script to generate amplicon coverage plot
Project description
amplicon_coverage_plot
The script will generate an interactive barplot given amplicon info in bed6/bedpe format and coverage information in cov/bam file.
Dependencies
Programming/Scripting languages
- Python >=v3.8
- The pipeline has been tested in v3.8.10
Python packages
Third party softwares/packages
- samtools >=1.9 - process bam file
Installation
Install by pip
pip install amplicov
Install by conda
conda install -c bioconda amplicon_coverage_plot
Install from source
Clone the amplicon_coverage_plot repository.
git clone https://github.com/chienchi/amplicon_coverage_plot
Then change directory to amplicon_coverage_plot and install.
cd amplicon_coverage_plot
python setup.py install
If the installation was succesful, you should be able to type amplicov -h and get a help message on how to use the tool.
amplicov -h
Usage
usage: amplicov [-h] (--bed [FILE] | --bedpe [FILE])
(--bam [FILE] | --cov [FILE]) [-o [PATH]] [-p [STR]] [--pp]
[--mincov [INT]] [--version]
Script to parse amplicon region coverage and generate barplot in html
optional arguments:
-h, --help show this help message and exit
--pp process proper paired only reads from bam file
(illumina)
--count_primer count overlapped primer region to unqiue coverage
--mincov [INT] minimum coverage to count as ambiguous N site
[default:10]
-r [STR], --refID [STR]
reference accession (bed file first field)
--depth_lines DEPTH_LINES [DEPTH_LINES ...]
Add option to display lines at these depths (provide depths as a list of integers) [default:5 10 20 50]
--gff [FILE] gff file for data hover info annotation
--version show program's version number and exit
