An AI ensemble model for predicting chemical classes
Project description
python-chebifier
An AI ensemble model for predicting chemical classes in the ChEBI ontology. It integrates deep learning models, rule-based models and generative AI-based models.
A web application for Chebifier is available at https://chebifier.hastingslab.org/.
Installation
Not all models can be installed automatically at the moment:
chebai-graphand its dependencies. To install them, follow the instructions in the chebai-graph repository.chemlog-extracan be installed withpip install git+https://github.com/ChEB-AI/chemlog-extra.git- The automatically installed version of
c3pmay not work under Windows. If you want to run chebifier on Windows, we recommend using this forked version:pip install git+https://github.com/sfluegel05/c3p.git
You can get the package from PyPI:
pip install chebifier
or get the latest development version from GitHub:
# Clone the repository
git clone https://github.com/yourusername/python-chebifier.git
cd python-chebifier
# Install the package
pip install -e .
Usage
Command Line Interface
The package provides a command-line interface (CLI) for making predictions using an ensemble model.
The ensemble configuration is given by a configuration file (by default, this is chebifier/ensemble.yml). If you
want to change which models are included in the ensemble or how they are weighted, you can create your own configuration file.
Trained deep learning models are automatically downloaded from Hugging Face.
To access a model from Hugging face, add the load_model key in your configuration file. For example:
my_electra:
type: electra
load_model: "electra_chebi50-3star_v244"
Available model weights:
resgated-aug_chebi50-3star_v244gat-aug_chebi50_v244electra_chebi50-3star_v244gat_chebi50_v244electra_chebi50_v241resgated_chebi50_v241c3p_with_weights
You can also supply your own model checkpoints (see configs/example_config.yml for an example).
# Make predictions
python -m chebifier predict --smiles "CC(=O)OC1=CC=CC=C1C(=O)O" --smiles "C1=CC=C(C=C1)C(=O)O"
# Make predictions using SMILES from a file
python -m chebifier predict --smiles-file smiles.txt
# Make predictions using a configuration file
python -m chebifier predict --ensemble-config configs/my_config.yml --smiles-file smiles.txt
# Get all available options
python -m chebifier predict --help
Python API
You can use the package programmatically as well:
from chebifier import BaseEnsemble
# Instantiate ensemble model. Optionally, you can pass
# a path to a configuration, like 'configs/example_config.yml'
ensemble = BaseEnsemble()
# Make predictions
smiles_list = ["CC(=O)OC1=CC=CC=C1C(=O)O", "C1=CC=C(C=C1)C(=O)O"]
predictions = ensemble.predict_smiles_list(smiles_list)
# Print results
for smiles, prediction in zip(smiles_list, predictions):
print(f"SMILES: {smiles}")
if prediction:
print(f"Predicted classes: {prediction}")
else:
print("No predictions")
The models
Currently, the following models are supported:
| Model | Description | #Classes | Publication | Repository |
|---|---|---|---|---|
electra |
A transformer-based deep learning model trained on ChEBI SMILES strings. | 1531* | Glauer, Martin, et al., 2024: Chebifier: Automating semantic classification in ChEBI to accelerate data-driven discovery, Digital Discovery 3 (2024) 896-907 | python-chebai |
resgated |
A Residual Gated Graph Convolutional Network trained on ChEBI molecules. | 1531* | python-chebai-graph | |
gat |
A Graph Attention Network trained on ChEBI molecules. | 1531* | python-chebai-graph | |
chemlog_peptides |
A rule-based model specialised on peptide classes. | 18 | Flügel, Simon, et al., 2025: ChemLog: Making MSOL Viable for Ontological Classification and Learning, arXiv | chemlog-peptides |
chemlog_element, chemlog_organox |
Extensions of ChemLog for classes that are defined either by the presence of a specific element or by the presence of an organic bond. | 118 + 37 | chemlog-extra | |
c3p |
A collection Chemical Classifier Programs, generated by LLMs based on the natural language definitions of ChEBI classes. | 338 | Mungall, Christopher J., et al., 2025: Chemical classification program synthesis using generative artificial intelligence, Journal of Cheminsformatics | c3p |
In addition, Chebifier also includes a ChEBI lookup that automatically retrieves the ChEBI superclasses for a class matched by a SMILES string. This is not activated by default, but can be included by adding
chebi_lookup:
type: chebi_lookup
model_weight: 10 # optional
to your configuration file.
The ensemble
For an extended description of the ensemble, see Flügel, Simon, et al., 2025: Chebifier 2: An Ensemble for Chemistry.
