wrapper for brain segmentation tools
Project description
BrainSeg-container
This repository provides a streamlined Python wrapper and CLI tool to automatically download, run, and standardize outputs from state-of-the-art brain segmentation tools using Apptainer / Singularity containers.
Brain segmentation tools often have conflicting dependencies, complex installation steps, or require specific versions of system libraries. This package solves that problem by containerizing the tools and handling the execution, file binding, and label standardization for you.
The Tools
This pipeline currently supports the following deep-learning-based segmentation tools. We may add more in the future.
-
GOUHFI This tool was designed to handle the challenges of Ultra-High Field MRI (7T+). It utilizes "domain randomization" during training, which allows it to remain robust across different MRI contrasts and resolutions, including standard clinical scans.
-
Resolution: Native (preserves input resolution).
-
CSF Availability: Yes (segments ventricles and subarachnoid space CSF).
-
-
SynthSeg Developed by the FreeSurfer team, this tool is famous for working "out of the box" on almost any kind of MRI scan (different contrasts, resolutions, or messy clinical data) without needing retraining.
-
Resolution: Fixed 1mm isotropic (always resamples input to 1mm).
-
CSF Availability: Yes.
-
-
FastSurfer A rapid deep-learning-based segmentation tool that is optimized for speed, finishing in minutes rather than hours.
-
Resolution: Native (but experimental below 0.7mm).
-
CSF Availability: No (segments ventricles, but ignores subarachnoid space CSF).
-
-
SimNIBS (Charm) The "Complete Head Anatomy Reconstruction Method" from the SimNIBS suite. While designed for modeling brain stimulation (TMS/TES), it produces high-quality segmentation of extra-cerebral tissues (skull, scalp, etc.) in addition to the brain.
-
Resolution: Native (pipeline uses the upsampled output to match input).
-
CSF Availability: Yes.
-
Segmented regions: Charm provides the following segmentation labels: White-Matter, Gray-Matter, CSF, Bone, Scalp, Eye_bals, Compact_bone, Spongy_bone, Blood, Muscle, Cartilage, Fat, Electrode, Saline_or_gel
-
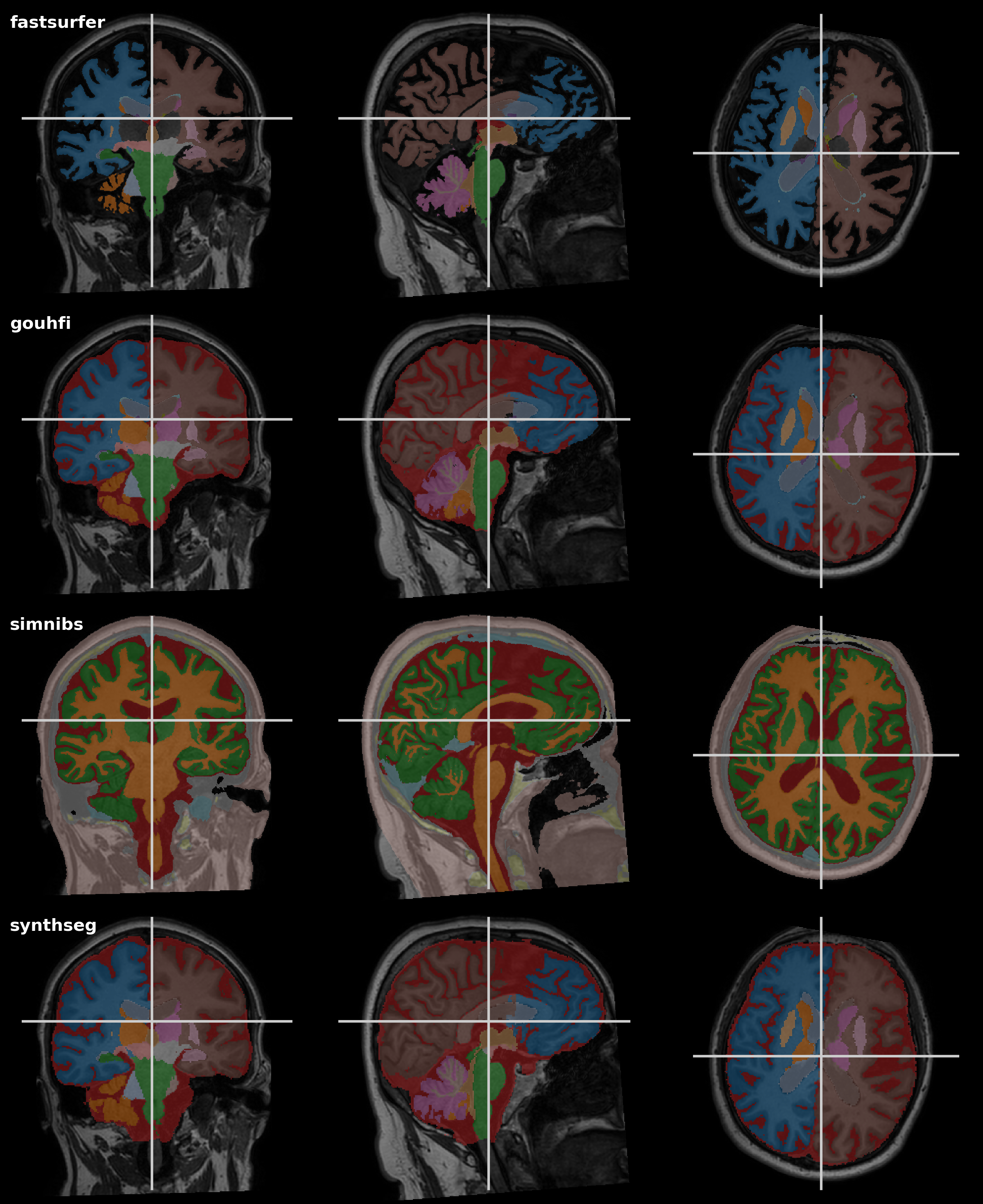
Comparison Output
The pipeline can automatically generate a comparison grid so you can quickly inspect the differences between the tools.
