AlphaFold3 ZIP → Standalone HTML Report with PAE heatmaps, confidence scores, and interface analysis
Project description
alpha_fold_viewer
AF3 ZIP → Standalone HTML Report — a single-file tool that converts AlphaFold3 output ZIP files into beautiful, self-contained HTML reports.
What it does
Takes an AlphaFold3 prediction ZIP file and produces a single HTML file containing:
- Input summary — sequences in FASTA format with copy buttons, chain types, lengths
- Confidence overview — all models ranked by
ranking_scorewith ipTM, pTM, fraction disordered, clash status - Chain sequences — full sequences with per-residue pLDDT coloring and interface residue highlighting
- Sequence heatmap strips — linear pLDDT bars with interface position markers
- PAE heatmaps — full predicted aligned error matrices with chain boundaries (embedded as base64 images)
- Interface analysis — inter-chain contacts with residue counts, mean PAE, pLDDT, and high-confidence percentages
- Per-model details — collapsible sections with chain info and interface residue ranges
The HTML is fully standalone — all images are embedded as data URIs, CSS is inline, no external dependencies. Open it in any browser, share via email, or include in presentations.
Works with any AlphaFold3 output: protein homodimers, heterodimers, protein+DNA complexes, multi-chain assemblies.
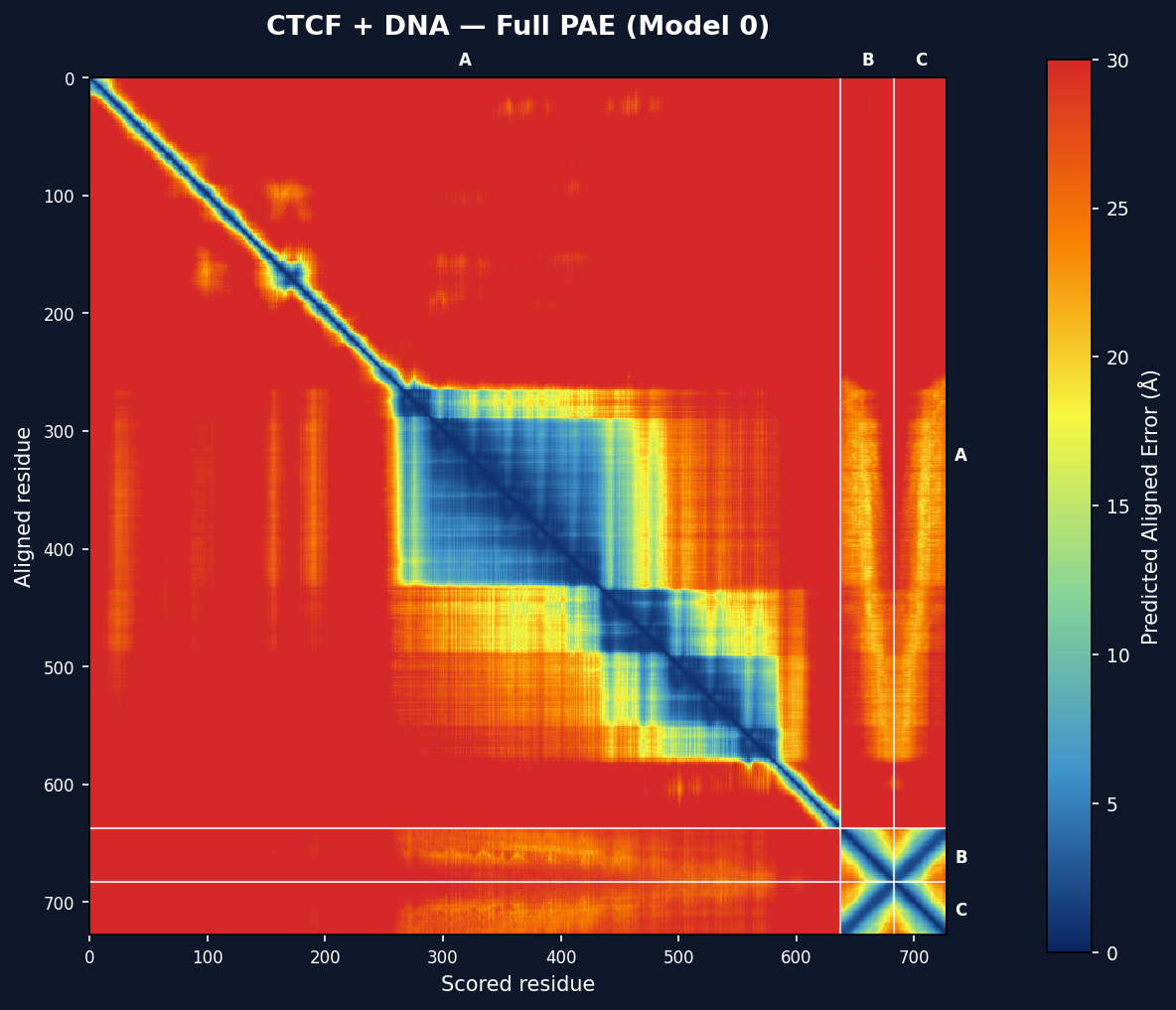
PAE Heatmap (CTCF protein + DNA complex, 3 chains)
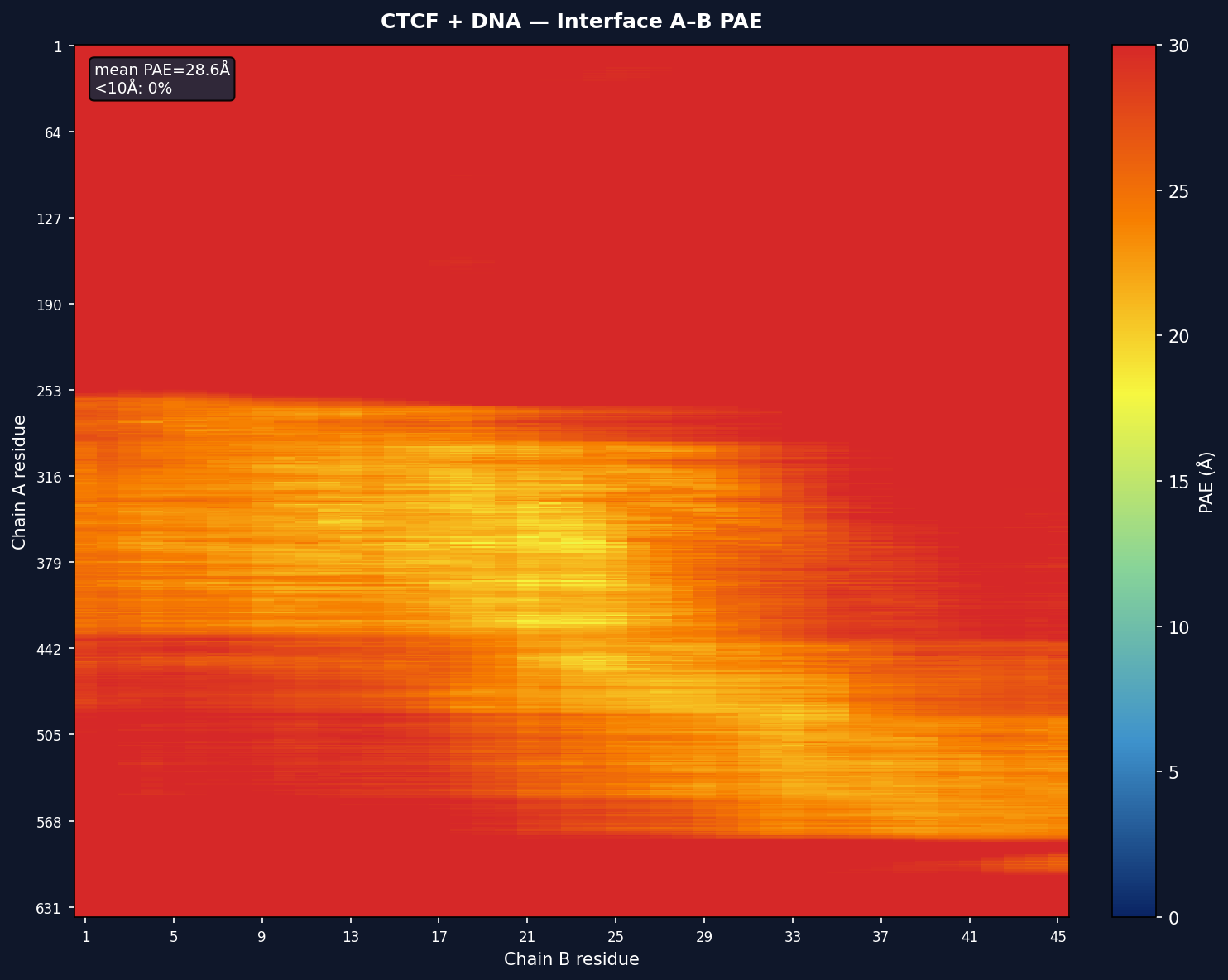
Interface PAE Sub-matrix
Sequence Strip with pLDDT and Interface Markers
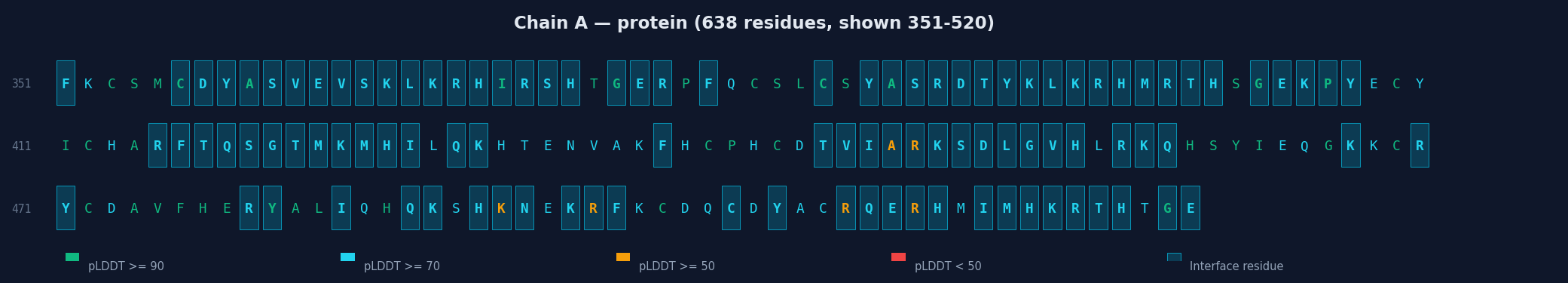
Colored Sequence with Interface Residues
Installation
From PyPI
pip install alpha-fold-viewer
From source
git clone https://github.com/aglabx/alpha_fold_viewer.git
cd alpha_fold_viewer
pip install .
