No project description provided
Project description
MartiniGlass: seeing your martini is believing
MartiniGlass uses vermouth to stably rewrite your input topology files as ones that can be used for visualisation in VMD. Although much of the functionality is focused on proteins, and the program has a particular focus on being able to visualise protein secondary/tertiary structure networks, MartiniGlass can in fact be used to reconstruct bonded networks of any Martini molecule!
Previously, cg_bonds-vX.tcl was able to read in Martini system topology information and draw it directly in VMD.
However, many Martini models now make extensive use of interaction types like virtual sites, which can't be handled
by the topology reading capabilities of this script directly. Martini_vis handles these and more by rewriting
virtual sites as bonded atoms for visualisation purposes.
Thanks to Jan Stevens for vis.vmd
If the solution here isn't working for you, please open an issue!
Documentation
Documentation for MartiniGlass is available on the readthedocs site. The documentation covers multiple use cases and runs through the tutorials in the examples folder step by step.
Installation
python3 -m venv venv && source venv/bin/activate # Not required, but often convenient.
pip install git+https://github.com/Martini-Force-Field-Initiative/MartiniGlass
Usage
Most likely, you can use MartiniGlass with a simple command:
martiniglass -p topol.top
But if you have proteins with complex tertiary structure networks that you also want to see, you might need to give some extra options. More comprehensive documentation and tutorials are available on the readthedocs site. If you think something is broken or have a feature request, please open an issue.