Getting Started
Prerequisites
You must have Apptainer (or Singularity) installed on your system to run the containers.
- Ubuntu/Debian:
sudo apt install apptainer - Conda/Mamba:
conda install -c conda-forge apptainer
Installation
You can install the package directly via pip:
pip install brainseg-containers
(Optional) If you want to use the plotting and comparison features, install with the plot extras:
pip install brainseg-containers[plot]
Usage
The package provides a simple command-line interface. The first time you run a specific tool, the wrapper will automatically download the corresponding container from the GitHub Container Registry and store it in ~/.brainseg_containers/.
Basic Command
brainseg-containers -t <tool_name> -i <input_file.nii.gz> -o <output_file.nii.gz>
Available Tools: synthseg, gouhfi, fastsurfer, simnibs
Examples
Run GOUHFI on a single subject:
brainseg-containers -t gouhfi -i inputs/sub-01_T1w.nii.gz -o results/sub-01_gouhfi.nii.gz
Run SynthSeg on the same subject:
brainseg-containers -t synthseg -i inputs/sub-01_T1w.nii.gz -o results/sub-01_synthseg.nii.gz
Note: You can optionally provide a custom path to a pre-downloaded .sif image using the --container flag.
Note on Labels
Different tools use different numbers to represent brain regions. To make comparison easier, this pipeline automatically remaps the output labels of FastSurfer and GOUHFI to match the standard FreeSurfer lookup table.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file brainseg_containers-1.0.0.dev0.tar.gz.
File metadata
- Download URL: brainseg_containers-1.0.0.dev0.tar.gz
- Upload date:
- Size: 28.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
bc03950b81a024ce7a4af38d701903dbc99205c0684ff6fde3365c90063af484
|
|
| MD5 |
07fab57399682eccebafffbc54a5bd7c
|
|
| BLAKE2b-256 |
9cb6d13ea0e66e8a9801a7e67b6bb7330661d8925940fdebc97af27b41f0257a
|
File details
Details for the file brainseg_containers-1.0.0.dev0-py3-none-any.whl.
File metadata
- Download URL: brainseg_containers-1.0.0.dev0-py3-none-any.whl
- Upload date:
- Size: 29.7 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a919a22ddea17336aa1acaff2dc17542c80121568de0c17d229832a02fac46f0
|
|
| MD5 |
3e578a1a6cec3a6ab619f07248451e18
|
|
| BLAKE2b-256 |
e2cd2298b885f2f419b9ab5ea7c75dac22a325626cf8df9a44f3830d55c30530
|