Lightweight molecular structure viewer for Jupyter notebooks using py3Dmol
Project description
atomview
A lightweight, standalone molecular structure viewer for Jupyter notebooks. Visualize biotite AtomArray structures interactively using py3Dmol (3Dmol.js) — with chain-coloured cartoons, ligand sticks, ion spheres, surfaces, and hover labels out of the box.
atomview extracts and refactors the excellent view() function from atomworks (atomworks.io.utils.visualize) into a focused, minimal package. If all you need is a quick way to render structures in a notebook, you no longer have to pull in atomworks and its full dependency tree (RDKit, ML tooling, dataset infrastructure, etc.).
Installation
pip install atomview
Or with uv:
uv add atomview
Quickstart
# from urllib.request import urlretrieve
# urlretrieve("https://files.rcsb.org/download/4HHB.cif", "4hhb.cif")
from atomview import load_structure, view
structure = load_structure("4hhb.cif")
view(structure)
Gallery
| Default protein view | Surface overlay |
|---|---|
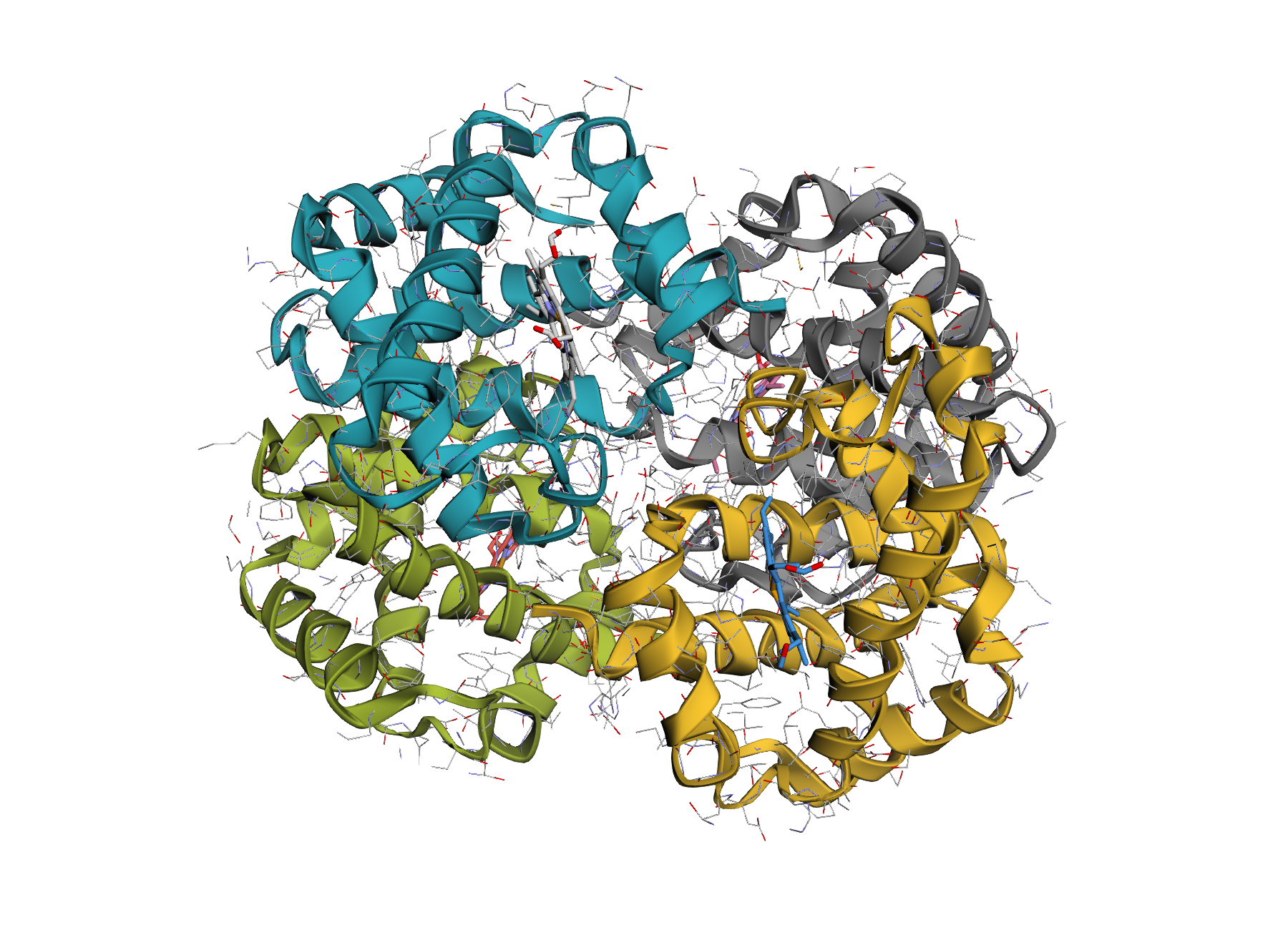 |
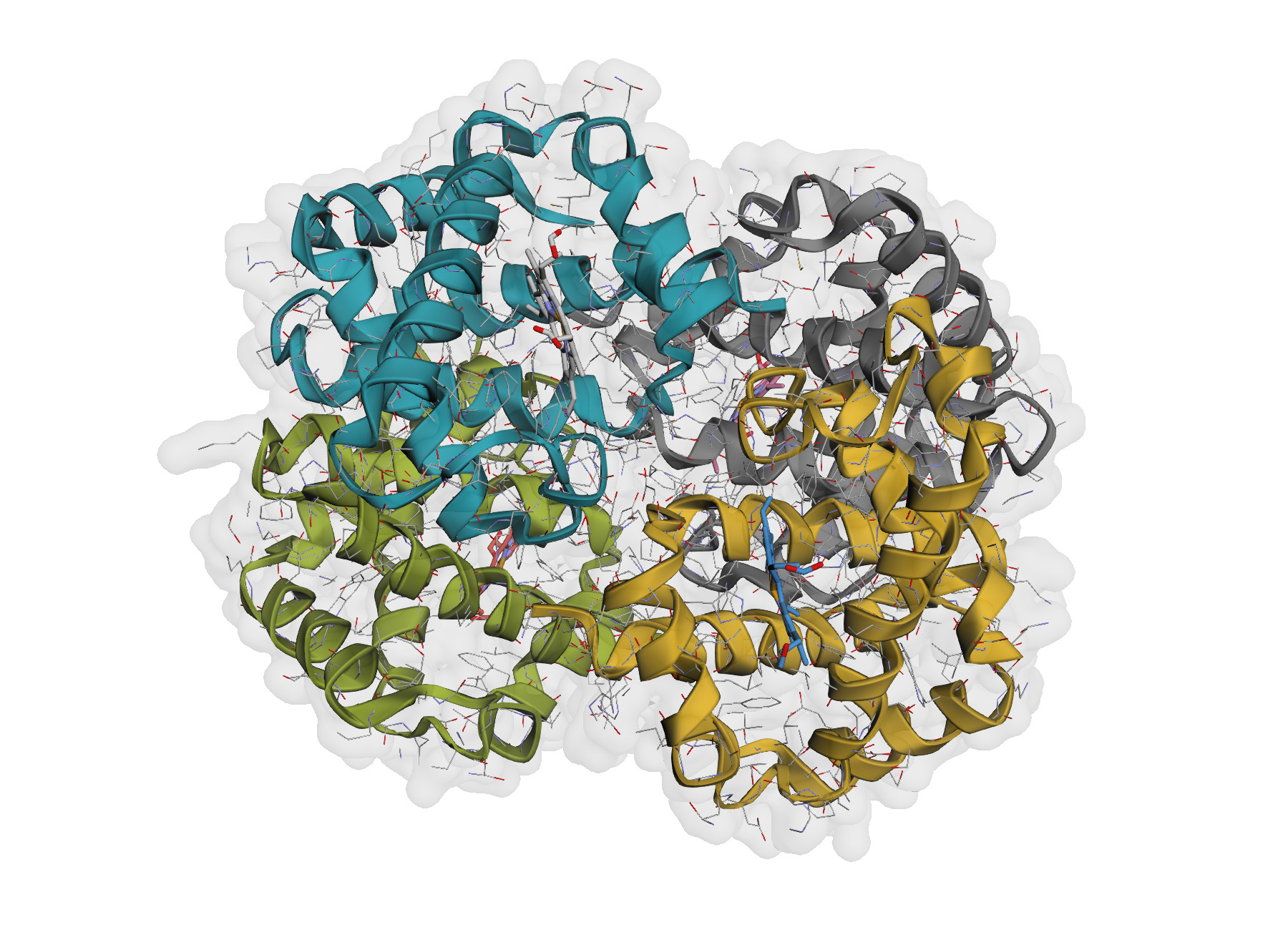 |
