Web interface for Google DeepMind's AlphaGenome genomic prediction model
Project description

A web interface for Google DeepMind's AlphaGenome genomic prediction model.
Quick Start · Features · Deployment · License
What is this?
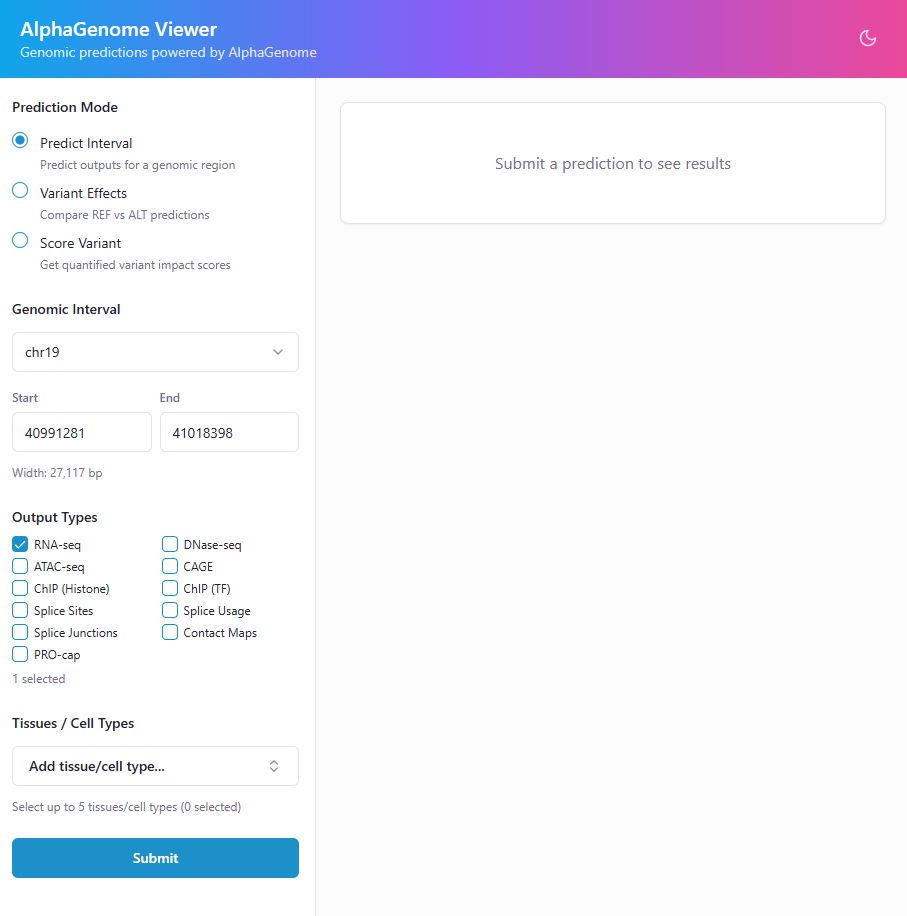
AlphaGenome Viewer gives you a point-and-click interface to query AlphaGenome predictions — no notebook or SDK boilerplate required. Enter a genomic region or variant, pick your output types and tissues, and get back publication-ready plots and scores in seconds.
Features
Interval Predictions — Visualize gene expression, chromatin accessibility, histone marks, and more across any genomic region.
Variant Effect Comparison — Overlay REF vs ALT predictions side-by-side to see exactly how a variant changes the signal.
Variant Scoring — Score variants with AlphaGenome's recommended scorers and get quantile rankings across the genome.
Supported output types
| Category | Types |
|---|---|
| Expression | RNA-seq, CAGE, PRO-cap |
| Accessibility | ATAC-seq, DNase-seq |
| Chromatin | ChIP-seq (histones), ChIP-seq (TFs), Contact maps |
| Splicing | Splice sites, Splice site usage, Splice junctions |
Quick Start
Option A: pip install (simplest)
pip install alphagenome-viewer
alphagenome-viewer
Open http://localhost:8000 and enter your API key when prompted. The command accepts --port, --host, --workers, and --plots-dir flags.