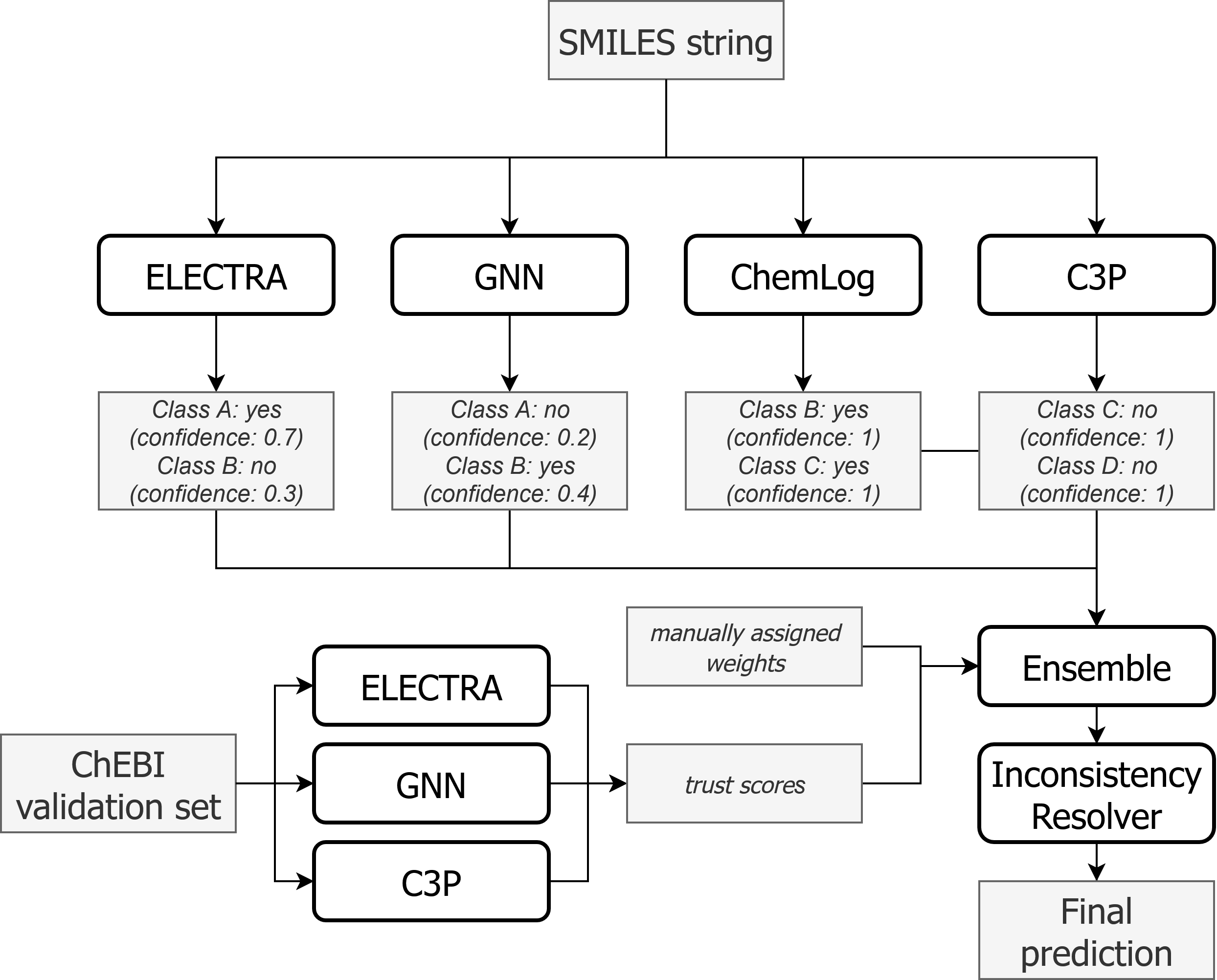
Given a sample (i.e., a SMILES string) and models $m_1, m_2, \ldots, m_n$, the ensemble works as follows:
- Get predictions from each model $m_i$ for the sample.
- For each class $c$, aggregate predictions $p_c^{m_i}$ from all models that made a prediction for that class. The aggregation happens separately for all positive predictions (i.e., $p_c^{m_i} \geq 0.5$) and all negative predictions ($p_c^{m_i} < 0.5$). If the aggregated value is larger for the positive predictions than for the negative predictions, the ensemble makes a positive prediction for class $c$:
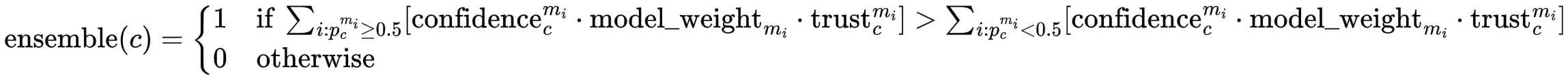
Here, confidence is the model's (self-reported) confidence in its prediction, calculated as $ \text{confidence}_c^{m_i} = 2|p_c^{m_i} - 0.5| $ For example, if a model makes a positive prediction with $p_c^{m_i} = 0.55$, the confidence is $2|0.55 - 0.5| = 0.1$. One could say that the model is not very confident in its prediction and very close to switching to a negative prediction. If another model is very sure about its negative prediction with $p_c^{m_j} = 0.1$, the confidence is $2|0.1 - 0.5| = 0.8$. Therefore, if in doubt, we are more confident in the negative prediction.
