Generated from aind-library-template
Project description
aind_hierarchical_bootstrap
Implementation of a hierarchical bootstrap for pandas dataframes. This package can be used to perform two useful things:
- Estimate the confidence interval that takes into account the hierarchical nature of the data
- Perform a statistical test to compare the mean value between two groups that have hierachical structure.
Everything is generally applicable to any nested dataset, although my use case is neuroscience specific. I made some simple attempts to speed up the implementation, but it is still time consuming. I followed the procedure outlined in:
Application of the hierarchical bootstrap to multi-level data in neuroscience (2020). https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7906290/
Contact: alexpiet [at] gmail [dot] com
Installation
I recommend installing within a conda environment.
git clone https://github.com/AllenNeuralDynamics/aind_hierarchical_bootstrap.git
cd aind_hierarchical_bootstrap
pip install .
Demonstration
I developed a simple demonstration of why a hierarchical approach is important when data is nested. The demonstration function will generate synthetic data with a nested structure. Importantly, there is no difference between the generative means of the two processes, but the variability at the two levels of the nested data means that a naive t-test will return a signficant result, while a hierarchical approach will show no effect.
In this synthetic dataset there are two groups, colored black and mageneta. Each group has a hierarchical structure. As a specific example we might consider the groups two be an experimental manipulation, and the levels as nested observations (Level 1 could be behavioral sessions, level 2 could be cells, and level 3 is each measurement from a single cell).
import aind_hierarchical_bootstrap.make_data as md
df,bootstraps, stats_df = md.demonstrate_levels()
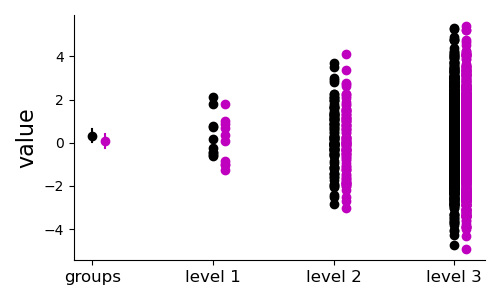
A naive t-test results in a false positive, whereas the hierarchical sampling approach correctly finds no significant difference between the groups.
General use
Organize your data in the pandas "tidy" format, where each row is a single observation. Each level of the nested structure should be defined in a column, as well as any top level groups. The observation variable should be its own column.
Estimating the standard error of the mean with nested data
For example if we have a dataframe of observations "response" and nested hierarchies "level_1" and "level_2", then we can compute the bootstraps with different levels of hierachical bootstrapping. You can use the df variable from the demonstration script above to follow along these examples. To compute non-hierarchical bootstraps, which is just sampling with replacement from all observations, regardless of hierarchy (sample once):
import aind_hierarchical_bootstrap.bootstrap as hb
bootstraps = hb.bootstrap(df, metric='response',levels=[], nboots=10000)
bootstraps is a dictionary with:
<metric>the sampled value of the observation metric for each bootstrap iteration, will have length ofnboots<metric>_semthe estimated standard error of the mean of the observation metric, computed as the standard deviation of the bootstrapped samples. In this case this should be very close todf['response'].sem()groups, the list of top-levels groups, in this case just['response']
To sample with one level of hierarchy, which means we sample with replacement from elements of "level_1", then sample with replacement from all observations within that element of level_1 but ignoring all structure below that (level_2, level_3). Note we are sampling twice here. Note now that the estimated SEM differs significantly from the naive approach above.
import aind_hierarchical_bootstrap.bootstrap as hb
bootstraps = hb.bootstrap(df, metric='response',levels=['level_1'], nboots=10000)
To sample with two levels of the hierarchy, which means we sample with replacement from elements of "level_1", then sample with replacement from "level_2" elements within that element of level_1, then finally sample from all observations within that level_1, level_2 element. Note we sample three times.
import aind_hierarchical_bootstrap.bootstrap as hb
bootstraps = hb.bootstrap(df, metric='response',levels=['level_1','level_2'], nboots=10000)
How many nesting steps you should take depends on the variance at each level of your dataset. The code should work for as many levels as you want, but performance will suffer greatly from each additional level.
Comparing groups and hypothesis testing
If you have multiple top-level manipulations you want to compare, then you can specify that too:
bootstraps = hb.bootstrap(df, metric='response',levels=['level_1','level_2'], top_level='group',nboots=10000)
Now bootstraps is a dictionary with two entries for each unique group:
<group label>the sampled value of the observation metric for each bootstrap iteration, only for this group. This will have levelnboots.<group label>_semthe estimatd standard error of the mean of the observation metric for this group.
You can perform statistical testing with:
import aind_hierarchical_bootstrap.stats as stats
stats_df = stats.compute_stats(bootstraps)
stats_df is a dataframe with one row for each hypothesis test performed. Each row has the following columns:
- name, which is the combination of the two groups compared
- p, the fraction of bootstrap iterations where group1 > group2
- nboots, the number of bootstrap iterations used in the test (inherited from
bootstraps)
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file aind_hierarchical_bootstrap-0.0.7.tar.gz.
File metadata
- Download URL: aind_hierarchical_bootstrap-0.0.7.tar.gz
- Upload date:
- Size: 12.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
58d6a37cfb244f2e14304e42318509ee303fadaccc52d0dd77ed42eddedaa663
|
|
| MD5 |
1370ccbc99bc2d23e61ee4a9a4f48475
|
|
| BLAKE2b-256 |
ed27d4178e595b6bb27cb77081224ec54c1230b2ab8dec048c21a13e2a39d498
|
File details
Details for the file aind_hierarchical_bootstrap-0.0.7-py3-none-any.whl.
File metadata
- Download URL: aind_hierarchical_bootstrap-0.0.7-py3-none-any.whl
- Upload date:
- Size: 11.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0fb88729ea966316e0b476796b724f9121f72e4342372813087b0c2eb5d25db3
|
|
| MD5 |
0b64efb4b548ad8f42db41eba4ea1368
|
|
| BLAKE2b-256 |
d210029e5c8b1e8364384e5c26aa0ec5e3d51afeec50c87ba7b3f78af7b56572
|