A manager of singularity containers and their singularity recipes (NGS applications)
Project description








- Python version:
Python 3.9, 3.10, 3.11, 3.12
- Source:
- Documentation:
- Issues:
- Platform:
Linux with bash, zsh, or fish shell
Overview

Damona is a conda-style package and environment manager built on top of Apptainer/Singularity containers. It lets you install bioinformatics (and other) tools as isolated containers, manage multiple versions side-by-side, and run them exactly like any other command-line tool — with no dependency conflicts and no root privileges required.
Think of Damona as conda for Singularity images: the same familiar create / activate / install workflow you already know, but with the rock-solid isolation and reproducibility that containers provide.
Why Damona?
Managing scientific software is notoriously painful:
Conda environments break when incompatible packages are installed together.
Raw Singularity/Apptainer requires verbose singularity exec invocations and manual management of image files.
Docker is unavailable or restricted on most HPC clusters.
Damona solves all three problems at once:
Feature |
Conda |
Singu- larity |
Damona |
|---|---|---|---|
Familiar install/activate workflow |
✔ |
✗ |
✔ |
Tools callable as plain commands |
✔ |
✗ |
✔ |
Full container isolation (no dep. conflicts) |
✗ |
✔ |
✔ |
No root required |
✔ |
✔ |
✔ |
Works on HPC/clusters without Docker |
✔ |
✔ |
✔ |
Multiple tool versions in separate envs |
✔ |
✗ |
✔ |
Images shared across environments |
✗ |
✗ |
✔ |
Central + custom registries |
✔ |
✗ |
✔ |
Key strengths at a glance
One command to install — damona install bwa downloads the container and creates a wrapper script so that bwa just works in your shell.
Zero dependency conflicts — every tool runs inside its own container, completely isolated from everything else on the system.
No root, no Docker — Apptainer/Singularity runs fully unprivileged, making Damona ideal for shared HPC clusters where Docker is not available.
Multiple versions, one system — need BWA 0.7.17 in one pipeline and BWA 0.7.18 in another? Create two Damona environments and switch instantly.
Images are shared — re-installing a tool in a second environment reuses the already-downloaded image, saving disk space and time.
Reproducible by design — pin exact versions in an environment file and export/import it to reproduce results anywhere.
Custom registries — host your own registry to distribute in-house containers to your team, just like a private conda channel.
Installation
Step 1 — Install Apptainer
Damona is a manager for Apptainer/Singularity images, so Apptainer must be present on your system first. Follow the official Apptainer installation guide, or — if you already use conda — install it with:
conda install -c conda-forge apptainer
Step 2 — Install Damona
pip install damona
Step 3 — Initialise Damona
Run damona once to create the configuration directory and shell helpers:
damona
Follow the on-screen instructions. To make the shell integration permanent, add one of the following lines to your shell start-up file:
bash — add to ~/.bashrc:
source ~/.config/damona/damona.sh
zsh — add to ~/.zshrc:
source ~/.config/damona/damona.zsh
fish — add to ~/.config/fish/config.fish:
source ~/.config/damona/damona.fish
Then open a new shell and run damona again. You should see the help screen:
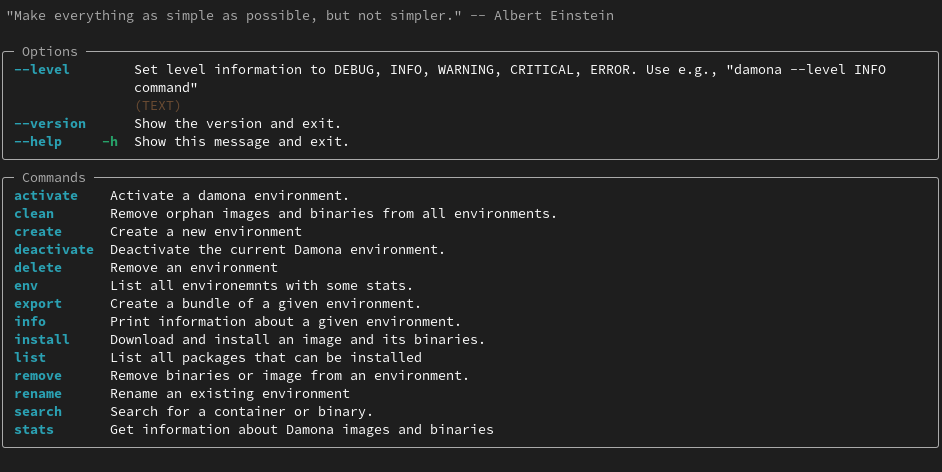
Quick Start
The full workflow takes under a minute:
# 1. Create a named environment
damona create TEST
# 2. Activate it (installed tools go here)
damona activate TEST
# 3. Install a tool — container + wrapper created automatically
damona install fastqc:0.11.9
# 4. Use it just like any other command
fastqc --help
# 5. Rename or remove the environment when you're done
damona rename TEST --new-name prod
damona remove prodFor more examples see the User Guide.