Gallery
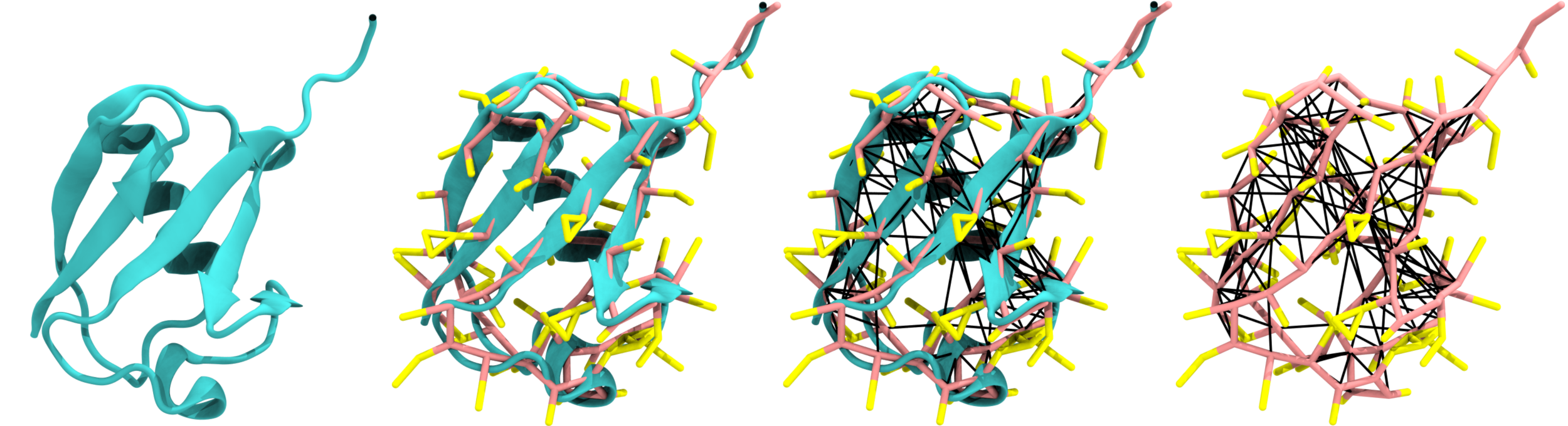
For a practical example of how MartiniGlass can be used to generate visualisable topologies, and see the expected output, there is an example of the 1UBQ system shown below in the examples folder. Here are some illustrations of how MartiniGlass can be used to visualise your molecules
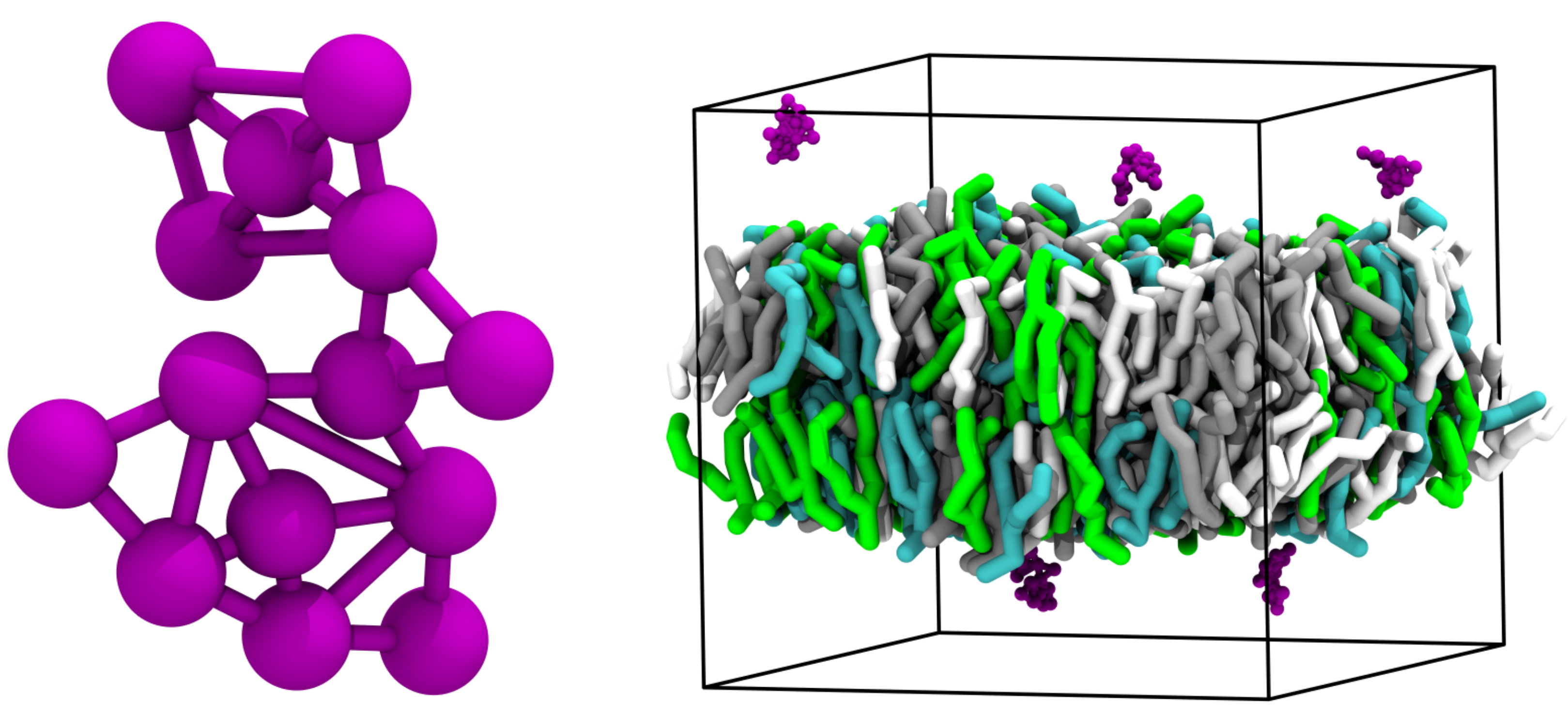
Non-protein systems. Left: the molecular motor of Vainikka and Marrink, a synthetic molecule with complex topology, showing how the beads are connected using the CPK mode in VMD. Right: Several molecular motors together with a lipid bilayer of several different lipid types.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file martiniglass-1.1.0.tar.gz.
File metadata
- Download URL: martiniglass-1.1.0.tar.gz
- Upload date:
- Size: 17.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0b3808ec4f34699f893df9355c70bd4af6abb3fe7730970eb22960444d3d3713
|
|
| MD5 |
58b175b9fac7ffaf640e0bb4884a5850
|
|
| BLAKE2b-256 |
478be4708b226643bb2c09973cb59c134f673f861837755a1c649bb99b6f03a0
|
File details
Details for the file MartiniGlass-1.1.0-py3-none-any.whl.
File metadata
- Download URL: MartiniGlass-1.1.0-py3-none-any.whl
- Upload date:
- Size: 23.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.10.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6f883aa64bae0e77b6fd88834470bbdaf0002fd383cf94906b78ae7cfcc1e77b
|
|
| MD5 |
eb9ea5b386d236ce2581e9d4ae26c998
|
|
| BLAKE2b-256 |
9bacc34944e5f1ccfcbe987ce1985a67f8b17920e885ddc706449fbc80b897ff
|