Adaptive Mixture ICA in Python
Project description
AMICA-Python
Yes, it's fast.
A Python implementation of the AMICA (Adaptive Mixture Independent Component Analysis) algorithm for blind source separation, that was originally developed in FORTRAN by Jason Palmer at the Swartz Center for Computational Neuroscience (SCCN).
AMICA-Python is pre-alpha but is tested against the Fortran implementation and is ready for test driving.
| Python | Fortran |
|---|---|
 |
 |
Installation
For now, AMICA-Python should be installed from source, and you will have to manually install PyTorch (see below) yourself:
git clone https://github.com/scott-huberty/amica-python.git
cd amica-python
pip install -e .
[!IMPORTANT] You must install PyTorch before using AMICA-Python.
Installing PyTorch
Depending on your system and preferences, you can install PyTorch with or without GPU support.
To install the standard version of PyTorch, run:
python -m pip install torch
To install the CPU-only version of PyTorch, run:
python -m pip install torch --index-url https://download.pytorch.org/whl/cu113
Or for Conda users:
conda install -c conda-forge pytorch-cpu
[!WARNING] If you are using an Intel Mac, you cannot install Pytorch via pip, because there are no precompiled wheels for that platform. Instead, you must install PyTorch via Conda, e.g.:
conda install pytorch -c conda-forge
If you use UV, you can also just install torch while installing AMICA-Python:
uv pip install -e ".[torch-cpu]"
uv pip install -e ".[torch-cuda]"
Usage
AMICA-Python exposes a scikit-learn style interface. Here is an example of how to use it:
import numpy as np
from scipy import signal
from amica import AMICA
rng = np.random.default_rng(0)
n_samples = 2000
time = np.linspace(0, 8, n_samples)
s1 = np.sin(2 * time) # Sinusoidal
s2 = np.sign(np.sin(3 * time)) # Square wave
s3 = signal.sawtooth(2 * np.pi * time) # Sawtooth
S = np.c_[s1, s2, s3]
S += 0.2 * rng.standard_normal(S.shape) # Add noise
S /= S.std(axis=0) # Standardize
A = np.array([[1, 1, 1],
[0.5, 2, 1.0],
[1.5, 1.0, 2.0]]) # Mixing matrix
X = S @ A.T # Observed mixtures
ica = AMICA(random_state=0)
X_new = ica.fit_transform(X)
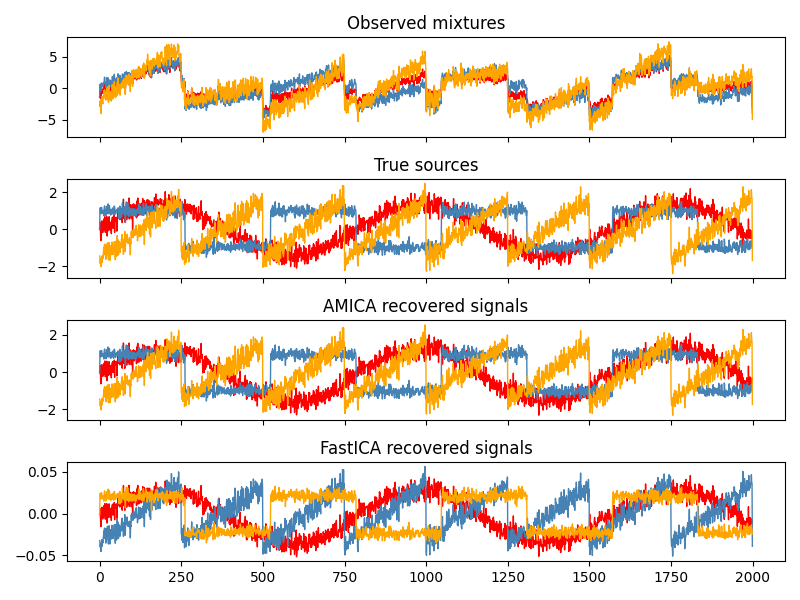
GPU acceleration
If PyTorch was installed with CUDA support, you can fit AMICA on GPU:
ica = AMICA(device='cuda', random_state=0)
For more examples and documentation, please see the documentation.
What is AMICA?
AMICA is composed of two main ideas, which are hinted at by the name and the title of the original paper: AMICA: An Adaptive Mixture of Independent Component Analyzers with Shared Components.
1. Adaptive Mixture ICA
Standard ICA assumes each source is independent and non-Gaussian. Extended Infomax ICA improves on this by handling both sub-Gaussian and super-Gaussian sources. AMICA goes further by modeling each source as a mixture of multiple Gaussians. This flexibility lets AMICA represent virtually any source shape - super-Gaussian, sub-Gaussian, or even some funky bimodal distribution:

In practice, the authors argue that this leads to a more accurate approximation of the source signals.
2. Shared Components
AMICA can learn multiple ICA decompositions (i.e. models). This is a work around to the assumption of ICA that the sources are stationary (they do not change over time). AMICA will decide which model best explains the data at each sample, effectively allowing the sources to change over time. The "shared components" part of the paper title refers to AMICA's ability to allow the various ICA models to share some components (i.e. sources) between them, to reduce computational load.
What does AMICA-Python implement?
In short, AMICA-Python implements point 1 above (Adaptive Mixture ICA), but does not implement point 2 (running multiple ICA models simultaneously).
AMICA-Python is powered by Torch and wrapped in an easy-to-use scikit-learn style interface.
The outputs are numerically tested against the original FORTRAN implementation to ensure correctness and minimize bugs.
What wasn't implemented?
- The ability to model multiple ICA decompositions simultaneously.
- The ability to reject unlikely samples based on a thresholded log-likelihood (in the FORTRAN implementation, this is a strategy to deal with artifacts in the data).
- AMICA-Python does not expose all the hyper-parameters available in the original FORTRAN implementation. Instead I have tried to pick sensible defaults that should work well in most cases, thus reducing the complexity of the interface.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file amica_python-0.1.0.tar.gz.
File metadata
- Download URL: amica_python-0.1.0.tar.gz
- Upload date:
- Size: 61.8 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b8eb17dc978e3d3b800c2cecd3ef0849849c1ce441caad9eb6d7ef1359e43d52
|
|
| MD5 |
199379528ff66daf16039a1076d4e237
|
|
| BLAKE2b-256 |
d33feb1f98f26e48b96422a1e1839ee9c22e2095773cf2b0cf7b177568941822
|
File details
Details for the file amica_python-0.1.0-py3-none-any.whl.
File metadata
- Download URL: amica_python-0.1.0-py3-none-any.whl
- Upload date:
- Size: 66.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.16
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
59ffde9a8893e065cd01a0b09dd0833d0f173aeb54b989af5563bc3258e521f6
|
|
| MD5 |
ce459e9a64250b7d8a940e80d6683842
|
|
| BLAKE2b-256 |
85ff464b32a75a375e7213068089a3ddd639e45ff01b6231a836e7622bddaf31
|