Example without conda (pyenv + minimap2)
If you manage Python with pyenv instead of conda, the workflow is identical — Damona only requires Python ≥ 3.9 and Apptainer.
# --- Prerequisites ---------------------------------------------------
# 1. Install Apptainer (once, system-wide or via your package manager).
# On Debian/Ubuntu:
sudo apt-get install -y apptainer
# Or follow https://apptainer.org/docs/admin/main/installation.html
# --- Python environment ----------------------------------------------
# 2. Create and activate a pyenv virtualenv with Python 3.10
pyenv install 3.10.14 # skip if already installed
pyenv virtualenv 3.10.14 damona-env
pyenv activate damona-env
# 3. Install Damona
pip install damona
# --- First-time initialisation ---------------------------------------
# 4. Run once to create the configuration directory and shell helpers
damona
# 5. Add the shell integration to your start-up file (bash example):
echo 'source ~/.config/damona/damona.sh' >> ~/.bashrc
source ~/.bashrc
# --- Install minimap2 ------------------------------------------------
# 6. Create and activate a Damona environment
damona create my-env
damona activate my-env
# 7. Install minimap2 — the container is downloaded automatically
damona install minimap2:2.24.0
# 8. Use it like any other command
minimap2 --versionMotivation
Conda/Bioconda is excellent for distributing pre-compiled scientific software, but dependency conflicts are a real-world problem: installing one package can silently break another, and rebuilding environments is time-consuming.
Singularity/Apptainer solves the isolation problem perfectly — each image is self-contained and reproducible — but using it directly requires verbose singularity exec commands and manual bookkeeping of image files and wrapper scripts.
Damona bridges the gap: it wraps Singularity images in the familiar conda-style environment model so that containers are as easy to install, activate, and use as conda packages, while retaining full container isolation and reproducibility.
Damona was originally developed for the Sequana project, but it is completely general-purpose and can be used to distribute any Singularity-compatible software.
Commands (Full CLI Reference)
Run damona --help to see all available commands:
Environment management: create Create a new environment remove Remove an environment and all its binaries. rename Rename an existing environment. env List all environments with their size and binary counts. activate Activate a damona environment. deactivate Deactivate the current Damona environment. Package management: install Download and install an image and its binaries into the active environment. uninstall Uninstall a binary or an image from an environment. clean Find and remove orphaned images and binaries across all environments. export Export an environment as a YAML file or a tar bundle. info Show images and binaries installed in an environment. Registry: search Search the registry for a container image or binary. list List all containers available in the local registry. stats Show registry statistics and local installation summary. Developer tools: check Check that all binaries in a built image are functional. build Build a Singularity image from a local recipe, a Damona recipe, or a Docker image. catalog Show a developer overview: latest version, size, and base image for every container.
For command-specific help (e.g. install):
damona install --help
1. List available environments
By default there is one environment called base. Unlike conda’s base, it is not essential and may be altered freely (but it cannot be removed or re-created). List all environments with:
damona env
2. Create environments
All environments are stored in ~/.config/damona/envs/. Create a new one:
damona create TEST
Verify it was created:
damona env
The last line should confirm that TEST is the current environment.
