TCRcloud is a TCR repertoire visualization and comparison tool
Project description





TCRcloud is a TCR repertoire visualization and comparison tool
Instalation
TCRcloud is written in python and can be installed from PyPI using pip:
pip3 install TCRcloud
Currently it is only compatible with Linux (x86-64) and macOS (x86-64) because one of the dependencies used is also only compatible with those operating systems. It does work on WSL if you want to use Windows.
TCRcloud uses the AIRR Data Commons API and needs AIRR compliant data as input.
TCRcloud was initially developed for TCR repertoires but it is also compatible with BCR repertoires.
Usage
To create a word cloud
TCRcloud cloud repertoire.airr.rearrangements.tsv
By default TCRcloud colours the CDR3 based on the Vgene but you can provide a json file that atributes colours in Hex format to specific sequences:
{
"#FF0000":["CAASITGNQFYF","CAVREDGTSGSARQLTF"],
"#0000FF":["CAVMDSNYQLIW"]
}
The sequences not in the file will be coloured grey.
Only the colours for human TCR and BCR variable genes are coded into TCRcloud.
To use your custom colours for the word cloud
TCRcloud cloud repertoire.airr.rearrangements.tsv -c colours.json
To create a word cloud without a legend
TCRcloud cloud repertoire.airr.rearrangements.tsv -l False
To create a radar plot comparing diversity metrics
TCRcloud radar repertoire.airr.rearrangements.tsv
By default TCRcloud uses repertoire_id but you can create a legend with the text you want by providing a json file:
{
"2839362682105696746-242ac113-0001-012":"Twin 2A",
"2939134772391776746-242ac113-0001-012":"Twin 2B"
}
To create a radar plot with your desired legend
TCRcloud radar repertoire.airr.rearrangements.tsv -c legend.json
To create a radar plot without a legend
TCRcloud radar repertoire.airr.rearrangements.tsv -l False
To export the calculated metrics from the radar to a text file
TCRcloud radar repertoire.airr.rearrangements.tsv -e True
The metrics utilized in the radar
Distinct CDR3: A simple count of the unique CDR3 sequences in the sample
D50 Index: Developed by Dr Jian Han, this metric represents the percent of dominant and unique T or B cell clones that account for the cumulative 50% of the total CDR3 counted in the sample
Convergence: Frequency of CDR3 amino acid sequences that are coded by more than one nucleotide sequence
Gini Index: Originally developed by Dr Corrado Gini to represent the wealth inequality within a social group, this metric is a measure of distribution, with 0 representing perfect equality and 1 representing perfect inequality between CDR3
Shannon Index: Originally developed by Dr Claude Shannon to quantify the entropy in strings of text, this metric takes into account the number of CDR3 present, as well as, the relative abundance of each CDR3 and higher the number, the higher is the species diversity
Simpson Index: Originally developed by Dr Edward H. Simpson to measure diversity in a ecosystem, this metric measures the probability that two randomly selected CDR3 are different
Chao1 index: Originally developed by Dr Anne Chao to estimate richness in a community, this metric indicates the estimated number of CDR3 in a sample
Using TCRcloud you can download rearragements files from the AIRR compliant databases based on AIRR repertoire metadata files
To download AIRR rearrangements files
TCRcloud download repertoire.airr.json
TCRcloud provides some test data to experiment the tool. The data is one twin pair from the monozygotic twins study from the Mark Davis lab (DOI: 10.1038/ncomms11112)
To download the test data repertoire file
TCRcloud testdata
After having the testdata.airr.json file you can use the download function included in TCRcloud to get the matching rearragements file.
