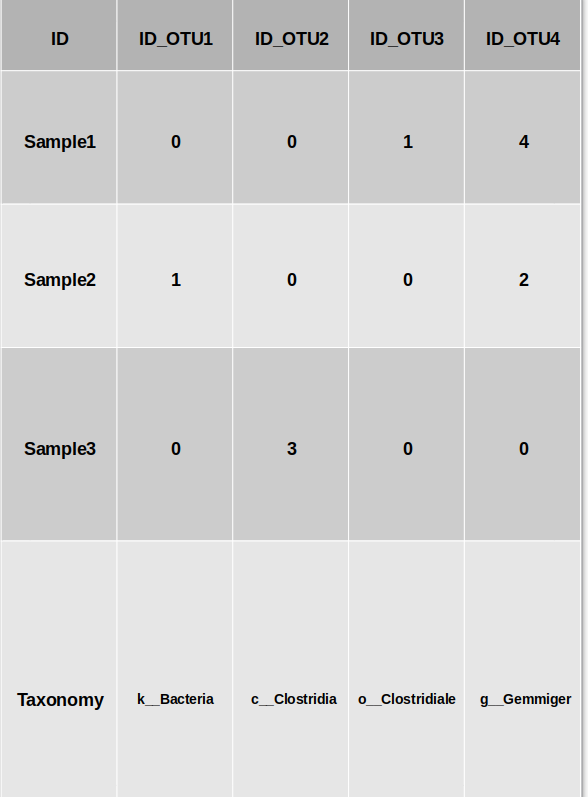
Parameters to the preprocessing
Now you will have to select the parameters for the preprocessing.
- The taxonomy level used - taxonomy sensitive dimension reduction by grouping the bacteria at a given taxonomy level. All features with a given representation at a given taxonomy level will be grouped and merged using three different methods: Average, Sum or Merge (using PCA then followed by normalization).
- Normalization - after the grouping process, you can apply two different normalization methods. the first one is the log (10 base)scale. in this method
x → log10(x + ɛ),where ɛ is a minimal value to prevent log of zero values.
The second methos is to normalize each bacteria through its relative frequency.
If you chose the Log normalization, you now have four standardization
possibilities:
a) No standardization
b) Z-score each sample
c) Z-score each bacteria
d) Z-score each sample, and Z-score each bacteria (in this order)
When performing relative normalization, we either dont standardize the results or performe only a standardization on the bacteria.
- Dimension reduction - after the grouping, normalization and standardization you can choose from two Dimension reduction method: PCA or ICA. If you chose to apply a Dimension reduction method, you will also have to decide the number of dimensions you want to leave.
How to use
use MIPMLP.preprocess(input_df)
####parameters:
taxonomy_level 4-7 , default is 7
taxnomy_group : sub PCA, mean, sum, default is mean
epsilon: 0-1
z_scoring: row, col, both, No, default is No
pca: (0, 'PCA') second element always PCA. first is 0/1
normalization: log, relative, default is log
norm_after_rel: No, relative, default is No
output
The output is the processed file.
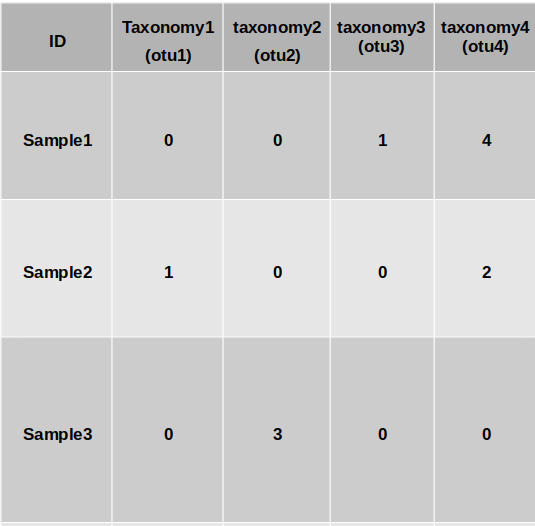
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file MIPMLP-1.1.4.tar.gz.
File metadata
- Download URL: MIPMLP-1.1.4.tar.gz
- Upload date:
- Size: 9.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dfaded357f3b382f2ab88b715d9a79567bd515e5528ed596ffc4a4cf26c1b12c
|
|
| MD5 |
f825204df9262ccf12005f21a4e696e9
|
|
| BLAKE2b-256 |
bd3b0aa8649a02c858862aab209cd2981c0e8f49781f2510114f82c79c724246
|
File details
Details for the file MIPMLP-1.1.4-py3-none-any.whl.
File metadata
- Download URL: MIPMLP-1.1.4-py3-none-any.whl
- Upload date:
- Size: 11.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
87e586318c03210d9c966bc68a9c37b75314e9c93ada37765806e67b339319f1
|
|
| MD5 |
1dab467052b67a6e69227f679ef92e14
|
|
| BLAKE2b-256 |
9d5b8bbe8f012d9e093a0e11786c7ca0d874e8bcc00480a0ab98d48b3ff22aa7
|