Option B: From source
Prerequisites
- Python 3.10+
- Node.js 18+
- AlphaGenome API key
Install & Run
# Clone the repo
git clone https://github.com/Abrar-Abir/alphagenome-viewer.git
cd alphagenome-viewer
# Install dependencies
cd backend && pip install -r requirements.txt && cd ..
cd frontend && npm install && cd ..
# Start both services
./start.sh
You should see output like this:
Starting AlphaGenome Viewer...
Starting backend on http://localhost:8000...
Backend started (PID 20094, logging to backend/app.log)
Starting frontend on http://localhost:5173...
Frontend started (PID 20134, logging to frontend/app.log)
Both services running. Press Ctrl+C to stop.
Backend: http://localhost:8000 (logs: backend/app.log)
Frontend: http://localhost:5173 (logs: frontend/app.log)
Open http://localhost:5173 and enter your API key when prompted. That's it.
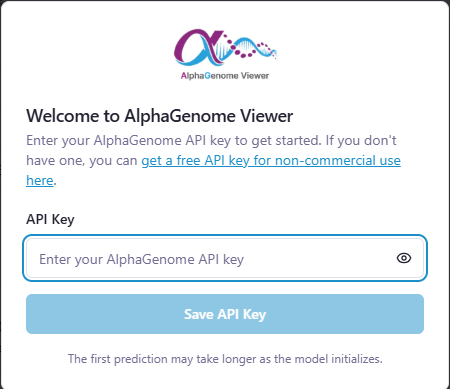
Press Ctrl+C to stop both services:
^C
Shutting down...
[2]+ Terminated npm run dev > app.log 2>&1
Shutdown complete.
You can also run the services separately — see backend/README.md and frontend/README.md for details.
Deployment
Docker
docker compose up
Open http://localhost:8000. Generated plots persist across restarts via a Docker volume. Your API key is stored in your browser's localStorage.
To build and run manually:
docker build -t alphagenome-viewer .
docker run -p 8000:8000 -v agplots:/app/data/plots alphagenome-viewer
Apptainer / Singularity
For shared servers or HPC environments, build a single container — no Node.js needed at runtime:
apptainer build alphagenome-viewer.sif alphagenome-viewer.def
mkdir -p ./data/plots
apptainer run \
--bind ./data/plots:/opt/alphagenome-viewer/data/plots \
alphagenome-viewer.sif
Open http://localhost:8000. Change the port with APPTAINERENV_AGVIEWER_PORT=9000.
See apptainer run-help alphagenome-viewer.sif for the full list of environment variables and bind mount paths.
Tech Stack
- Frontend: React, Vite, Tailwind CSS, shadcn/ui
- Backend: FastAPI, Matplotlib
- API: AlphaGenome SDK
Contributing
Contributions are welcome! Please open an issue first to discuss what you'd like to change.
License
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file alphagenome_viewer-1.0.0.tar.gz.
File metadata
- Download URL: alphagenome_viewer-1.0.0.tar.gz
- Upload date:
- Size: 766.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
05e2a62bffa9622384649f5df5f89ecb763b1d2a32cc2a78b2001ed378103a3f
|
|
| MD5 |
52773e02625783fe47af43140c4a73be
|
|
| BLAKE2b-256 |
e5036a410025de88be7ebf90e825062f5bc9efeccd0442f1bef0e645a88eae38
|
File details
Details for the file alphagenome_viewer-1.0.0-py3-none-any.whl.
File metadata
- Download URL: alphagenome_viewer-1.0.0-py3-none-any.whl
- Upload date:
- Size: 770.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d7dfd9ac65198938563113398b6b8290bc2c0dd9fdb4334fa9f49cd9bc09227a
|
|
| MD5 |
38744a85e84267e9f4069bbd83f0d022
|
|
| BLAKE2b-256 |
b927a2aa5652f98964a59db4b188fe6889d30e7c79555b5c22de4a86e62b80c8
|