Requirements: Python 3.8+, numpy, scipy, matplotlib.
Usage
# Basic usage — generates fold_ctcf_report.html
af3-report fold_ctcf_dimer.zip
# Custom output path
af3-report fold_ctcf_dimer.zip -o ctcf_report.html
# Stricter contact distance (default: 8.0 Å)
af3-report fold_ctcf_dimer.zip --contact-dist 6.0
# Keep extracted temp files for debugging
af3-report fold_ctcf_dimer.zip --keep-tmp
# Also works as a Python script
python af3_report.py fold_ctcf_dimer.zip
CLI Reference
af3-report INPUT_ZIP [-o OUTPUT_HTML] [--contact-dist 8.0] [--keep-tmp]
Positional:
INPUT_ZIP Path to AlphaFold3 output ZIP file
Options:
-o, --output Output HTML file (default: {zip_name}_report.html)
--contact-dist Inter-atomic contact threshold in Å (default: 8.0)
--keep-tmp Keep extracted temporary files
Output Description
Confidence Overview
Models are sorted by ranking_score (highest first). The best model is highlighted in green. Columns:
| Column | Description |
|---|---|
| Ranking Score | AF3 composite confidence metric (higher = better) |
| ipTM | Interface predicted TM-score (0–1, higher = better interface) |
| pTM | Predicted TM-score for overall structure |
| Frac. Disordered | Fraction of residues predicted as disordered |
| Clash | Whether the model has steric clashes |
Chain Sequences
Each chain is displayed with:
- Full sequence colored by per-residue pLDDT (green ≥90, cyan ≥70, yellow ≥50, red <50)
- Interface residues highlighted with cyan background
- Linear heatmap strip showing pLDDT along the sequence with interface markers
Interface Analysis
For multi-chain models, inter-chain contacts are detected using a KDTree spatial search. Each interface reports:
| Metric | Description |
|---|---|
| Res. A / Res. B | Number of residues at the interface per chain |
| Atom Contacts | Total inter-chain atom pairs within contact distance |
| pLDDT A / pLDDT B | Mean predicted local confidence at interface residues |
| Avg PAE | Mean predicted aligned error across interface residue pairs |
| PAE <10Å | Percentage of PAE values below 10Å (higher = more confident) |
| High-conf | Percentage of contacts where both atoms have pLDDT ≥ 70 |
PAE Heatmaps
Each model gets a full PAE matrix heatmap with chain boundary lines. The colormap runs from dark blue (low PAE = high confidence) through green/yellow to red (high PAE = low confidence). Scale: 0–30 Å.
For multi-chain models, per-interface sub-matrices are also shown with mean PAE and <10Å percentage annotations.
How it works
- Extracts the AF3 ZIP to a temporary directory
- Auto-discovers model files (
*_model_*.cif,*_full_data_*.json,*_summary_confidences_*.json) - Parses mmCIF structures to extract atom coordinates, chain IDs, pLDDT values
- Loads PAE matrices from full_data JSONs
- Detects inter-chain interfaces using scipy KDTree
- Cross-references interfaces with PAE data
- Generates PAE heatmaps in-memory using matplotlib (→ base64 PNGs)
- Assembles everything into a single standalone HTML file
License
MIT
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file alpha_fold_viewer-0.1.2.tar.gz.
File metadata
- Download URL: alpha_fold_viewer-0.1.2.tar.gz
- Upload date:
- Size: 19.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
21e0456e5bb35aeed25989fc49c06d314ff0ee0d9e542714077f74a701a4f8a9
|
|
| MD5 |
69ac07b1aafbc524c713ab39f142c2ae
|
|
| BLAKE2b-256 |
bb534fc9925541e7d14f7e7449487a865d6a2d22768acde5e3036db534be7f6b
|
File details
Details for the file alpha_fold_viewer-0.1.2-py3-none-any.whl.
File metadata
- Download URL: alpha_fold_viewer-0.1.2-py3-none-any.whl
- Upload date:
- Size: 17.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
45c3526df68dde0445cdef215821fee5904bf5bd3bef0950d9f634f848144295
|
|
| MD5 |
d39b533a0c20c6f4136f1c444ec29929
|
|
| BLAKE2b-256 |
4bea92c0f2987bc8409db0fe8a94c32f32f04396e73a01927428dd24d07dc3eb
|