A scikit-learn meta-estimator for computing tight conformal predictions
Project description
👖 Conformal Tights
A scikit-learn meta-estimator that adds conformal prediction of coherent quantiles and intervals to any scikit-learn regressor. Features:
- 🍬 Meta-estimator: add prediction of quantiles and intervals to any scikit-learn regressor
- 🌡️ Conformally calibrated: accurate quantiles, and intervals with reliable coverage
- 🚦 Coherent quantiles: quantiles increase monotonically instead of crossing each other
- 👖 Tight quantiles: selects the lowest dispersion that provides the desired coverage
- 🎁 Data efficient: requires only a small number of calibration examples to fit
- 🐼 Pandas support: optionally predict on DataFrames and receive DataFrame output
Using
Installing
First, install this package with:
pip install conformal-tights
Predicting quantiles
Conformal Tights exposes a meta-estimator called ConformalCoherentQuantileRegressor that you can use to wrap any scikit-learn regressor, after which you can use predict_quantiles to predict conformally calibrated quantiles. Example usage:
from conformal_tights import ConformalCoherentQuantileRegressor
from sklearn.datasets import fetch_openml
from sklearn.model_selection import train_test_split
from xgboost import XGBRegressor
# Fetch dataset and split in train and test
X, y = fetch_openml("ames_housing", version=1, return_X_y=True, as_frame=True, parser="auto")
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.15, random_state=42)
# Create a regressor, wrap it, and fit on the train set
my_regressor = XGBRegressor(objective="reg:absoluteerror")
conformal_predictor = ConformalCoherentQuantileRegressor(estimator=my_regressor)
conformal_predictor.fit(X_train, y_train)
# Predict with the wrapped regressor
ŷ_test = conformal_predictor.predict(X_test)
# Predict quantiles with the conformal wrapper
ŷ_test_quantiles = conformal_predictor.predict_quantiles(X_test, quantiles=(0.025, 0.05, 0.1, 0.9, 0.95, 0.975))
When the input data is a pandas DataFrame, the output is also a pandas DataFrame. For example, printing the head of ŷ_test_quantiles yields:
| house_id | 0.025 | 0.05 | 0.1 | 0.9 | 0.95 | 0.975 |
|---|---|---|---|---|---|---|
| 1357 | 114784 | 120894 | 131618 | 175761 | 188052 | 205449 |
| 2367 | 67417 | 80074 | 86754 | 117854 | 127583 | 142322 |
| 2822 | 119423 | 132048 | 138725 | 178526 | 197246 | 214206 |
| 2126 | 94031 | 99850 | 110891 | 150249 | 164703 | 182528 |
| 1544 | 68996 | 81516 | 88232 | 121774 | 132425 | 147110 |
Let's visualize the predicted quantiles on the test set:
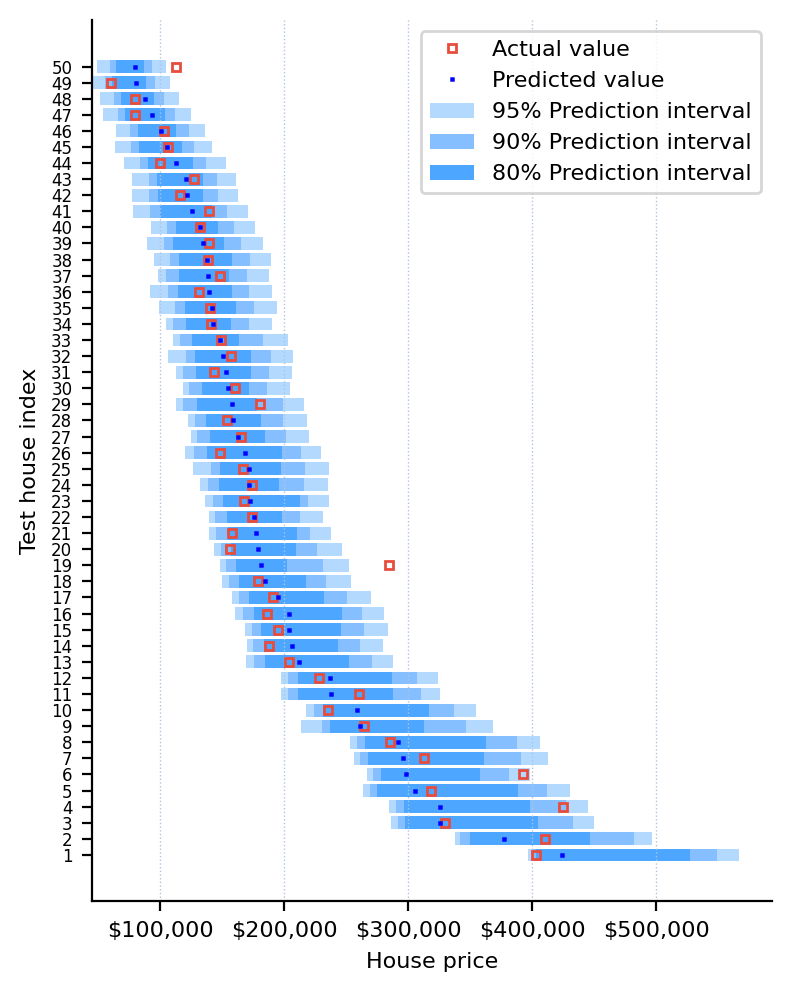
Expand to see the code that generated the graph above
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
%config InlineBackend.figure_format = "retina"
plt.rcParams["font.size"] = 8
idx = (-ŷ_test.sample(50, random_state=42)).sort_values().index
y_ticks = list(range(1, len(idx) + 1))
plt.figure(figsize=(4, 5))
for j in range(3):
end = ŷ_test_quantiles.shape[1] - 1 - j
coverage = round(100 * (ŷ_test_quantiles.columns[end] - ŷ_test_quantiles.columns[j]))
plt.barh(
y_ticks,
ŷ_test_quantiles.loc[idx].iloc[:, end] - ŷ_test_quantiles.loc[idx].iloc[:, j],
left=ŷ_test_quantiles.loc[idx].iloc[:, j],
label=f"{coverage}% Prediction interval",
color=["#b3d9ff", "#86bfff", "#4da6ff"][j],
)
plt.plot(y_test.loc[idx], y_ticks, "s", markersize=3, markerfacecolor="none", markeredgecolor="#e74c3c", label="Actual value")
plt.plot(ŷ_test.loc[idx], y_ticks, "s", color="blue", markersize=0.6, label="Predicted value")
plt.xlabel("House price")
plt.ylabel("Test house index")
plt.yticks(y_ticks, y_ticks)
plt.tick_params(axis="y", labelsize=6)
plt.grid(axis="x", color="lightsteelblue", linestyle=":", linewidth=0.5)
plt.gca().xaxis.set_major_formatter(ticker.StrMethodFormatter("${x:,.0f}"))
plt.gca().spines["top"].set_visible(False)
plt.gca().spines["right"].set_visible(False)
plt.legend()
plt.tight_layout()
plt.show()
Predicting intervals
In addition to quantile prediction, you can use predict_interval to predict conformally calibrated prediction intervals. Compared to quantiles, these focus on reliable coverage over quantile accuracy. Example usage:
# Predict an interval for each example with the conformal wrapper
ŷ_test_interval = conformal_predictor.predict_interval(X_test, coverage=0.95)
# Measure the coverage of the prediction intervals on the test set
coverage = ((ŷ_test_interval.iloc[:, 0] <= y_test) & (y_test <= ŷ_test_interval.iloc[:, 1])).mean()
print(coverage) # 96.6%
When the input data is a pandas DataFrame, the output is also a pandas DataFrame. For example, printing the head of ŷ_test_interval yields:
| house_id | 0.025 | 0.975 |
|---|---|---|
| 1357 | 107203 | 206290 |
| 2367 | 66665 | 146005 |
| 2822 | 115592 | 220315 |
| 2126 | 85288 | 183038 |
| 1544 | 67890 | 150646 |
Contributing
Prerequisites
1. Set up Git to use SSH
- Generate an SSH key and add the SSH key to your GitHub account.
- Configure SSH to automatically load your SSH keys:
cat << EOF >> ~/.ssh/config Host * AddKeysToAgent yes IgnoreUnknown UseKeychain UseKeychain yes EOF
2. Install Docker
- Install Docker Desktop.
- Enable Use Docker Compose V2 in Docker Desktop's preferences window.
- Linux only:
- Export your user's user id and group id so that files created in the Dev Container are owned by your user:
cat << EOF >> ~/.bashrc export UID=$(id --user) export GID=$(id --group) EOF
- Export your user's user id and group id so that files created in the Dev Container are owned by your user:
3. Install VS Code or PyCharm
- Install VS Code and VS Code's Dev Containers extension. Alternatively, install PyCharm.
- Optional: install a Nerd Font such as FiraCode Nerd Font and configure VS Code or configure PyCharm to use it.
Development environments
The following development environments are supported:
- ⭐️ GitHub Codespaces: click on Code and select Create codespace to start a Dev Container with GitHub Codespaces.
- ⭐️ Dev Container (with container volume): click on Open in Dev Containers to clone this repository in a container volume and create a Dev Container with VS Code.
- Dev Container: clone this repository, open it with VS Code, and run Ctrl/⌘ + ⇧ + P → Dev Containers: Reopen in Container.
- PyCharm: clone this repository, open it with PyCharm, and configure Docker Compose as a remote interpreter with the
devservice. - Terminal: clone this repository, open it with your terminal, and run
docker compose up --detach devto start a Dev Container in the background, and then rundocker compose exec dev zshto open a shell prompt in the Dev Container.
Developing
- This project follows the Conventional Commits standard to automate Semantic Versioning and Keep A Changelog with Commitizen.
- Run
poefrom within the development environment to print a list of Poe the Poet tasks available to run on this project. - Run
poetry add {package}from within the development environment to install a run time dependency and add it topyproject.tomlandpoetry.lock. Add--group testor--group devto install a CI or development dependency, respectively. - Run
poetry updatefrom within the development environment to upgrade all dependencies to the latest versions allowed bypyproject.toml. - Run
cz bumpto bump the package's version, update theCHANGELOG.md, and create a git tag.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file conformal_tights-0.2.1.tar.gz.
File metadata
- Download URL: conformal_tights-0.2.1.tar.gz
- Upload date:
- Size: 17.4 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.2 CPython/3.10.14 Linux/6.5.0-1016-azure
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
0940f933045645bc9ed77f4f2573360e296e80005cbdd79d3141b07e6ad62409
|
|
| MD5 |
238a0921034e59f48bec08ba444a4eb0
|
|
| BLAKE2b-256 |
90e85ad544053fa22bb2ebae39fa92b399e8c5cf15afeba514819c6a751a7011
|
File details
Details for the file conformal_tights-0.2.1-py3-none-any.whl.
File metadata
- Download URL: conformal_tights-0.2.1-py3-none-any.whl
- Upload date:
- Size: 14.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.2 CPython/3.10.14 Linux/6.5.0-1016-azure
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1d5be3fe395460c5182cb9d36c3b16f532a92684e1864be62ab707286e5b6c08
|
|
| MD5 |
a76d01190a5fd114cc830a0a2f938e81
|
|
| BLAKE2b-256 |
2c569c93dc17294c790a8981e42ebe3140c06ec3b20a75295db8f862c499a5b7
|













