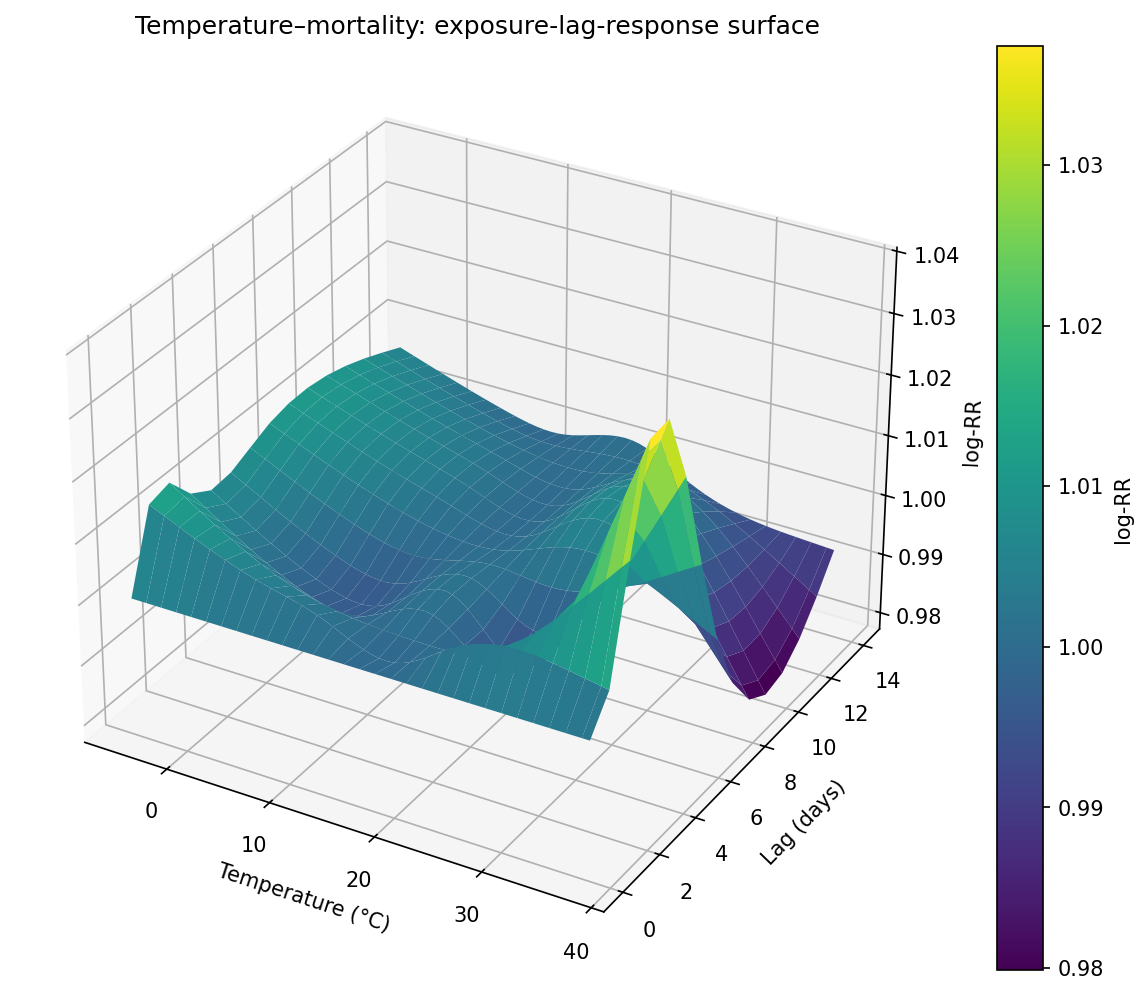
Distributed Lag Non-linear Models (DLNMs) in Python
Project description
crossbasis
Python implementation of Distributed Lag Non-linear Models (DLNMs) based on Gasparrini, Armstrong & Kenward (2010), Statistics in Medicine 29:2224–2234.
DLNMs model simultaneously non-linear and delayed effects of an exposure (e.g. temperature) on an outcome (e.g. daily deaths) — the standard approach in environmental epidemiology.