3. Activate and deactivate environments
Activating an environment appends its bin directory to your $PATH:
damona activate TEST damona env # confirms TEST is active
Deactivating removes it from $PATH:
damona deactivate TEST
Multiple environments can be active simultaneously; they follow a Last-In-First-Out order when deactivated without a name:
damona activate base damona activate TEST damona deactivate # removes TEST (last activated) damona deactivate base # removes base by name
4. Install a tool
With the target environment active, install any available package:
damona install fastqc:0.11.9
Damona downloads the Singularity image, registers it in ~/.config/damona/images (shared by all environments), and creates a wrapper script so that fastqc is available as a plain command.
5. Inspect an environment
List the binaries installed in an environment together with the underlying images:
damona info TEST
6. Search the registry
Search for available packages:
damona search PATTERN
Search an external registry (e.g. the official Damona registry):
damona search fastqc --url damona
The damona URL alias is pre-configured in ~/.config/damona/damona.cfg. You can add your own registry URLs there to distribute in-house containers.
7. Combine multiple environments
Images are shared across environments, so re-using an already-downloaded image in a new environment is instant and costs no extra disk space:
damona create test1 damona activate test1 damona install fastqc:0.11.9 # reuses the cached image
Activate several environments at once to mix tool sets:
damona activate base damona activate test1
For more details see the User Guide and the Developer Guide.
Contributors
Maintaining Damona would not have been possible without users and contributors. Each contribution has been an encouragement to pursue this project. Thanks to all:

Changelog
From version 0.16 onwards, we will not mention the new software and their versions but only changes made to the code itself. Entire list of software is available using the command:
damona list
Version |
Description |
|---|---|
0.19.1 |
|
0.19.0 |
|
0.18.0 |
|
0.17.2 |
|
0.17.1 |
|
0.17.0 |
|
0.16.0 |
|
0.15.2 |
|
0.15.1 |
|
0.15.0 |
|
0.14.7 |
|
0.14.6 |
|
0.14.5 |
|
0.14.4 |
|
0.14.3 |
|
0.14.2 |
|
0.14.1 |
|
0.14 |
|
0.13 |
|
0.12.3 |
|
0.12.2 |
|
0.12.1 |
|
0.12.0 |
|
0.11.1 |
|
0.11.0 |
|
0.10.1 |
|
0.10.0 |
|
0.9.1 |
|
0.9.0 |
|
0.8.4 |
|
0.8.3 |
|
0.8.2 |
|
0.8.1 |
|
0.8.0 |
|
0.7.1 |
|
0.7.0 |
|
0.6.0 |
|
0.5.3 |
|
0.5.2 |
|
0.5.1 |
|
0.5.0 |
|
0.4.3 |
|
0.4.2 |
|
0.4.1 |
|
0.4.0 |
|
0.3.X |
|
0.3.0 |
|
0.2.3 |
|
0.2.2 |
|
0.2.1 |
fixed manifest |
0.2.0 |
first working version of damona to pull image locally with binaries |
0.1.1 |
small update to fix RTD, travis, coveralls |
0.1 |
first release to test feasibility of the project |
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file damona-0.19.1.tar.gz.
File metadata
- Download URL: damona-0.19.1.tar.gz
- Upload date:
- Size: 989.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
748a3d3e014a43f884f09cd8342dd17172b665c657802189418bbf6f3052dbff
|
|
| MD5 |
bae5d47c1c0bf1b3d8e35c0a97defdcb
|
|
| BLAKE2b-256 |
c52c8d89a84a459b9f267784f67c4db0b1867b6dcc7d09bdb03ed7618f7ccb0a
|
File details
Details for the file damona-0.19.1-py3-none-any.whl.
File metadata
- Download URL: damona-0.19.1-py3-none-any.whl
- Upload date:
- Size: 1.2 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
02d09ba895d994422ae6483694a0e1cbc5669e96fb032122ab9d35ad4f04fa30
|
|
| MD5 |
33d368967f290ceb6f9287c5065b914c
|
|
| BLAKE2b-256 |
7d7e7df2abbc38748633ced3929405950e5abf75a5d8f3fc0c7ca880a94d2478
|