| One-line rendering with automatic chain colouring. | Optional translucent VDW surface for shape context. |
| Ligand focus | Mixed complex |
|---|---|
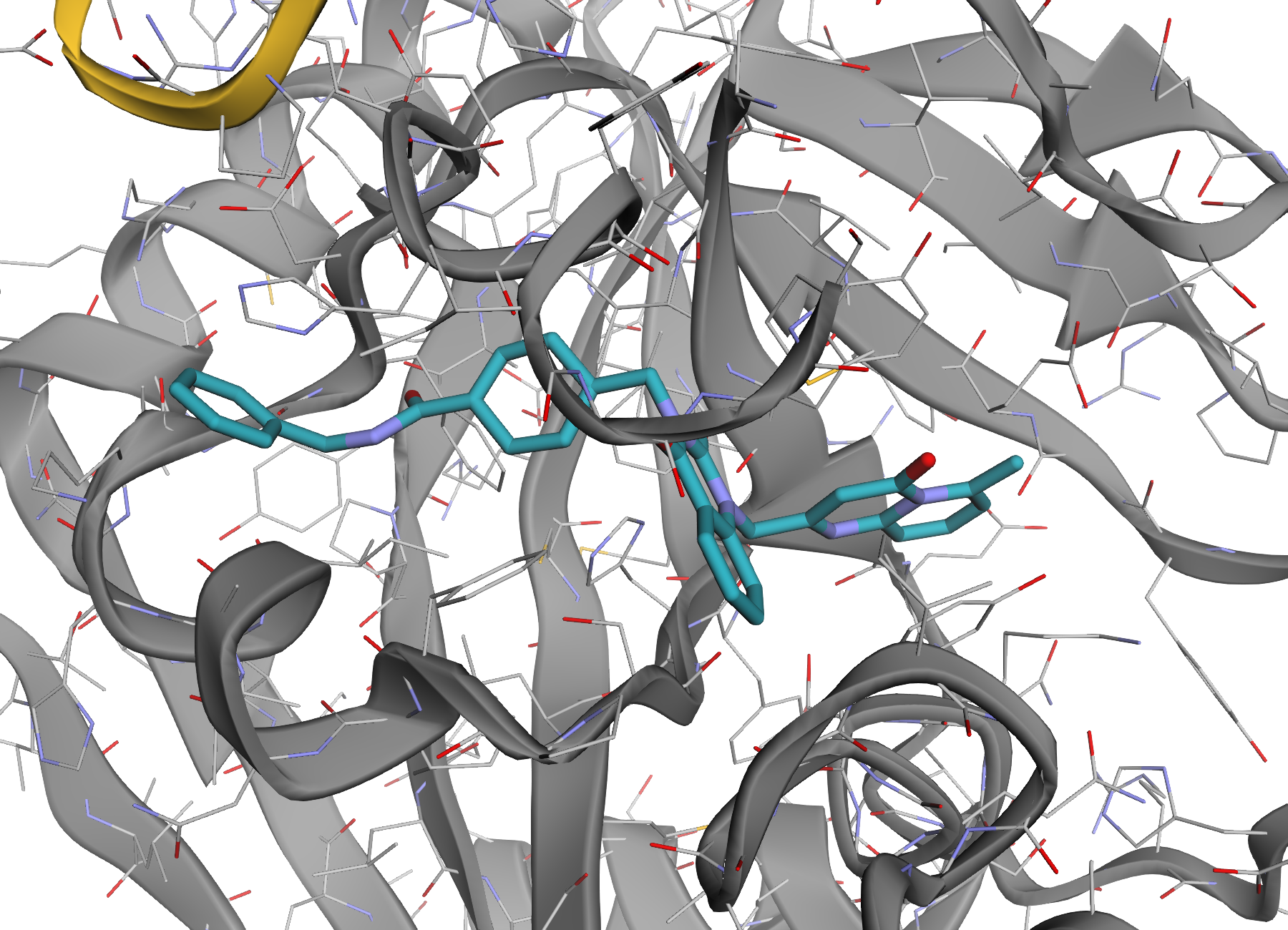 |
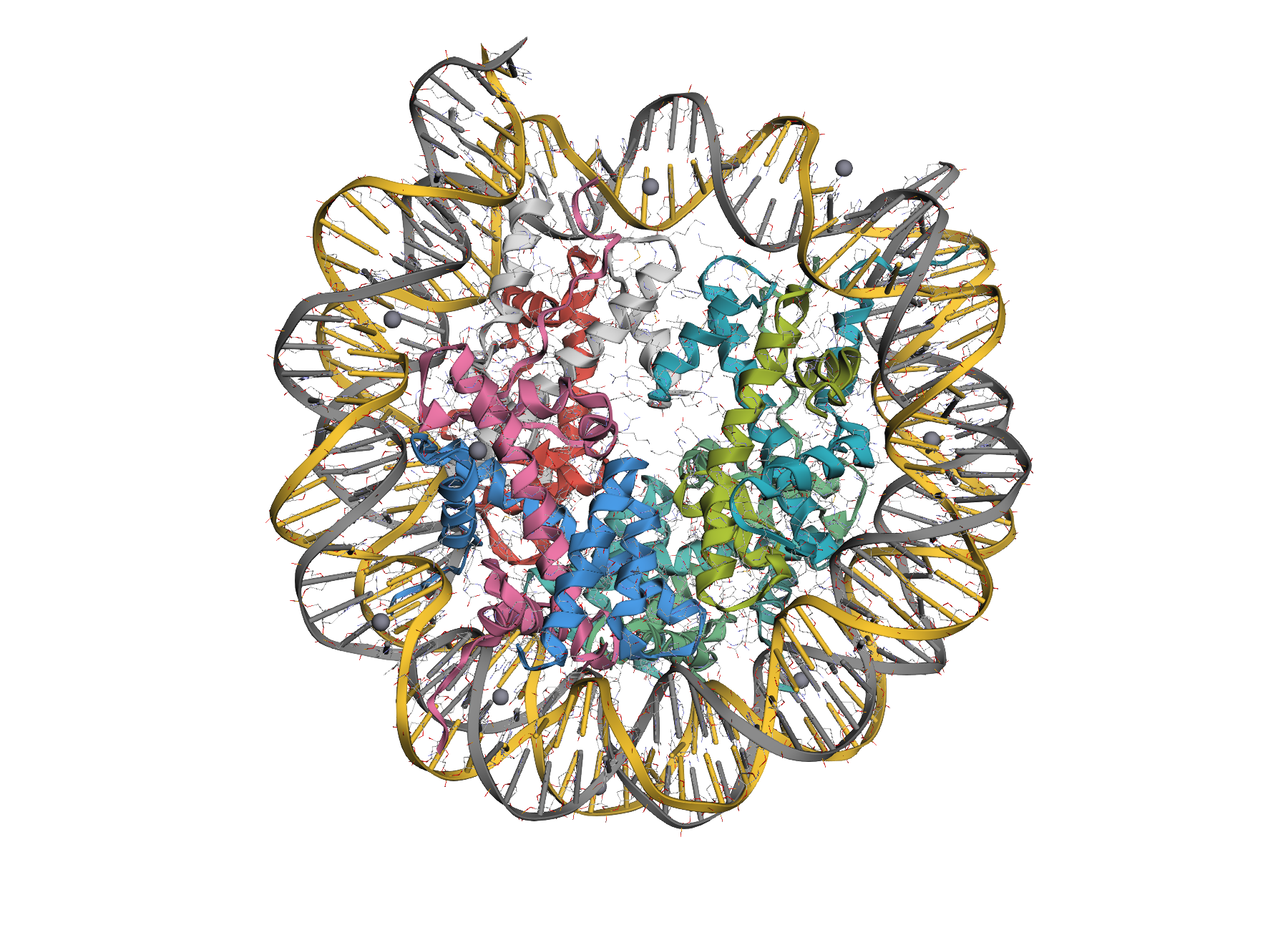 |
| Zoom to a ligand or binding-site selection. | Proteins, nucleic acids, ligands, and ions styled together. |
API
view()
The main entry point. Returns a py3Dmol.view object that renders inline in Jupyter.
