A package to extract the causal graph from continuous tabular data.
Project description
CausalExplain - A library to infer causal-effect relationships from tabular data
'CausalExplain' is a library that implements methods to extract the causal graph, from tabular data, specifically the ReX method, and other compared methods like GES, PC, FCI, LiNGAM, CAM, and NOTEARS. At present, the public, supported path is centered on ReX and the more complete comparison methods; the PC and CAM implementations remain in the repository for research/reference purposes but are not supported as production-ready public APIs.
This repository contains the implementation of ReX and all necessary tools to reproduce the results presented in our accompanying paper. ReX supports diverse data generation processes, including non-linear and additive noise models, and has demonstrated robust performance on synthetic and real-world datasets.
About ReX
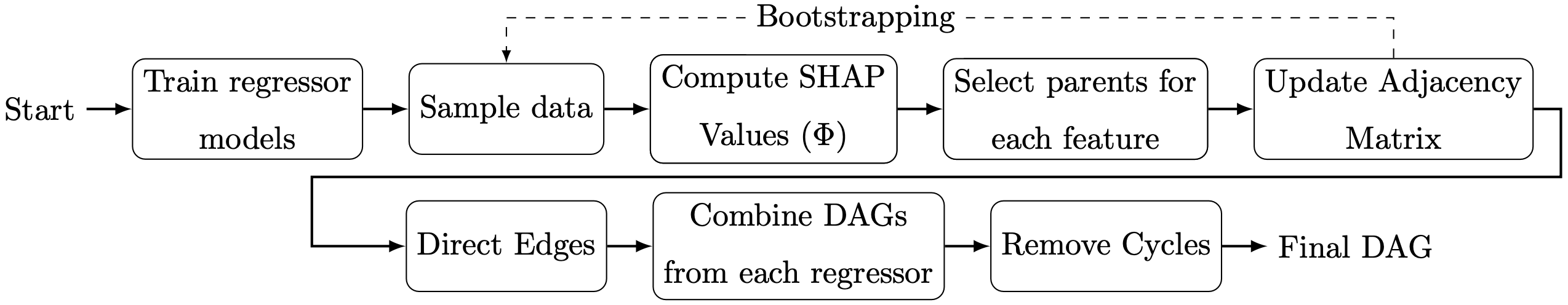
ReX is a causal discovery method that leverages machine learning (ML) models coupled with explainability techniques, specifically Shapley values, to identify and interpret significant causal relationships among variables. Comparative evaluations on synthetic datasets comprising tabular data reveal that ReX outperforms state-of-the-art causal discovery methods across diverse data generation processes, including non-linear and additive noise models. Moreover, ReX was tested on the Sachs single-cell protein-signaling dataset, achieving a precision of 0.952 and recovering key causal relationships with no incorrect edges. Taking together, these results showcase ReX's effectiveness in accurately recovering true causal structures while minimizing false positive predictions, its robustness across diverse datasets, and its applicability to real-world problems. By combining ML and explainability techniques with causal discovery, ReX bridges the gap between predictive modeling and causal inference, offering an effective tool for understanding complex causal structures.
Our experimental results, conducted on five families of synthetic datasets with varying complexity, demonstrate that REX consistently recovers true causal relationships with high precision while minimizing false positives and orientation errors, comparing favorably to existing methods. Additionally, REX was tested on the Sachs single-cell protein-signaling dataset (Sachs et al., 2005), achieving a competitive performance with no false positives and recovering important causal relationships. This further validates the applicability of REX to real-world datasets, highlighting its robustness across different types of data.
Prerequisites without Docker
- Operating System: Linux or macOS
- Environment Manager: PyEnv or Conda
- Programming Language: Python 3.10+
- Hardware: CPU (CUDA/MPS optional)
Installation
The project can be installed using pip:
$ pip install causalexplain
This installs the package together with its core runtime dependencies for the CLI, plotting, and bundled causal-discovery methods.
What's new in v0.9.4
- GUI: the Train tab now mirrors the weighted progress tracked by the ReX training pipeline, so the progress legend shows the current phase and the bar stays purely visual.
- Release hygiene: version references, changelogs, and citation metadata were synchronized for this patch release.
Data
The datasets used in the paper and the examples can be generated using the
generators module, which is also part of this library. In case you want to
reproduce results from the articles that we used as reference, you can find
the datasets in the data folder.
Executing causalexplain
Option 1: Command Line
After installation, you can use either the installed causalexplain command
or the module entry point:
$ causalexplain --help
$ python -m causalexplain --help
___ _ _ _
/ __\__ _ _ _ ___ __ _| | _____ ___ __ | | __ _(_)_ __
/ / / _` | | | / __|/ _` | |/ _ \ \/ / '_ \| |/ _` | | '_ \
/ /__| (_| | |_| \__ \ (_| | | __/> <| |_) | | (_| | | | | |
\____/\__,_|\__,_|___/\__,_|_|\___/_/\_\ .__/|_|\__,_|_|_| |_|
|_|
usage: causalexplain [-h] {run,generate,gui} ...
