Cell segmentation, tracking and event annotation
Project description
 Welcome to Cell-ACDC!
Welcome to Cell-ACDC!
A GUI-based Python framework for segmentation, tracking, cell cycle annotations and quantification of microscopy data
Written in Python 3 by Francesco Padovani and Benedikt Mairhoermann.
Core developers: Francesco Padovani, Timon Stegmaier, and Benedikt Mairhoermann.
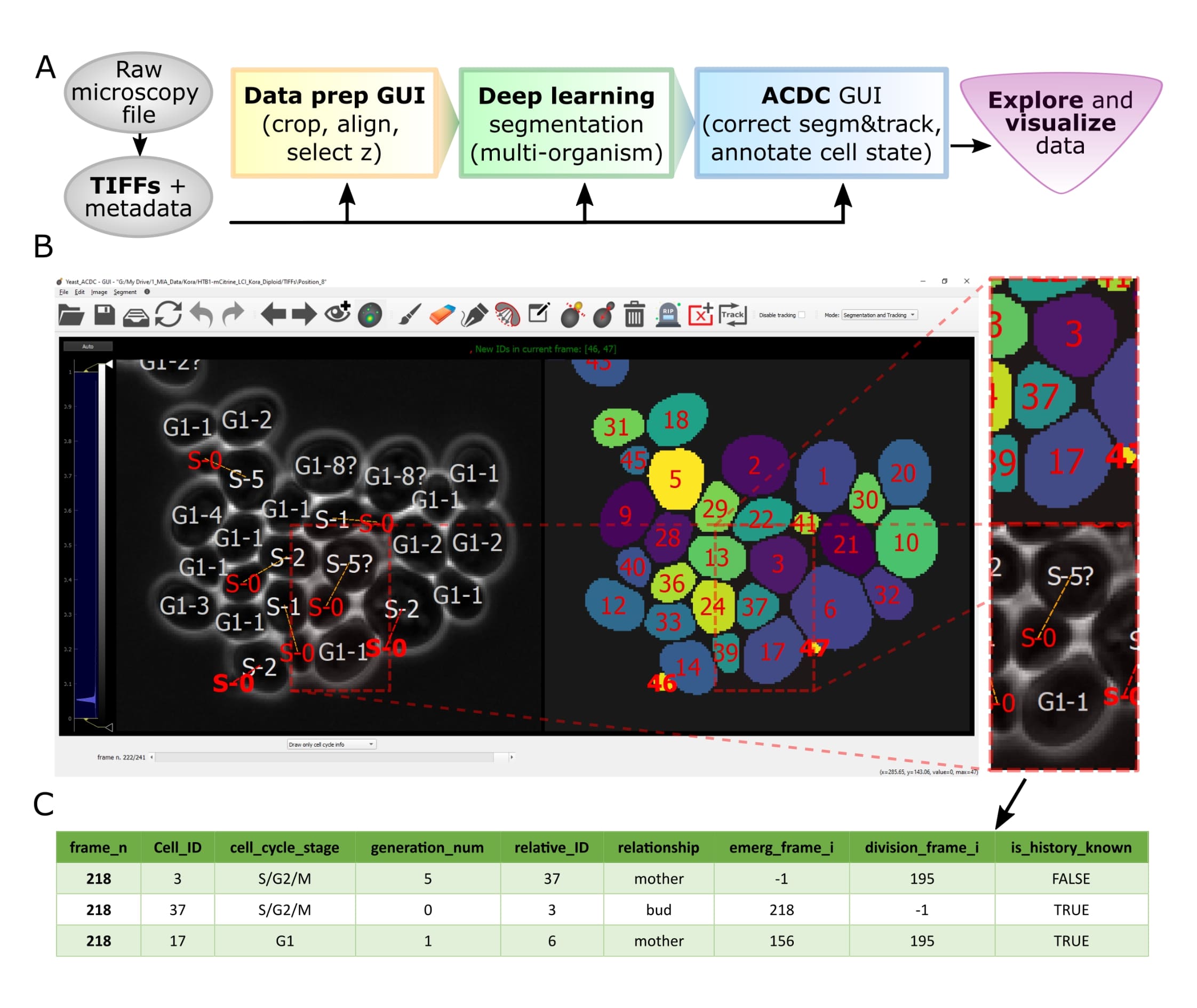
Overview of pipeline and GUI
Overview
Let’s face it, when dealing with segmentation of microscopy data we often do not have time to check that everything is correct, because it is a tedious and very time consuming process. Cell-ACDC comes to the rescue! We combined the currently best available neural network models (such as Segment Anything Model (SAM), YeaZ, cellpose, StarDist, YeastMate, omnipose, delta, DeepSea, etc.) and we complemented them with a fast and intuitive GUI.
We developed and implemented several smart functionalities such as real-time continuous tracking, automatic propagation of error correction, and several tools to facilitate manual correction, from simple yet useful brush and eraser to more complex flood fill (magic wand) and Random Walker segmentation routines.
See the table below how it compares to other popular tools available (Table 1 of our publication).
