Density–Diversity Coresets (DDC) for dataset summarization and distributional approximation
Project description
dd-coresets
Density–Diversity Coresets (DDC): a small weighted set of real data points that approximates the empirical distribution of a large dataset.
This library exposes a simple API (in the spirit of scikit-learn) to:
- build an unsupervised density–diversity coreset (
fit_ddc_coreset); - compare against random, stratified, and k-medoids baselines (
fit_random_coreset,fit_stratified_coreset,fit_kmedoids_coreset).
The goal is pragmatic: help data scientists work with large datasets using small, distribution-preserving subsets that are easy to simulate, visualise, and explain.
Motivation
The Problem
"We have 30M rows. We want 500 points + weights for EDA/simulation/stress testing."
Large datasets are ubiquitous in data science, but many workflows require small, interpretable subsets:
- Exploratory plots and dashboards need small, representative samples.
- Scenario analysis and simulations need few representative points with weights.
- Prototyping models is faster on coresets than on full data.
- Distributional stress testing requires faithful approximation of the empirical distribution.
The Solution: Distributional Fidelity
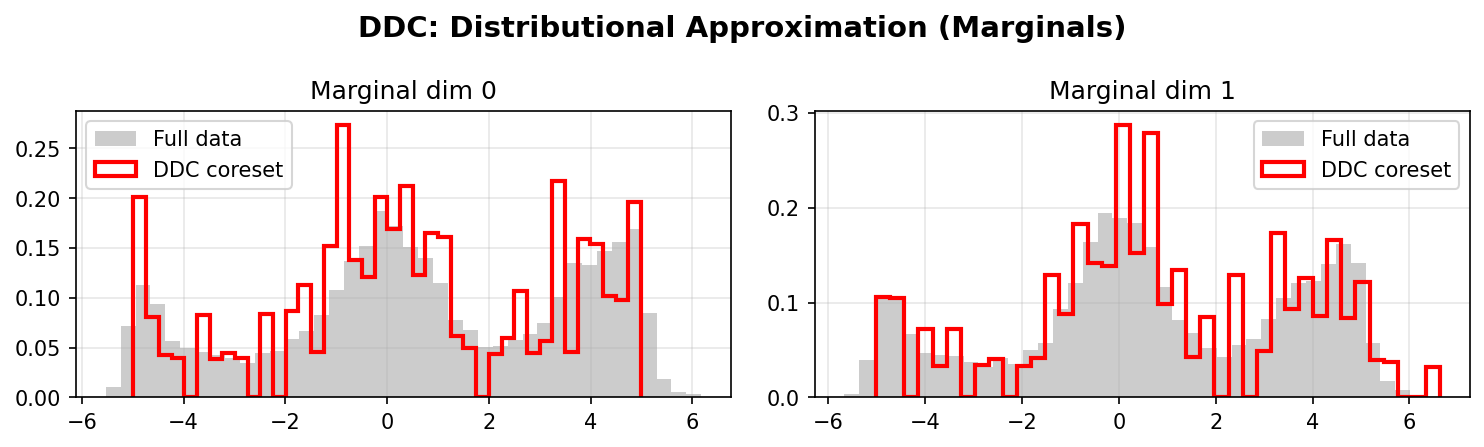
DDC selects real data points that preserve the distributional properties of your dataset. The weighted coreset closely matches the original distribution across all dimensions:
Left: Full data distribution (gray histogram) vs DDC weighted coreset (red outline).
Right: The coreset preserves marginal distributions, capturing modes, tails, and correlations.
Why Not Just Random Sampling?
Common approaches have limitations:
- Random sampling: Simple, but can miss important modes or tails, leading to biased estimates.
- Stratified sampling: Good when you already know the right strata (segments, classes, products), but needs domain knowledge and alignment with stakeholders.
- Cluster centroids (e.g., k-means): Minimize reconstruction error, but centroids are not real data points and are not explicitly distributional.
DDC sits in between:
- Unsupervised and geometry-aware (no domain knowledge required).
- Selects real points (medoids) that cover dense regions and diverse modes.
- Learns weights via soft assignments, approximating the empirical distribution.
When to Use / When Not to Use
- Use DDC when:
- You have a large tabular dataset and need a small, distribution-preserving subset.
- You want faithful mini-distributions for EDA, simulation, or stress testing.
- You don't yet know the right strata or segments (low-knowledge regime).
- You need real data points (not synthetic centroids) for interpretability.
- Your data has multiple modes, complex geometries, or non-convex structures.
- Don't use DDC when:
- You need strong theoretical guarantees (e.g., coreset-specific for Bayesian inference, Wasserstein DRO, or model-specific coresets).
- Your state space is non-Euclidean (e.g., graphs, strings, manifolds without a good embedding).
- You have well-defined strata that are aligned with your task (use
fit_stratified_coresetinstead). - You need deterministic, reproducible coresets (DDC uses randomness in initialization and working sample selection).
Complexity and Scale
High-level complexity: After a working sample of size n₀, the main costs are:
- O(n₀ log n₀): k-NN graph construction for density estimation
- O(k n₀ d): Greedy selection of k representatives
- O(n₀ k d): Soft assignment and weighting
- O(n k d): Optional reweighting step on the full dataset (if
reweight_full=True)
Practical scaling: For n₀ = 20,000, k = 500, d = 50:
- Working sample phase: ~1-2 seconds
- Full reweighting (if enabled): ~5-10 seconds for
n = 1M
The algorithm is sublinear in the full dataset size when n₀ << n, making it practical for very large datasets.
