Package for building co-accessibility networks from ATAC-seq data.
Project description

CIRCE: Cis-regulatory interactions between chromatin regions
Description
This repo contains a python package for inferring co-accessibility networks from single-cell ATAC-seq data, using skggm for the graphical lasso and scanpy for data processing.
It is based on the pipeline and hypotheses presented in the manuscript "Cicero Predicts cis-Regulatory DNA Interactions from Single-Cell Chromatin Accessibility Data" by Pliner et al. (2018). This R package Cicero is available here.
Results may slitghly vary between both packages, notably due to the different implementations of graphical lasso.
Currently, scores are very close when applied to the same metacells, computed from Cicero's methodology. (cf comparison plots below). It should run significantly faster than Cicero (e.g.: running time of 5 sec instead of 17 min for the dataset 2).
If you have any suggestion, don't hesitate ! This package is still a work in progress :)
Installation
The package can be installed using pip:
pip install circe
and from github
pip install "git+https://github.com/cantinilab/circe.git"
Minimal example
import anndata as ad
import circe as an
atac = ad.read_h5ad('atac_data.h5ad')
an.add_region_infos(atac)
an.compute_atac_network(atac)
an.extract_atac_links(atac)
Comparison to Cicero R package
On the same metacells obtained from Cicero code.
All tests can be found in the circe benchmark repo
Toy dataset 1 (fake data):
- Pearson correlation coefficient: 0.999126
- Spearman correlation coefficient: 0.99838
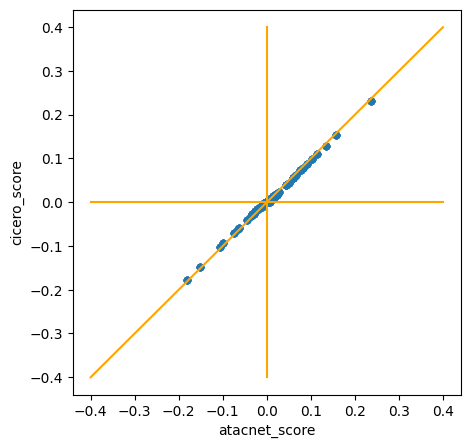
Real dataset 2 (subsample of neurips PBMC)
- Pearson correlation coefficient: 0.999958
- Spearman correlation coefficient: 0.999911
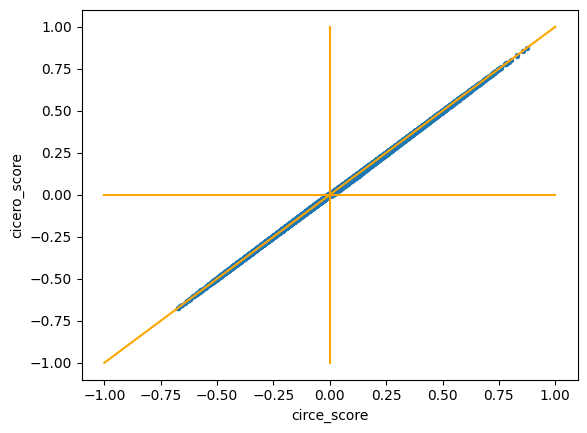
Performance on real dataset 2:
- Runtime: ~100x faster
- Memory usage: ~5x less
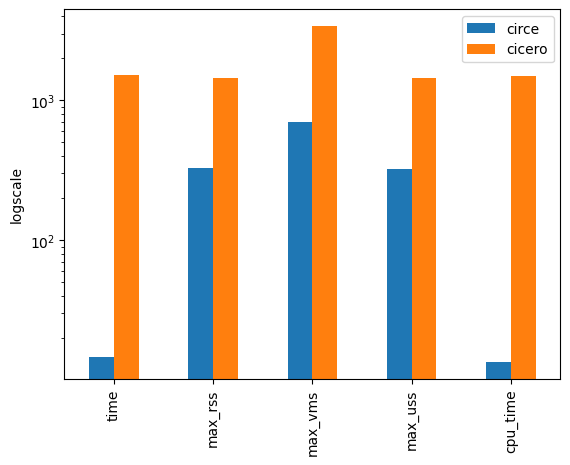
Coming:
- Calculate metacells !
- Add stats on similarity on large datasets.
- Add stats on runtime, memory usage.
- This package can be run on multiple cores. Currently working to speed up the mutlithreding use.
- Fix seed for reproducibility.
Usage
It is currently developped to work with AnnData objects. Check Example1.ipynb for a simple usage example.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file circe_py-0.1.8.tar.gz.
File metadata
- Download URL: circe_py-0.1.8.tar.gz
- Upload date:
- Size: 235.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.4.2 CPython/3.10.9 Linux/6.5.0-41-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ff9a174890de21126fc31d59ac8d10dbe3e6284365462ada75027a134d21d155
|
|
| MD5 |
2db8e67b84ce05355681b5f65ae78690
|
|
| BLAKE2b-256 |
4cf3ee266ba7aa78abf40b7a04ec01f5a09febb9000e95d3fcf4eb4cbaa84099
|
File details
Details for the file circe_py-0.1.8-cp310-cp310-manylinux_2_35_x86_64.whl.
File metadata
- Download URL: circe_py-0.1.8-cp310-cp310-manylinux_2_35_x86_64.whl
- Upload date:
- Size: 236.8 kB
- Tags: CPython 3.10, manylinux: glibc 2.35+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.4.2 CPython/3.10.9 Linux/6.5.0-41-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
149a11018e75577e35d67dbcf8faac53b2024c47d4c6c41e280d39b9d7a16837
|
|
| MD5 |
b02060fc37cd3e8be56a59b6641f0a94
|
|
| BLAKE2b-256 |
70f21bd9351838a11a157185ce728858253a5cc202b8ddae377bb3a5e9248f81
|















