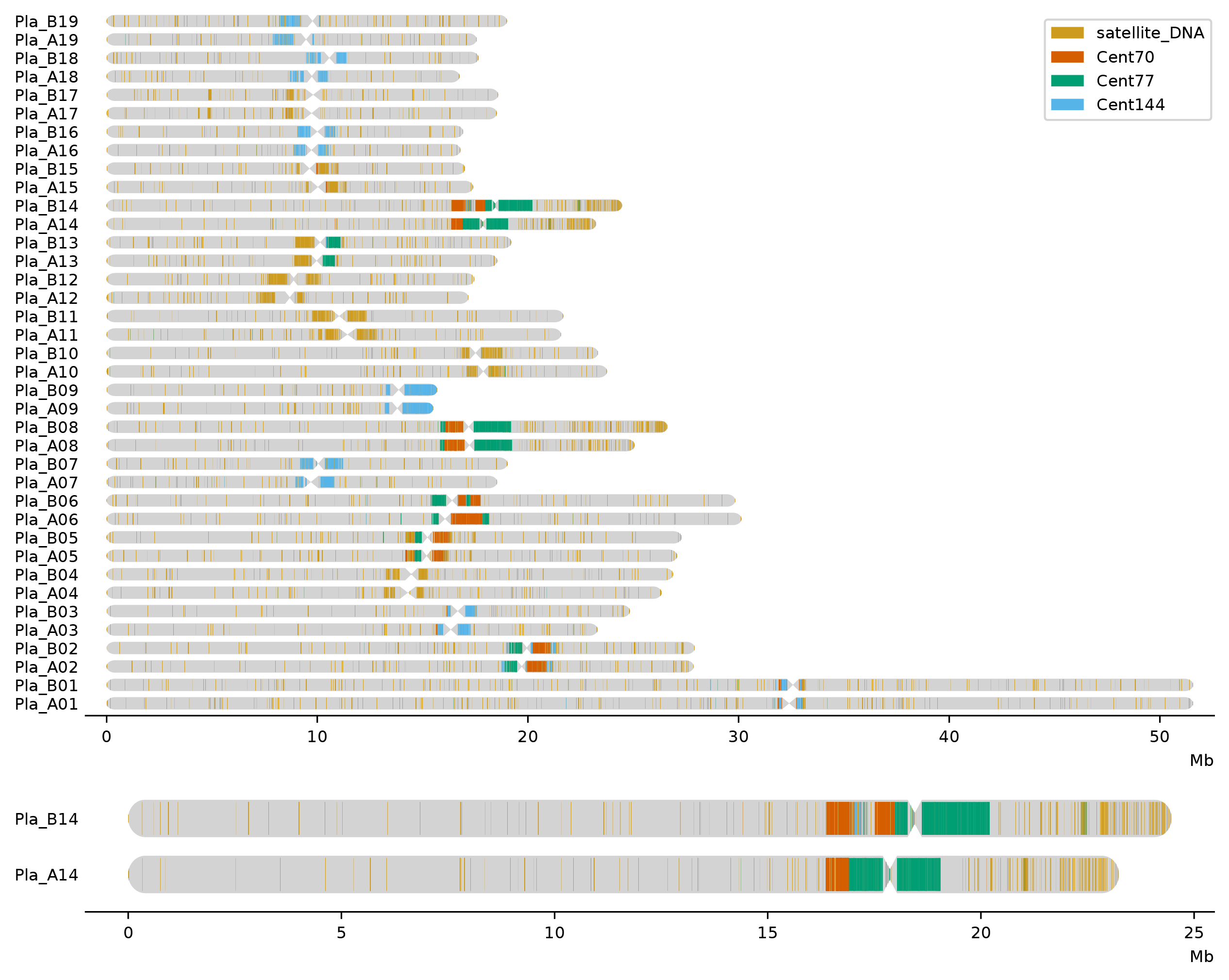
A python package for drawing genome-wide data on idiograms.
Project description
ChromosomeIdeogram
A python package for drawing genome-wide data on ideograms.
Install
pip install chromosome-ideogram --upgrade
Usage example
import matplotlib.pyplot as plt
from matplotlib import gridspec
from chromosome_ideogram import ChromosomeIdeogram
plt.rcParams['font.size'] = 8
plt.rcParams['pdf.fonttype'] = 42
fig = plt.figure(figsize=(10, 8))
gs = gridspec.GridSpec(nrows=10, ncols=1)
axes1 = fig.add_subplot(gs[:8, :])
axes2 = fig.add_subplot(gs[8:, :])
ci1 = ChromosomeIdeogram(
karyotype='example/karyotype.xls',
radius=0.4, # Radius of chromosome (it is not recommended to make any changes)
space=3 # Chromosome center coordinate interval (radius x space, also not recommended for modification)
)
ci1.draw_chromh(axes=axes1, centromere_angle=60, color='lightgrey')
ci1.annotations(axes=axes1, annotations='example/satellite_DNA.bed')
# Create a second karyotype diagram object (containing only chromosome 14)
ci2 = ChromosomeIdeogram(
karyotype='example/Chr14_karyotype.xls',
radius=0.4,
space=3
)
ci2.draw_chromh(axes=axes2, centromere_angle=60, color='lightgrey')
ci2.annotations(axes=axes2, annotations='example/Chr14_satellite_DNA.bed')
axes1.legend(loc=(0.85, 0.85)) # Set axes1 legend, you can also set axes2 legend
plt.subplots_adjust(hspace=6) # Adjust the vertical distance between the two subgraphs
plt.savefig('example/satellite_DNA.pdf', bbox_inches='tight')
plt.savefig('example/satellite_DNA.png', bbox_inches='tight')
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file chromosome_ideogram-0.0.3.tar.gz.
File metadata
- Download URL: chromosome_ideogram-0.0.3.tar.gz
- Upload date:
- Size: 4.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.8.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
92803970181b167cf77f8f031898001248f06776489f327b997858bfec78f44b
|
|
| MD5 |
b64da9f37f1e5cfcf8666f112dfd2cd9
|
|
| BLAKE2b-256 |
c9d4d4d281860b56cbf1d2ebc1d6c1e72eb0c2785bde3f9cd4f1dc727dd37d60
|
File details
Details for the file chromosome_ideogram-0.0.3-py3-none-any.whl.
File metadata
- Download URL: chromosome_ideogram-0.0.3-py3-none-any.whl
- Upload date:
- Size: 5.0 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.8.20
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
fa8a93f985d6aaa3455bb0f930df66a30a63d2f53b409cc5e23fe60d2389552a
|
|
| MD5 |
0e02b88a0c8f82cbea83f78992c4f9fb
|
|
| BLAKE2b-256 |
f41d919b830d149b3ec5f977fbda7c9670fb12debedaf1780fee86d58eb18bce
|