Confidence can be disabled by the use_confidence parameter of the predict method (default: True).
Themodel_weight can be set for each model in the configuration file (default: 1). This is used to favor a certain
model independently of a given class.
Trust is based on the model's performance on a validation set. After training, we evaluate the Machine Learning models
on a validation set for each class. If the ensemble_type is set to wmv-f1, the trust is calculated as F1-score $^{6.25}$.
If the ensemble_type is set to mv (the default), the trust is set to 1 for all models.
Inconsistency resolution
After a decision has been made for each class independently, the consistency of the predictions with regard to the ChEBI hierarchy and disjointness axioms is checked. This is done in 3 steps:
- (1) First, the hierarchy is corrected. For each pair of classes $A$ and $B$ where $A$ is a subclass of $B$ (following the is-a relation in ChEBI), we set the ensemble prediction of $A$ to $0$ if the absolute value of $B$'s score is large than that of $A$. For example, if $A$ has a net score of $3$ and $B$ has a net score of $-4$, the ensemble will set $A$ to $0$ (i.e., predict neither $A$ nor $B$).
- (2) Next, we check for disjointness. This is not specified directly in ChEBI, but in an additional ChEBI module (chebi-disjoints.owl).
We have extracted these disjointness axioms into a CSV file and added some more disjointness axioms ourselves (see
data>disjoint_chebi.csvanddata>disjoint_additional.csv). If two classes $A$ and $B$ are disjoint and we predict both, we select one with the higher class score and set the other to 0. - (3) Since the second step might have introduced new inconsistencies into the hierarchy, we repeat the first step, but with a small change. For a pair of classes $A \subseteq B$ with predictions $1$ and $0$, instead of setting $B$ to $1$, we now set $A$ to $0$. This has the advantage that we cannot introduce new disjointness-inconsistencies and don't have to repeat step 2.
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file chebifier-1.2.1.tar.gz.
File metadata
- Download URL: chebifier-1.2.1.tar.gz
- Upload date:
- Size: 33.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cf1295953e4f30245c63af243c825e7ac2911da4c04ffca45687b8f266c1d624
|
|
| MD5 |
faa86e1f91718e92af050fe00c7ad91a
|
|
| BLAKE2b-256 |
9b859ffba4f7d0977ae1ef0b8911f563ed006b9e1dd1c95a1b9f8127782c99b3
|
Provenance
The following attestation bundles were made for chebifier-1.2.1.tar.gz:
Publisher:
python-publish.yml on ChEB-AI/python-chebifier
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
chebifier-1.2.1.tar.gz -
Subject digest:
cf1295953e4f30245c63af243c825e7ac2911da4c04ffca45687b8f266c1d624 - Sigstore transparency entry: 956352217
- Sigstore integration time:
-
Permalink:
ChEB-AI/python-chebifier@3474ec6b41b8b46a24ebe90c37ab2d1d625a96d1 -
Branch / Tag:
- Owner: https://github.com/ChEB-AI
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@3474ec6b41b8b46a24ebe90c37ab2d1d625a96d1 -
Trigger Event:
release
-
Statement type:
File details
Details for the file chebifier-1.2.1-py3-none-any.whl.
File metadata
- Download URL: chebifier-1.2.1-py3-none-any.whl
- Upload date:
- Size: 35.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7dcc6823c3e35946412bb517f4ac54d3067bcdecd6f79c28f31ec3d53c0a9b50
|
|
| MD5 |
9cc7e316fb659a662e99acf40f767e56
|
|
| BLAKE2b-256 |
c5708c3b3158819daa50563914f4fc6906ef7e59cfd66749749cda551a252dd2
|
Provenance
The following attestation bundles were made for chebifier-1.2.1-py3-none-any.whl:
Publisher:
python-publish.yml on ChEB-AI/python-chebifier
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
chebifier-1.2.1-py3-none-any.whl -
Subject digest:
7dcc6823c3e35946412bb517f4ac54d3067bcdecd6f79c28f31ec3d53c0a9b50 - Sigstore transparency entry: 956352220
- Sigstore integration time:
-
Permalink:
ChEB-AI/python-chebifier@3474ec6b41b8b46a24ebe90c37ab2d1d625a96d1 -
Branch / Tag:
- Owner: https://github.com/ChEB-AI
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
python-publish.yml@3474ec6b41b8b46a24ebe90c37ab2d1d625a96d1 -
Trigger Event:
release
-
Statement type:












