Python client for characterization of clusters from single-cell RNA-seq data.
Project description
CyteType
Agentic, Evidence-Based Cell Type Annotation for Single-Cell RNA-seq
CyteType performs automated cell type annotation in single-cell RNA sequencing (scRNA-seq) data. It uses a multi-agent AI architecture to deliver transparent, evidence-based annotations with Cell Ontology mapping.
Integrates with Scanpy and Seurat workflows.
Preprint published: Nov. 7, 2025: bioRxiv link - Dive into benchmarking results
Why CyteType?
Cell type annotation is one of the most time-consuming steps in single-cell analysis. It typically requires weeks of expert curation, and the results often vary between annotators. When annotations do get done, the reasoning is rarely documented; this makes it difficult to reproduce or audit later.
CyteType addresses this with a novel agentic architecture: specialized AI agents collaborate on marker gene analysis, literature evidence retrieval, and ontology mapping. The result is consistent, reproducible annotations with a full evidence trail for every decision.
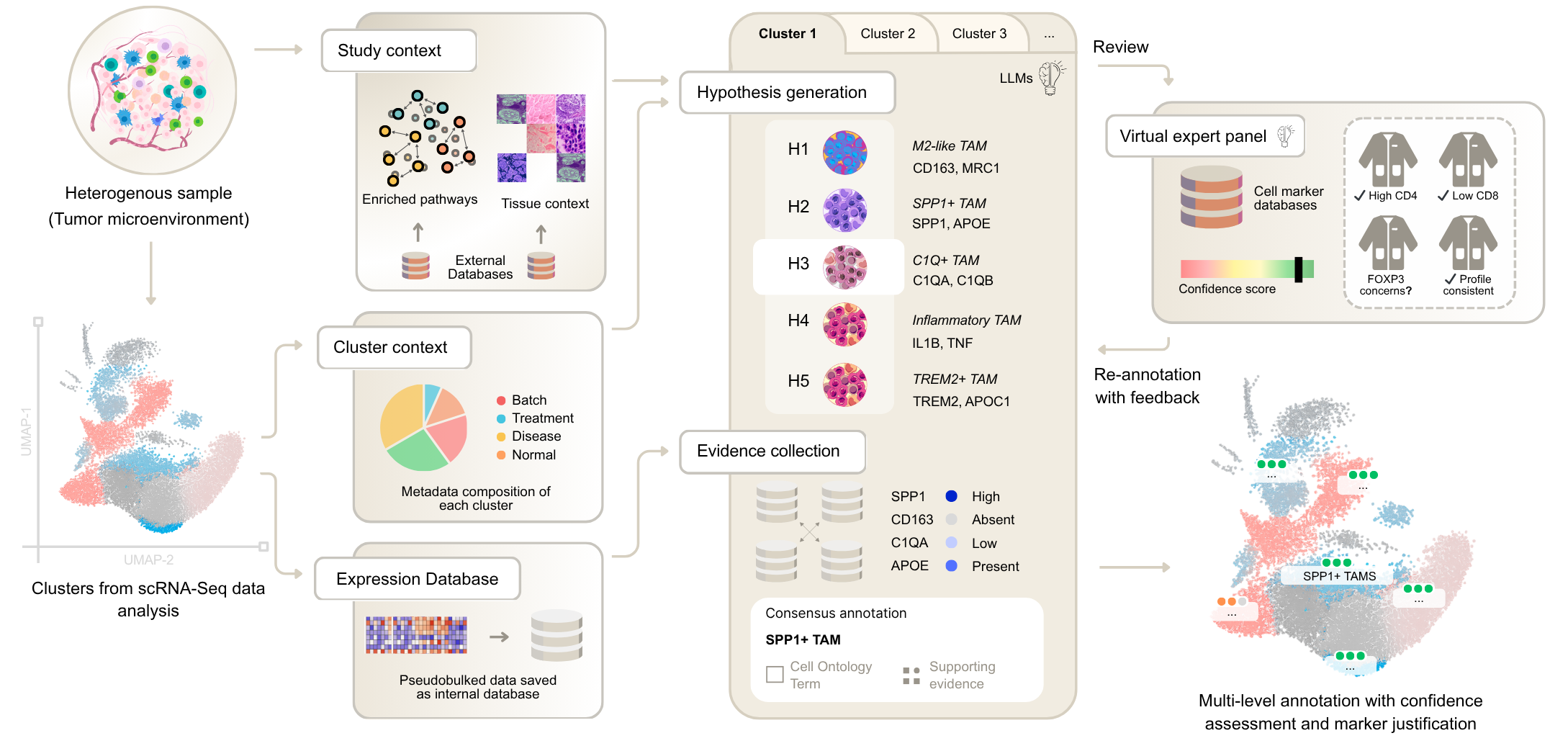
Key Features
| Feature | Description |
|---|---|
| Cell Ontology Integration | Automatic CL ID assignment for standardized terminology and cross-study comparison |
| Confidence Scores | Numeric certainty values (0–1) for cell type, subtype, and activation state — useful for flagging ambiguous clusters |
| Linked Literature | Each annotation includes supporting publications and condition-specific references — see exactly why a call was made |
| Annotation QC via Match Scores | Compare CyteType results against your existing annotations to quickly identify discrepancies and validate previous work |
| Embedded Chat Interface | Explore results interactively; chat is connected to your expression data for on-the-fly queries |
Also included: interactive HTML reports, Scanpy/Seurat compatibility (R wrapper via CyteTypeR), and no API keys required out of the box.
Quick Start
Installation
pip install cytetype
Basic Usage with Scanpy
import scanpy as sc
from cytetype import CyteType
# Assumes preprocessed AnnData with clusters and marker genes
group_key = 'clusters'
annotator = CyteType(
adata,
group_key=group_key,
rank_key='rank_genes_' + group_key,
n_top_genes=100
)
adata = annotator.run(study_context="Human PBMC from healthy donor")
sc.pl.umap(adata, color='cytetype_annotation_clusters')
Note: No API keys required for default configuration. See Configuration for LLM setup, artifact handling, and advanced options.
Using R/Seurat? → CyteTypeR
Documentation
| Resource | Description |
|---|---|
| Configuration | LLM settings, parameters, and customization |
| Output Columns | Understanding annotation results and metadata |
| Troubleshooting | Common issues and solutions |
| Development | Contributing and local setup |
| Discord | Community support |
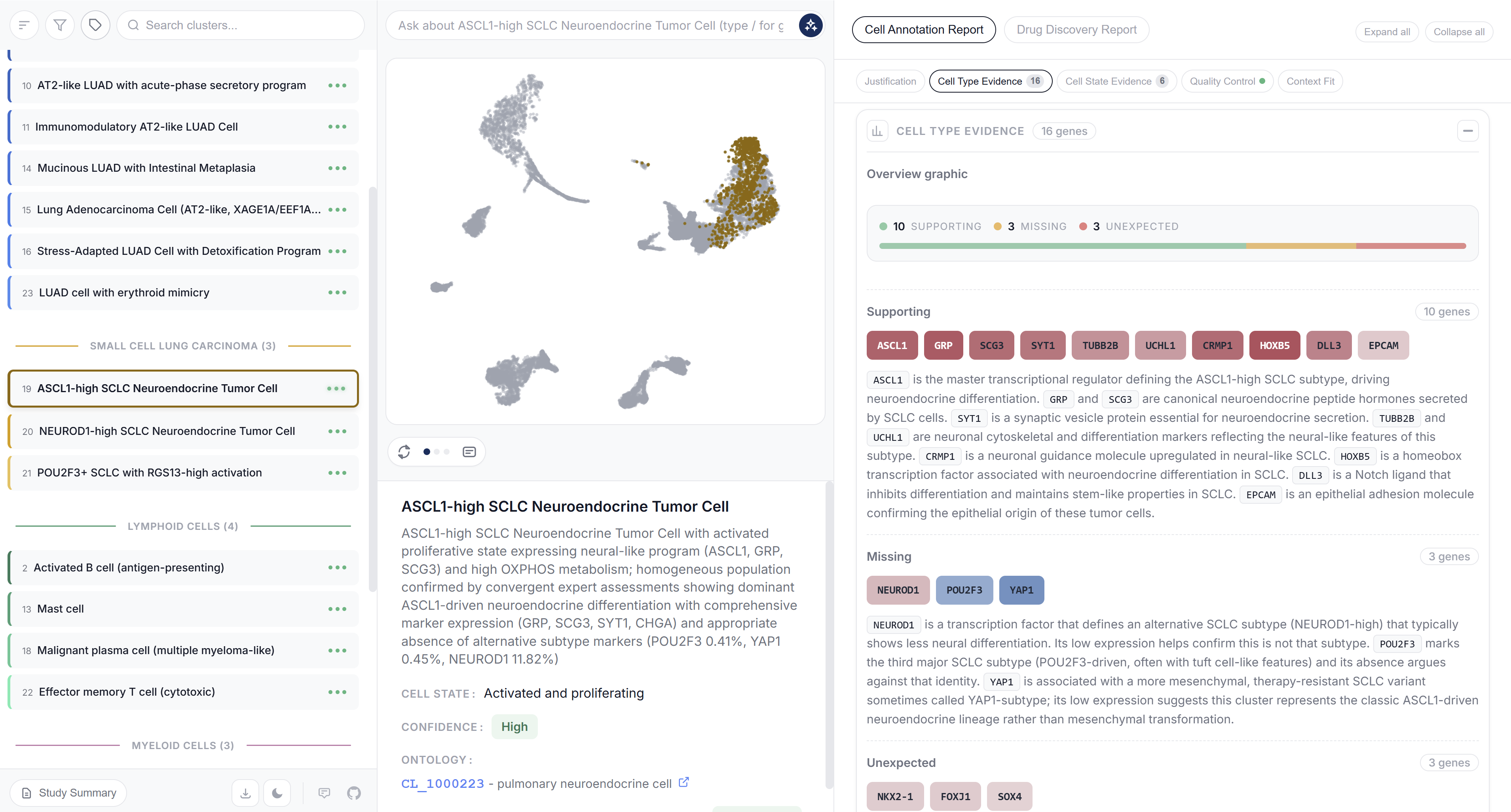
Output Reports
Each analysis generates an HTML report documenting annotation decisions, reviewer comments and an embedded chat interface for further exploration.
Benchmarks
Validated across PBMC, bone marrow, tumor microenvironment, and cross-species datasets. CyteType's agentic architecture consistently outperforms existing annotation methods:
| Comparison | Improvement |
|---|---|
| vs GPTCellType | +388% |
| vs CellTypist | +268% |
| vs SingleR | +101% |
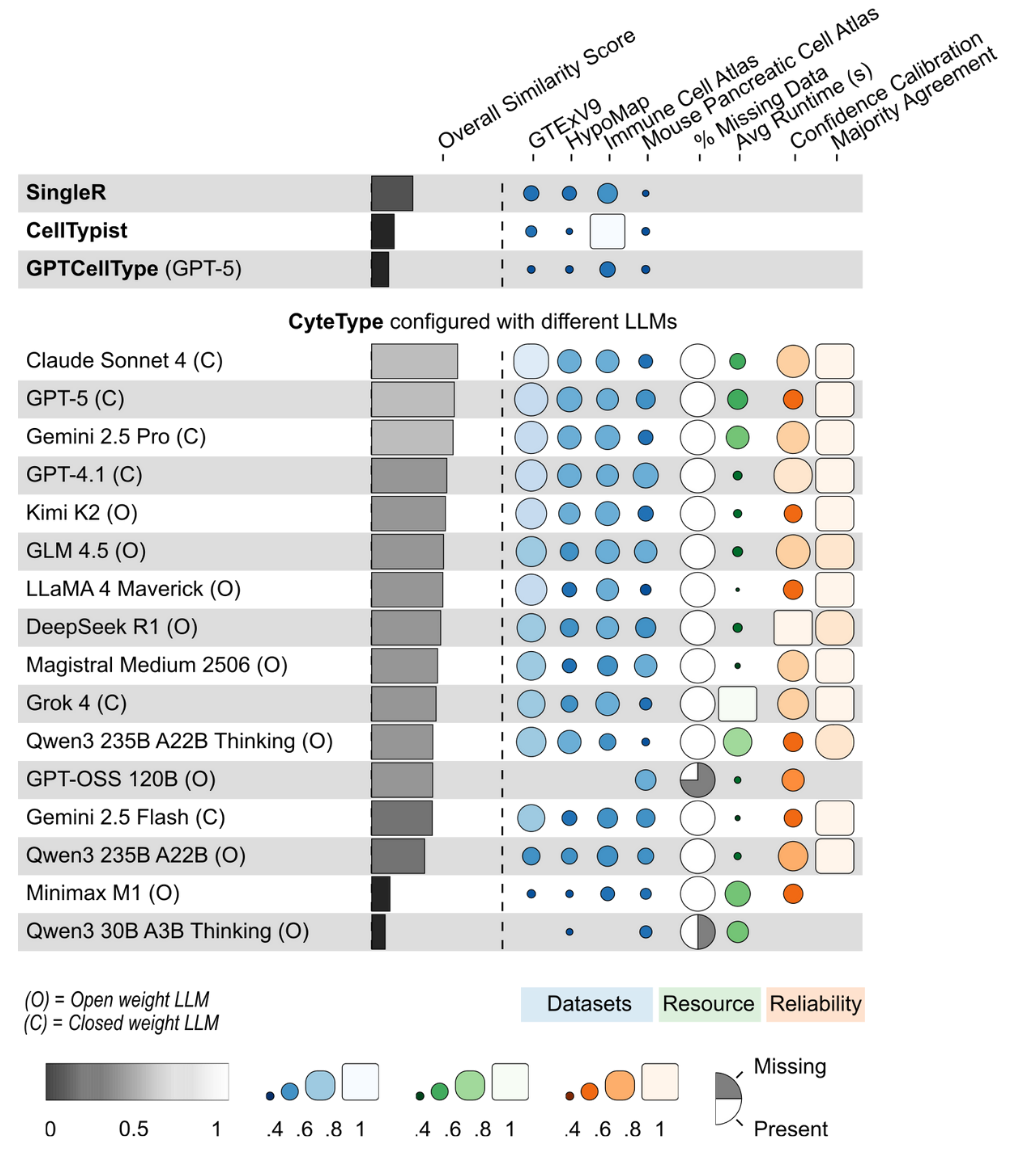

Browse CyteType results on atlas scale datasets
Citation
If you use CyteType in your research, please cite our preprint:
Ahuja G, Antill A, Su Y, Dall'Olio GM, Basnayake S, Karlsson G, Dhapola P. Multi-agent AI enables evidence-based cell annotation in single-cell transcriptomics. bioRxiv 2025. doi: 10.1101/2025.11.06.686964
@article{cytetype2025,
title={Multi-agent AI enables evidence-based cell annotation in single-cell transcriptomics},
author={Gautam Ahuja, Alex Antill, Yi Su, Giovanni Marco Dall'Olio, Sukhitha Basnayake, Göran Karlsson, Parashar Dhapola},
journal={bioRxiv},
year={2025},
doi={10.1101/2025.11.06.686964},
url={https://www.biorxiv.org/content/10.1101/2025.11.06.686964v1}
}
License
CyteType is free for academic and non-commercial research under CC BY-NC-SA 4.0.
For commercial licensing, contact contact@nygen.io.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file cytetype-0.19.3.tar.gz.
File metadata
- Download URL: cytetype-0.19.3.tar.gz
- Upload date:
- Size: 51.2 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
bd43b252fd88bd93183395161afbea25f0b3c69121d0eb3e738ffccb9a5158c6
|
|
| MD5 |
e0950bb9be6120c6cd41472059d6e797
|
|
| BLAKE2b-256 |
3ead63b80f81089cd2a5dd3dc03d4890665df068fa74392ef64e68d72a12687e
|
Provenance
The following attestation bundles were made for cytetype-0.19.3.tar.gz:
Publisher:
publish.yml on NygenAnalytics/CyteType
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
cytetype-0.19.3.tar.gz -
Subject digest:
bd43b252fd88bd93183395161afbea25f0b3c69121d0eb3e738ffccb9a5158c6 - Sigstore transparency entry: 1061823870
- Sigstore integration time:
-
Permalink:
NygenAnalytics/CyteType@e27bd80a4ff33ccc14a1e0f9fc1dd8799bc77a62 -
Branch / Tag:
refs/tags/0.19.3 - Owner: https://github.com/NygenAnalytics
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@e27bd80a4ff33ccc14a1e0f9fc1dd8799bc77a62 -
Trigger Event:
release
-
Statement type:
File details
Details for the file cytetype-0.19.3-py3-none-any.whl.
File metadata
- Download URL: cytetype-0.19.3-py3-none-any.whl
- Upload date:
- Size: 56.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
54aa982a88fccefed68787ff9b71879d3fa1eaa16fe63fd1746db5b1d093d244
|
|
| MD5 |
88031c969c59774fd8e5dc20f7cb3678
|
|
| BLAKE2b-256 |
3f910e207998c1aee01b8e0ba9ea2c5efe53f8daadc3155526c9806fea24ae6c
|
Provenance
The following attestation bundles were made for cytetype-0.19.3-py3-none-any.whl:
Publisher:
publish.yml on NygenAnalytics/CyteType
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
cytetype-0.19.3-py3-none-any.whl -
Subject digest:
54aa982a88fccefed68787ff9b71879d3fa1eaa16fe63fd1746db5b1d093d244 - Sigstore transparency entry: 1061823877
- Sigstore integration time:
-
Permalink:
NygenAnalytics/CyteType@e27bd80a4ff33ccc14a1e0f9fc1dd8799bc77a62 -
Branch / Tag:
refs/tags/0.19.3 - Owner: https://github.com/NygenAnalytics
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@e27bd80a4ff33ccc14a1e0f9fc1dd8799bc77a62 -
Trigger Event:
release
-
Statement type:
