from atomview import view
v = view(
structure,
show_cartoon=True, # cartoon for polymers
show_surface=False, # VDW surface overlay
show_hover=True, # hover labels
hide_solvent=True, # remove water
hide_crystallization_aids=True,
zoom_to_selection=None, # e.g. {"chain": "A", "resi": 35}
width=600,
height=400,
colors=None, # custom colour list, or use defaults
)
Why not just use atomworks?
atomworks is a fantastic, comprehensive framework for biomolecular modelling maintained by RosettaCommons. But it's a big library — it bundles I/O, transforms, ML dataset tooling, RDKit integration, and more. If you just want to call view(structure) in a notebook, that's a lot of overhead.
atomview gives you exactly that one thing: a great molecular viewer with sensible defaults, in a package that installs in seconds.
License
MIT
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file atomview-0.1.1.tar.gz.
File metadata
- Download URL: atomview-0.1.1.tar.gz
- Upload date:
- Size: 8.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
e0a0a83535b35a5eb6201f4e29940cd99ac7de7ef02003351b900a27a3ab3852
|
|
| MD5 |
08c77c2459564fb9c44352ee733c48cd
|
|
| BLAKE2b-256 |
bbf637a434199d192af1a5128f7c2f1a313eec517609bc0e6026c4c9e313cb6b
|
Provenance
The following attestation bundles were made for atomview-0.1.1.tar.gz:
Publisher:
deploy.yml on cch1999/atomview
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
atomview-0.1.1.tar.gz -
Subject digest:
e0a0a83535b35a5eb6201f4e29940cd99ac7de7ef02003351b900a27a3ab3852 - Sigstore transparency entry: 1435657267
- Sigstore integration time:
-
Permalink:
cch1999/atomview@3631b84ae4e7264f54972069872ad0ddb02eb4ab -
Branch / Tag:
refs/heads/main - Owner: https://github.com/cch1999
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
deploy.yml@3631b84ae4e7264f54972069872ad0ddb02eb4ab -
Trigger Event:
push
-
Statement type:
File details
Details for the file atomview-0.1.1-py3-none-any.whl.
File metadata
- Download URL: atomview-0.1.1-py3-none-any.whl
- Upload date:
- Size: 10.3 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
f573739e311d90da4517d68e4e807a7e2a295833c8c54296d1b66bd84634f982
|
|
| MD5 |
a3dea7b1eb54fe9a5bf4a8f85178738d
|
|
| BLAKE2b-256 |
cdaaf2c1de4ca835add4872faa589504944c9bd09c0842b729ac693b4fa7979e
|
Provenance
The following attestation bundles were made for atomview-0.1.1-py3-none-any.whl:
Publisher:
deploy.yml on cch1999/atomview
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
atomview-0.1.1-py3-none-any.whl -
Subject digest:
f573739e311d90da4517d68e4e807a7e2a295833c8c54296d1b66bd84634f982 - Sigstore transparency entry: 1435657314
- Sigstore integration time:
-
Permalink:
cch1999/atomview@3631b84ae4e7264f54972069872ad0ddb02eb4ab -
Branch / Tag:
refs/heads/main - Owner: https://github.com/cch1999
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
deploy.yml@3631b84ae4e7264f54972069872ad0ddb02eb4ab -
Trigger Event:
push
-
Statement type:

