The top-level help lists the available subcommands. Use
causalexplain run --help, causalexplain generate --help, and
causalexplain gui --help for mode-specific options.
The minimum required to run causalexplain run is a dataset file in CSV
format, with the first row containing the names of the variables, and the rest
of the rows containing the values of the variables. The method selected by
default is ReX, but you can also choose between PC, FCI, GES, LiNGAM, CAM, and
NOTEARS. At the end of the execution, the edges of the plausible causal graph
will be displayed along with the metrics obtained, if the true DAG is provided
(argument -t).
PC and CAM are still exposed in the CLI for reproducibility and internal comparison, but they are currently unsupported: parts of their helper API are unfinished, and they should not be treated as stable public interfaces.
Generate synthetic data from the CLI
The CLI can also generate a synthetic dataset and save both the .csv data
file and the .dot ground-truth DAG from a single output base path:
$ python -m causalexplain generate \
--mechanism linear \
--variables 10 \
--samples 500 \
--output /path/to/generated/toy_dataset
This writes /path/to/generated/toy_dataset.csv and
/path/to/generated/toy_dataset.dot.
The required arguments for generation mode are:
--mechanism--variables--samples--output
The remaining generation controls default to the same values used by the GUI:
--timeout 30, --max-retries 50, --min-edges 0, --max-edges 30,
--max-parents 3, --seed 1234, and --rescale.
GUI mode
To use the local GUI, run:
$ python -m causalexplain gui
This launches a browser-based app for training models, loading/evaluating saved runs, and generating synthetic datasets, all on your local machine (port 8080).
Option 2: Notebook
In case you want to run causalexplain from your code in a notebook, you can
use the GraphDiscovery class. The following example shows how to use
the GraphDiscovery class to train a model on a dataset using ReX method:
Note: If the notebook kernel cannot import causalexplain, run the notebook
from the repo root, or install the package (pip install -e .), or add the
repo root to sys.path (e.g.: sys.path.insert(0, str(pathlib.Path("..").resolve())) ).
For higher-quality math text in plots, install a LaTeX distribution; otherwise
pass usetex=False when plotting.
See examples/simple_experiment.ipynb for a working notebook example.
from causalexplain import GraphDiscovery
experiment = GraphDiscovery(
experiment_name='toy_experiment',
model_type='rex',
csv_filename='../data/toy_dataset.csv',
true_dag_filename='../data/toy_dataset.dot')
# Run the experiments
experiment.run(hpo_iterations=10, bootstrap_iterations=10, combine_op='union', quiet=True)
# Plot the resulting DAG (avoid LaTeX/Graphviz dependencies when running locally)
experiment.plot(show_metrics=True, layout='circular', usetex=False)
To load a model from a file, you can use the load method of the
GraphDiscovery class:
from causalexplain import GraphDiscovery
experiment = GraphDiscovery()
experiment.load("/path/to/model.pkl")
Adaptive SHAP sampling
For direct SHAP usage in notebooks, the explainability module exposes a high-level wrapper that defaults to adaptive sampling:
from causalexplain.explainability.shapley import compute_shap
# default: adaptive_shap_sampling=True
res, diag = compute_shap(X, model, backend="kernel", adaptive_shap_sampling=True)
# disable (may be slow for large m)
res, diag = compute_shap(X, model, backend="kernel", adaptive_shap_sampling=False)
CLI example (same executable shown above):
python -m causalexplain run --shap-sampling
python -m causalexplain run --no-shap-sampling
For GBT-based ReX runs, -gbt-optimization controls whether per-target
feature matrices are cached to reduce repeated slicing. Use
--no-gbt-optimization to disable and lower memory usage (disabled by default).
To speed up hyperparameter tuning, use --hpo-optimization to enable Optuna
pruning and a downsampled HPO objective. You can cap rows with
--hpo-optimization-limit (disabled by default).
Available SHAP backends are kernel, gradient, explainer, and tree.
ReX defaults to tree when running the GBT regressor.
When adaptive sampling is enabled, the key knob is the SHAP optimization
limit (--shap-optimization-limit, Python: shap_budget). It controls both
SHAP background size and the number of rows explained; omit it to disable the
limit. The legacy max_shap_samples name is deprecated.
Note on large datasets: if adaptive_shap_sampling=False and m > 2000, the
tool warns about potential non-termination (the threshold is conservative).
Why adaptive sampling is mathematically reasonable
Many SHAP explainers approximate an expectation over a background distribution; using $n$ background points gives a Monte Carlo estimate. The standard error scales approximately as $SE \sim \frac{1}{\sqrt{n}}$. So, when sampling without replacement from a finite dataset of size $m$, the finite population correction factor applies: $\frac{1}{\sqrt{1 - (n/m)}}$.