Installation
pip install crossbasis
Quick start
import numpy as np
import statsmodels.api as sm
from crossbasis import CrossPred
# Daily exposure and outcome (e.g. temperature and deaths)
temperature = np.random.randn(365) * 10 + 15
deaths = np.random.poisson(40, 365)
# 1. Fit: returns the cross-basis matrix W for use in your GLM
cp = CrossPred(
var_basis="ns", var_df=5, # exposure dimension: natural cubic spline
lag_basis="ns", lag_df=4, # lag dimension: natural cubic spline
max_lag=21, # model effects up to 21 days after exposure
cen="median", # reference value for relative risk
)
W = cp.fit(temperature)
# 2. Fit a Poisson GLM (add any confounders alongside W)
X = sm.add_constant(W)
model = sm.GLM(deaths, X, family=sm.families.Poisson()).fit()
# 3. Predict the full exposure-lag-response surface
result = cp.predict(model, at=np.arange(-5, 36))
# 4. Visualise
result.plot_overall() # cumulative effect across all lags
result.plot_slice(lag=0) # exposure-response at lag 0
result.plot_slice(var=30) # lag-response at 30 °C
result.plot_3d() # full 3D surface
Confounders in the model? When you include extra terms (seasonal splines,
day-of-week, etc.), extract only the cross-basis coefficients before calling
predict:
n_cb = W.shape[1]
result = cp.predict(
coef=model.params.values[1:n_cb + 1],
vcov=model.cov_params().values[1:n_cb + 1, 1:n_cb + 1],
at=np.arange(-5, 36),
)
Core API
CrossPred — high-level interface
CrossPred(
var_basis="ns", # "ns" | "bs" | "poly" | "linear" | callable
var_df=5,
lag_basis="ns",
lag_df=4,
max_lag=21, # required
na_action="drop", # "drop" | "fill_zero" | "fill_mean" | float
cen="median", # float | "median" | "mean" | "minimum_risk"
)
| Method | Returns |
|---|---|
cp.fit(X) |
Cross-basis matrix W — include in your GLM design matrix |
cp.predict(model, at=..., cen=...) |
PredictionResult |
cp.predict(coef=..., vcov=..., at=...) |
PredictionResult (no statsmodels needed) |
PredictionResult
| Attribute | Shape | Description |
|---|---|---|
matfit |
(m, L+1) |
log-RR at each (exposure, lag) |
matse |
(m, L+1) |
SE of matfit |
RR / RR_low / RR_high |
(m, L+1) |
Relative risk with 95% CI |
allfit |
(m,) |
Cumulative log-RR over all lags |
allRR / allRR_low / allRR_high |
(m,) |
Cumulative RR with 95% CI |
result.to_frame() # long-format DataFrame (exposure, lag, logRR, se, RR, …)
result.plot_overall() # cumulative effect curve
result.plot_slice(lag=7) # exposure-response at a specific lag
result.plot_slice(var=25) # lag-response at a specific exposure value
result.plot_3d() # 3D surface
CrossBasis — power-user transformer
Use this directly when you need the cross-basis matrix outside statsmodels (e.g. in scikit-learn, PyMC, or JAX).
from crossbasis import CrossBasis
cb = CrossBasis(var_basis="ns", var_df=5, lag_basis="ns", lag_df=4, max_lag=21)
W = cb.fit_transform(exposure_data) # (n, v_x * v_l) numpy array
Custom basis functions
Any callable that returns an (n, df) float64 array works:
def my_basis(x: np.ndarray, df: int, **kwargs) -> np.ndarray:
...
cp = CrossPred(var_basis=my_basis, var_df=3, lag_basis="ns", lag_df=3, max_lag=14)
Centering
All relative risks are expressed relative to a reference exposure value (cen).
cen= |
Behaviour |
|---|---|
float |
Fixed reference value |
"median" (default) |
Median of training exposure |
"mean" |
Mean of training exposure |
"minimum_risk" |
Approximate minimum of the cumulative RR curve (experimental) |
References
Gasparrini A, Armstrong B, Kenward MG (2010). Distributed lag non-linear models. Statistics in Medicine 29:2224–2234. doi:10.1002/sim.3940
License
MIT
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file crossbasis-0.1.0.tar.gz.
File metadata
- Download URL: crossbasis-0.1.0.tar.gz
- Upload date:
- Size: 23.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7f3d443f66fe0e1b178eb5a9a753ae0271cf054df7180a635dbf48ee8dcd5b65
|
|
| MD5 |
3346be360c6fb5c58fcf366dbea03acb
|
|
| BLAKE2b-256 |
94327c9f2ab8528e014633df96981bf8631ae645364fdb7fe48975e43a6911f4
|
File details
Details for the file crossbasis-0.1.0-py3-none-any.whl.
File metadata
- Download URL: crossbasis-0.1.0-py3-none-any.whl
- Upload date:
- Size: 16.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.13
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
560066a62669616e4e1430b023de78c979e9a0e93b95cac2ed22173308ab9430
|
|
| MD5 |
02293a52c39c32c20ffdea76850f3d01
|
|
| BLAKE2b-256 |
4e42e4044608dc05d4db8729d42f9efe74961b681f575ea49716c4fc188da504
|