No project description provided
Project description
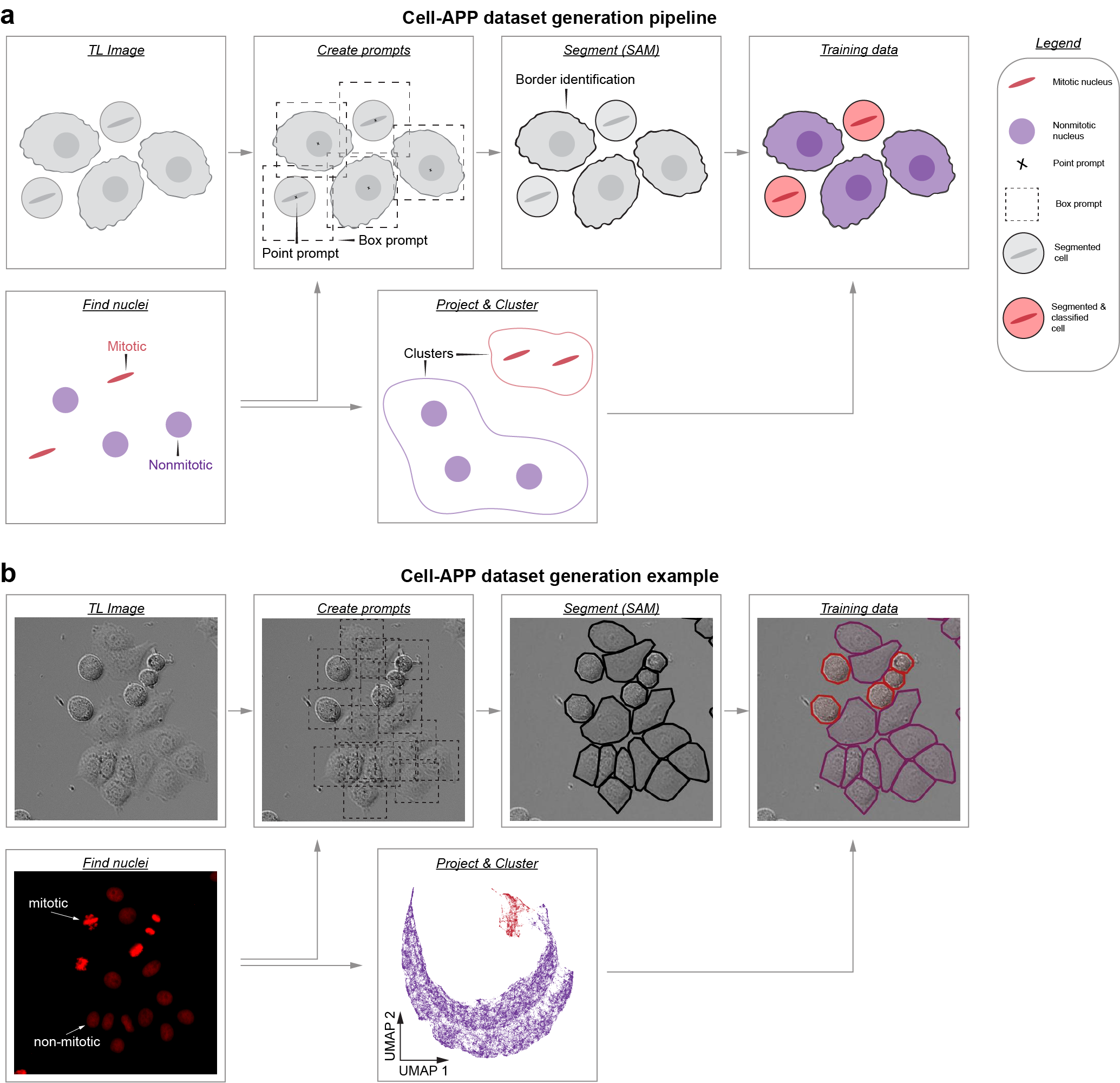
Cellular Annotation & Perception Pipeline
Description
Cell-APP automates the generation of cell masks (and classifications too!), enabling users to create custom instance segmentation training datasets in transmitted-light microscopy.
To learn more, read our preprint: https://www.biorxiv.org/content/10.1101/2025.01.23.634498v2.
For questions regarding installation or usage, contact: anishjv@umich.edu
Usage
-
Users who wish to segment HeLa, U2OS, HT1080, or RPE-1 cell lines may try our pre-trained model. These models can be used through our GUI (see Installation) and their weights can be downloaded at: https://zenodo.org/communities/cellapp/records?q=&l=list&p=1&s=10. To learn about using pre-trained models through the GUI, see this video:
-
Users who wish to segment their own cell lines may: (a) try our "general" model (GUI/weight download) or (b) train a custom model by creating an instance segmentation dataset via our Dataset Generation GUI (see Installation). To learn about creating custom datasets through the GUI, see this video:
Installation
cell-AAP requires Python 3.11–3.12. We recommend installing into a clean virtual environment (via conda or venv) to avoid dependency conflicts.
1. Create and activate an environment
With conda:
conda create -n cellapp -c conda-forge python=3.11
conda activate cellapp
Or with venv:
python -m venv cellapp
source cellapp/bin/activate # Linux/Mac
cellapp\Scripts\activate # Windows PowerShell
2. Install Pytorch:
conda install -c pytorch -c conda-forge pytorch torchvision #Mac
pip install torch torchvision #Linux/Windows
3. Install Cell-APP:
pip install cell-AAP
4. Finally, detectron2 must be built from source atop Cell-APP:
#Mac
CC=clang CXX=clang++ ARCHFLAGS="-arch arm64" python -m pip install 'git+https://github.com/facebookresearch/detectron2.git'
#Linux/Windows
python -m pip install 'git+https://github.com/facebookresearch/detectron2.git'
Napari Plugin Usage
- To open napari simply type "napari" into the command line, ensure that you are working the correct environment
- To instantiate the plugin, navigate to the "Plugins" menu and hover over "cell-AAP"
- You should see three plugin options; two relate to Usage 1; one relates to Usage 2.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file cell_aap-1.0.3.tar.gz.
File metadata
- Download URL: cell_aap-1.0.3.tar.gz
- Upload date:
- Size: 34.5 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
db79c56d9fd4f2c1e87654c0f8faa15aa041cd0d001b683d2bf98215a7067347
|
|
| MD5 |
fdd863a8d7e40acf2c2f50ac6ba3627b
|
|
| BLAKE2b-256 |
66fa8524649d1eeba790b13483d58a3d6582ccf9cfd6dc9d21b472d16b2ab674
|
File details
Details for the file cell_aap-1.0.3-py3-none-any.whl.
File metadata
- Download URL: cell_aap-1.0.3-py3-none-any.whl
- Upload date:
- Size: 60.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
26644dd0bfc01941039d43d131bd03f434e21c3e7d1e17958d8b46e06b797326
|
|
| MD5 |
c6e1de95b0a3a86cd4d8aa426228450c
|
|
| BLAKE2b-256 |
7f5c6bd6838dec17088dda0d311e25e66c33dcc765c8528ff8bb721ed1a2e4a5
|