This means increasing $n$ yields diminishing returns, so capping the background around 250 is a pragmatic speed/accuracy tradeoff. Repeating the sampling ($K$ runs) provides a stability diagnostic: compute a global importance vector per run as $\overline{\mid \text{SHAP} \mid}$ per feature, then check variability (CV) and rank stability (Spearman correlation) across runs.
Backend-aware note: Kernel SHAP is particularly sensitive and expensive, so
caps like max_explain_samples matter most there. Gradient and generic
explainers often have different performance profiles, but still benefit from
controlled baselines/background sizes.
This can be useful if you want to train a model on a dataset and then use it to predict causal graphs on other datasets, or train a model on different batches.
Once a model has been trained or loaded, you can plot the resulting DAG, save the trained model to a file, or export the predicted causal graph to a DOT file.
# Plot the resulting DAG
experiment.plot(show_metrics=True, layout='circular', usetex=False)
# Save the trained model to a file
experiment.save("/path/to/model.pkl")
To export the predicted causal graph to a DOT file, you can use the export
method of the GraphDiscovery class:
experiment.export("/path/to/my_predicted_graph.dot")
Output
The output of causalexplain is typically a graph with the edges of the
plausible causal graph and the metrics obtained from the evaluation of the
causal graph against the true DAG. These results are printed to the console,
unless the '-o' option is specified, in which case the DAG is saved to a
file in DOT format. Metrics are printed only if the true DAG is provided.
Example CLI commands
The following command illustrates how to run causalexplain on the toy dataset
using the ReX method:
$ python -m causalexplain run -d /path/to/toy_dataset.csv -t /path/to/toy_dataset.dot
The CLI still exposes -m pc and -m cam for research/reference workflows,
but those two methods are currently unsupported and are not considered
release-ready public interfaces.
For more information on command line options, run causalexplain -h or go to
the Quickstart
section in the documentation.
You can also launch the GUI locally:
$ python -m causalexplain gui
Prior knowledge (ReX)
ReX can optionally use prior knowledge to constrain edge directions when you
already know a rough ordering of variables (for example, temporal tiers).
The prior is a JSON file with a single prior key whose value is a list of
tiers; each tier is a list of column names. Variables in earlier tiers may
cause variables in later tiers, but not vice versa. All names must match the
dataset columns, but you can omit variables you have no prior knowledge about.
Example JSON file:
{
"prior": [
["A", "B"],
["C", "D"]
]
}
Use it from the CLI with -p/--prior (ReX only):
$ python -m causalexplain run -d /path/to/data.csv -p /path/to/prior.json
Or from a notebook:
prior = [["A", "B"], ["C", "D"]]
experiment.run(prior=prior, hpo_iterations=10, bootstrap_iterations=10)
Citation
If you use CausalExplain, please cite the software and/or the related publication below.
Software
Renero, J. (2026). CausalExplain (Version 0.8.0). Available at: https://github.com/renero/causalexplain
BibTeX
@software{causalexplain_software,
author = {Jesús Renero},
title = {CausalExplain},
version = {0.8.0},
url = {https://github.com/renero/causalexplain},
date = {2026-01-04}
}
Related Publication
Renero, J., Maestre, R., & Ochoa, I. (2026). ReX: Causal discovery based on machine learning and explainability techniques. Pattern Recognition, 172, 112491. https://doi.org/10.1016/j.patcog.2025.112491
BibTeX
@article{Renero2026ReX,
author = {Jesús Renero and Roberto Maestre and Idoia Ochoa},
title = {ReX: Causal discovery based on machine learning and explainability techniques},
journal = {Pattern Recognition},
volume = {172},
pages = {112491},
year = {2026},
doi = {10.1016/j.patcog.2025.112491},
url = {https://doi.org/10.1016/j.patcog.2025.112491}
}
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file causalexplain-0.9.4.tar.gz.
File metadata
- Download URL: causalexplain-0.9.4.tar.gz
- Upload date:
- Size: 992.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b48b7e35bc87dd1fca462aae92bb15f76b27320ce0b6cde9fa78930d2bdf7c4a
|
|
| MD5 |
86142b1ee91aa28c3cade4a1d0be44a1
|
|
| BLAKE2b-256 |
477dd3a7628a76a306425c464b32d2f293780bfae1fac0934c4530090d782bc5
|
File details
Details for the file causalexplain-0.9.4-py3-none-any.whl.
File metadata
- Download URL: causalexplain-0.9.4-py3-none-any.whl
- Upload date:
- Size: 966.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.10.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d41329ac0ca3ac4e537172cea25181231dd1a713dc34bb1ab66f008fa14d7ab4
|
|
| MD5 |
52d61025d4901a1eda8f0768cdf88868
|
|
| BLAKE2b-256 |
286ecd8d19cac81f9463ef92a8c0d27dd937d4b5b263a4f24d28c1b3912c4804
|