Feature |
Cell-ACDC |
YeaZ |
Cell-pose |
Yeast-Mate |
Deep-Cell |
Phylo-Cell |
Cell-Profiler |
ImageJ Fiji |
Yeast-Spotter |
Yeast-Net |
Morpho-LibJ |
|---|---|---|---|---|---|---|---|---|---|---|---|
Deep-learning segmentation |
✅ |
✅ |
✅ |
✅ |
✅ |
❌ |
✅ |
✅ |
✅ |
✅ |
❌ |
Traditional segmentation |
✅ |
❌ |
❌ |
❌ |
❌ |
✅ |
✅ |
✅ |
❌ |
❌ |
✅ |
Tracking |
✅ |
✅ |
❌ |
❌ |
✅ |
✅ |
✅ |
✅ |
❌ |
❌ |
❌ |
Manual corrections |
✅ |
✅ |
✅ |
✅ |
✅ |
✅ |
✅ |
✅ |
❌ |
❌ |
✅ |
Automatic real-time tracking |
✅ |
❌ |
❌ |
❌ |
❌ |
❌ |
❌ |
❌ |
❌ |
❌ |
❌ |
Automatic propagation of corrections |
✅ |
❌ |
❌ |
❌ |
❌ |
✅ |
❌ |
❌ |
❌ |
❌ |
❌ |
Automatic mother-bud pairing |
✅ |
❌ |
❌ |
✅ |
❌ |
✅ |
❌ |
❌ |
❌ |
❌ |
❌ |
Pedigree annotations |
✅ |
❌ |
❌ |
✅ |
✅ |
✅ |
✅ |
✅ |
❌ |
❌ |
❌ |
Cell division annotations |
✅ |
❌ |
❌ |
❌ |
❌ |
✅ |
✅ |
✅ |
❌ |
❌ |
❌ |
Downstream analysis |
✅ |
❌ |
❌ |
❌ |
✅ |
✅ |
✅ |
✅ |
❌ |
❌ |
❌ |
3D z-stacks |
✅ |
❌ |
✅ |
❌ |
✅ |
❌ |
✅ |
✅ |
❌ |
❌ |
✅ |
Multiple model organisms |
✅ |
❌ |
✅ |
❌ |
✅ |
❌ |
✅ |
✅ |
❌ |
❌ |
✅ |
Bio-formats |
✅ |
❌ |
❌ |
❌ |
❌ |
❌ |
✅ |
✅ |
❌ |
❌ |
✅ |
User manual |
✅ |
✅ |
✅ |
✅ |
✅ |
❌ |
✅ |
✅ |
✅ |
✅ |
✅ |
Open source |
✅ |
✅ |
✅ |
✅ |
✅ |
✅ |
✅ |
✅ |
✅ |
✅ |
✅ |
Does not require a licence |
✅ |
✅ |
✅ |
✅ |
✅ |
❌ |
✅ |
✅ |
✅ |
✅ |
✅ |
Is it only about segmentation?
Of course not! Cell-ACDC automatically computes several single-cell numerical features such as cell area and cell volume, plus the mean, max, median, sum and quantiles of any additional fluorescent channel’s signal. It even performs background correction, to compute the protein amount and concentration.
You can load and analyse single 2D images, 3D data (3D z-stacks or 2D images over time) and even 4D data (3D z-stacks over time).
Finally, we provide Jupyter notebooks to visualize and interactively explore the data produced.
Scientific publications where Cell-ACDC was used
Check here for a list of the scientific publications where Cell-ACDC was used.
Resources
Publication of Cell-ACDC
Image.sc Forum to ask any question. Make sure to tag the Topic with the tag cell-acdc
GitHub issues for reporting issues, request a feature or ask questions
Scientific publications where Cell-ACDC was used
Citing Cell-ACDC and the available models
If you find Cell-ACDC useful, please cite it as follows:
Padovani, F., Mairhörmann, B., Falter-Braun, P., Lengefeld, J. & Schmoller, K. M. Segmentation, tracking and cell cycle analysis of live-cell imaging data with Cell-ACDC. BMC Biology 20, 174 (2022). DOI: 10.1186/s12915-022-01372-6
IMPORTANT: when citing Cell-ACDC make sure to also cite the paper of the segmentation models and trackers you used! See here for a list of models currently available in Cell-ACDC.
Contact
Do not hesitate to contact us here on GitHub (by opening an issue) or directly at the email padovaf@tcd.ie for any problem and/or feedback on how to improve the user experience!
Contributing
At Cell-ACDC we encourage contributions to the code! Please read our contributing guide to get started.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file cellacdc-1.7.1.tar.gz.
File metadata
- Download URL: cellacdc-1.7.1.tar.gz
- Upload date:
- Size: 21.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
88b63ed911eba177c49a5fb41f8d75ed82e21939d5787f8b47941dbc54b38e9f
|
|
| MD5 |
442456b108b638dcba37e4fd50441bd6
|
|
| BLAKE2b-256 |
80a6ed13dad88b1cb5e0a65fd39123b32a35b6f4f05f900b35b1c67ae17b184f
|
File details
Details for the file cellacdc-1.7.1-py3-none-any.whl.
File metadata
- Download URL: cellacdc-1.7.1-py3-none-any.whl
- Upload date:
- Size: 16.3 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
33e4364d06643a31448066f863eac1941ae1039dfb26cacf6b1f8497c4206950
|
|
| MD5 |
86b77a98faf8503bfaa7bef72a122e0b
|
|
| BLAKE2b-256 |
4213d48d7f34b9275644b9a1372b39793c1e84a54e01fe6e6bc097e6fb514d5c
|
