Amplicon Input (required, mutually exclusive):
--bed [FILE] amplicon bed file (bed6 format)
--bedpe [FILE] amplicon bedpe file
Coverage Input (required, mutually exclusive):
--bam [FILE] sorted bam file (ex: samtools sort input.bam -o
sorted.bam)
--cov [FILE] coverage file [position coverage]
Output:
-o [PATH], --outdir [PATH]
output directory
-p [STR], --prefix [STR]
output prefix
Test
cd tests
./runTest.sh
Outputs
-- prefix_amplicon_coverage.txt
| ID | Whole_Amplicon | Unique | Whole_Amplicon_Ns(cov<10) | Unique_Amplicon_Ns(cov<10) |
|---|---|---|---|---|
| nCoV-2019_1 | 217.74 | 53.00 | 0.00 | 0.00 |
| nCoV-2019_2 | 1552.83 | 1235.50 | 0.00 | 0.00 |
| nCoV-2019_3 | 3164.22 | 2831.73 | 0.00 | 0.00 |
| nCoV-2019_4 | 2005.16 | 1658.00 | 0.00 | 0.00 |
| etc... |
Table Header Definition in the amplicon_coverage.txt
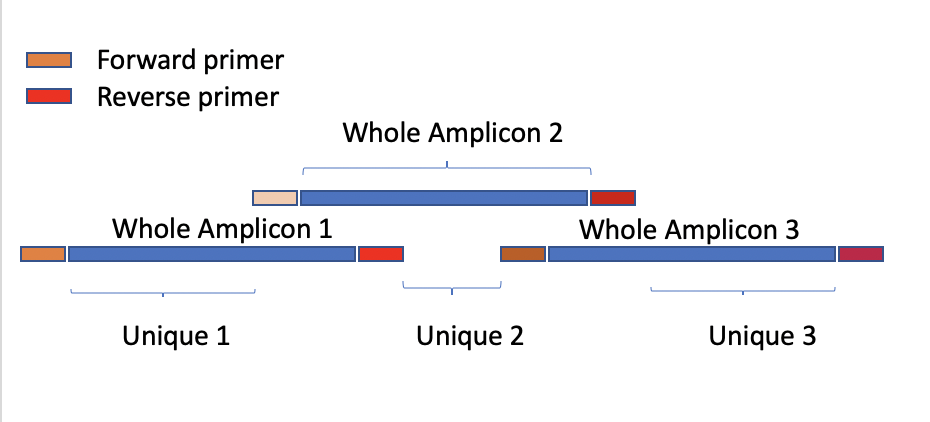
-
Whole_Amplicon_Ns(cov<10): The number of aligned position with coverage < 10 or (--mincov) in the Whole Amplicon region
-
Unique_Amplicon_Ns(cov<10): The number of aligned position with coverage < 10 or (--mincov) in the Unique region
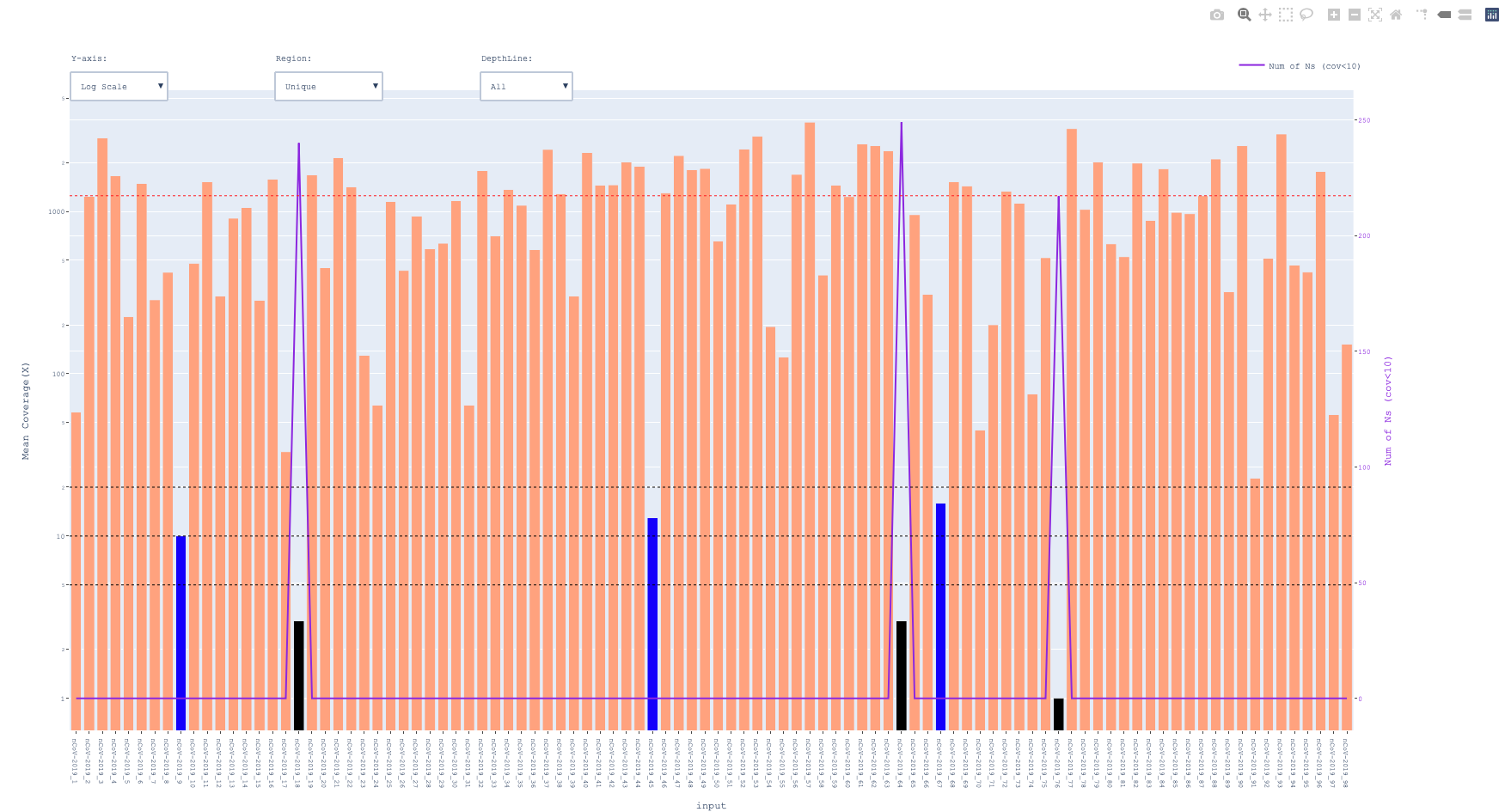
-- prefix_amplicon_coverage.html
color black for < 5x and blue for <20x
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file amplicov-0.3.4-py3-none-any.whl.
File metadata
- Download URL: amplicov-0.3.4-py3-none-any.whl
- Upload date:
- Size: 20.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.0.1 CPython/3.12.4
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1c787bb4a536e75bfc4ccbf30c62094420a527ba033070631e477b055017a8e4
|
|
| MD5 |
082d7be3c312bd91d69591ca34df1b69
|
|
| BLAKE2b-256 |
057d5306aff2d071ab644b6e7f0147527a63d35cd0399b328133663cab848f90
|