Examples:
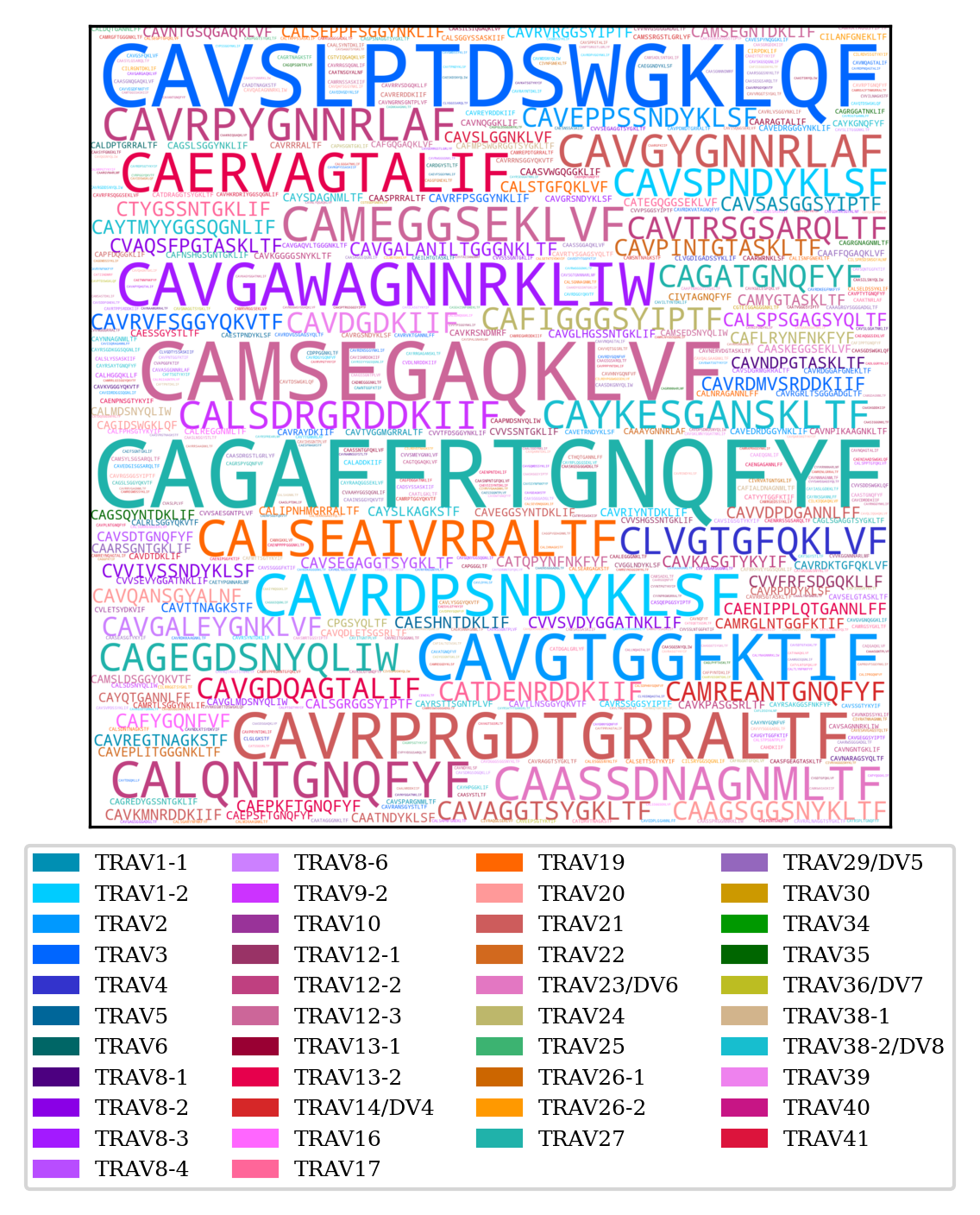
TRA CDR3 word cloud
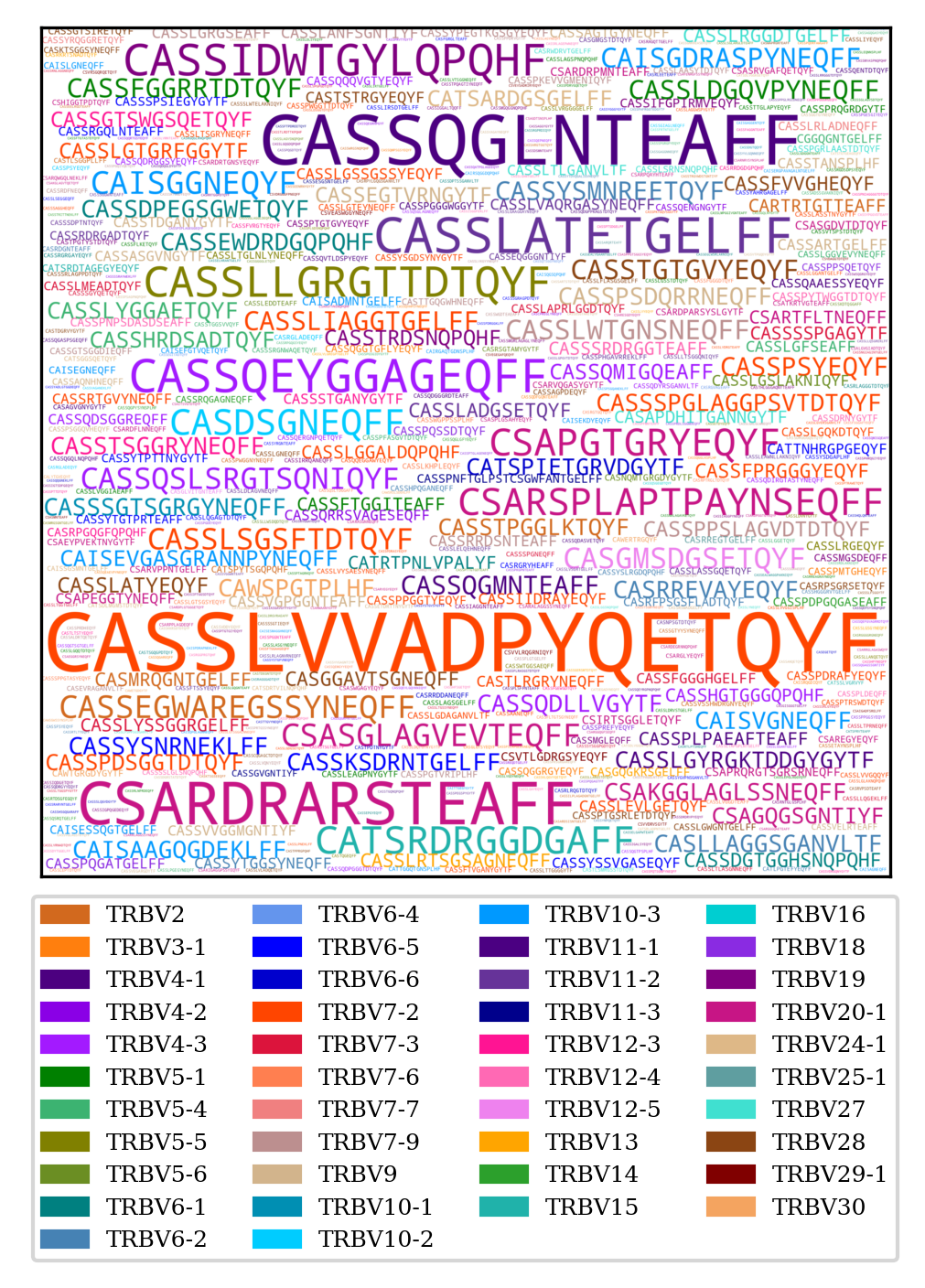
TRB CDR3 word cloud
TRG CDR3 word cloud
TRD CDR3 word cloud
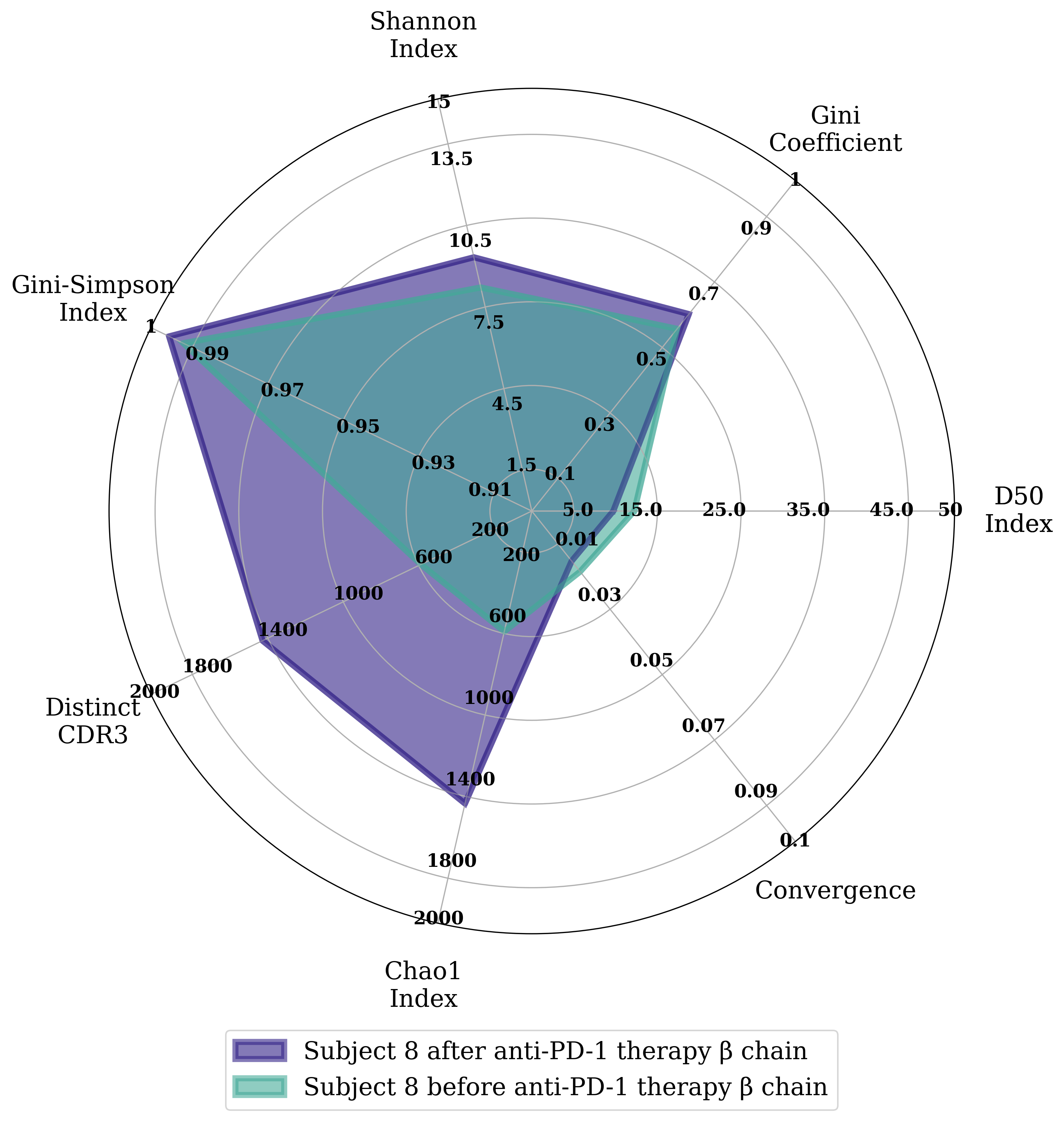
Diversity comparison
Comparing here the αβ repertoire from one twin pair from the monozygotic twins study from the Mark Davis lab (DOI:10.1038/ncomms11112)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file TCRcloud-1.3.1.tar.gz.
File metadata
- Download URL: TCRcloud-1.3.1.tar.gz
- Upload date:
- Size: 16.1 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.10.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
953a28ffe5081bd481e3f31cf218a67ae063d46035d0dceb6facc02d1a511882
|
|
| MD5 |
88d231042a46b9c09760f5599798ffdb
|
|
| BLAKE2b-256 |
79c16b94b29ae3e84d2bb2f3b8a3260e40cc05870d4d4b6232f96ad09d0ff0bf
|
File details
Details for the file TCRcloud-1.3.1-py3-none-any.whl.
File metadata
- Download URL: TCRcloud-1.3.1-py3-none-any.whl
- Upload date:
- Size: 17.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.10.6
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
bf4223a487c733c67a4b7ae9cd0f94f7d24c297c25d8060aaebf617dff657201
|
|
| MD5 |
4f996d5aabf38eaa5b240beec272dfeb
|
|
| BLAKE2b-256 |
429302283f28b9583d1f020ddda7858065415c5809dc60d3cd9551f9fbafd3db
|