Relation to Existing Methods
-
Random / Stratified sampling: DDC is unsupervised and geometry-aware, while stratified requires known strata. DDC often outperforms random sampling by 2-3x on distributional metrics.
-
k-means / k-medoids: DDC selects real points (like k-medoids) but optimizes for distributional fidelity rather than reconstruction error. DDC better preserves tails, correlations, and non-convex structures.
-
Wasserstein / Sinkhorn coresets: DDC is a practical heuristic that scales to large datasets, while optimal Wasserstein coresets are computationally expensive. DDC approximates distributional properties without solving full optimal transport.
-
DPP / Diversity sampling: DDC balances density (covering modes) and diversity (covering the space), while DPP focuses primarily on diversity. DDC's density component helps preserve distributional properties.
Installation
From PyPI
pip install dd-coresets
From Source
git clone https://github.com/crbazevedo/dd-coresets.git
cd dd-coresets
python -m venv .venv
source .venv/bin/activate # Windows: .venv\Scripts\activate
pip install -r requirements.txt
Dependencies (minimal):
numpy >= 1.24scikit-learn >= 1.2matplotlib >= 3.7(for examples/plots)pandas >= 2.0(for examples)
Quickstart
1. Fit a DDC coreset (unsupervised default)
import numpy as np
from dd_coresets import fit_ddc_coreset
# X: (n, d) preprocessed features (e.g. scaled, encoded, etc.)
X = ... # load your data here
S, w, info = fit_ddc_coreset(
X,
k=200, # number of representatives
n0=20000, # working sample size (None = use all)
m_neighbors=32, # kNN for density
alpha=0.3, # density–diversity trade-off
gamma=1.0, # kernel scale
refine_iters=1, # medoid refinement iters
reweight_full=True,
random_state=42,
)
# S: (k, d) real data points
# w: (k,) non-negative, sum to 1
# info: metadata (indices, parameters, etc.)
print(S.shape, w.shape)
print(info.method, info.k, info.n, info.n0)
You can now use (S, w) for:
- simulation / scenario analysis,
- plotting weighted histograms or KDEs,
- approximate distributional comparisons.
2. Baselines for comparison
from dd_coresets import (
fit_random_coreset,
fit_stratified_coreset,
fit_kmedoids_coreset,
)
# Random coreset (no domain knowledge)
S_rnd, w_rnd, info_rnd = fit_random_coreset(
X,
k=200,
n0=20000,
gamma=1.0,
reweight_full=True,
random_state=0,
)
# K-medoids coreset (clustering-based)
S_kmed, w_kmed, info_kmed = fit_kmedoids_coreset(
X,
k=200,
n0=20000,
gamma=1.0,
reweight_full=True,
max_iters=10,
random_state=0,
)
# Stratified coreset (when you have strata)
# strata: 1D array, same length as X, e.g. segment, class, product line
strata = ... # e.g. y labels or business segments
S_strat, w_strat, info_strat = fit_stratified_coreset(
X,
strata=strata,
k=200,
n0=20000,
gamma=1.0,
reweight_full=True,
random_state=0,
)
Use these baselines to benchmark DDC on your data (moment errors, Wasserstein distances, etc.).
3. Adaptive distances & presets (new in v0.2.0)
from dd_coresets import fit_ddc_coreset
# Simple: use auto mode with balanced preset
S, w, info = fit_ddc_coreset(X, k=500, mode="auto", preset="balanced")
# Check what pipeline was used
print(info["pipeline"])
# {"mode": "auto", "preset": "balanced", "adaptive": True, "pca_used": False, ...}
When to use adaptive distances:
- d < 20: Euclidean (default, fastest)
- 20 ≤ d < 50: Adaptive if
m_neighbors > d, else PCA → Adaptive - d ≥ 50: PCA (20–50 dims) → Adaptive/Euclidean
Presets:
"fast": Quick runs (fewer neighbors, 1 iteration)"balanced": Default (good trade-off)"robust": More neighbors, 2 iterations (better quality)
Label-wise wrapper (preserve class proportions):
from dd_coresets import fit_ddc_coreset_by_label
# X: features, y: class labels
S, w, info = fit_ddc_coreset_by_label(
X, y, k_total=500, mode="auto", preset="balanced"
)
# Class proportions preserved
print(info["k_per_class"]) # [150, 200, 150] for 3 classes
Examples / Notebooks
We provide example notebooks demonstrating DDC usage:
-
examples/basic_tabular.ipynb(coming soon)- Installation and basic
fit_ddc_coresetusage - Comparison with
fit_random_coresetandfit_stratified_coreset - Distributional metrics (Wasserstein-1, Kolmogorov-Smirnov) and visualizations
- Installation and basic
-
examples/multimodal_2d.ipynb(coming soon)- 2D multimodal dataset (3 Gaussians + ring structure)
- Spatial coverage visualization
- Marginal distribution comparison with histograms
For now, see the experiment scripts in experiments/ for working examples.
API Overview
All functions assume X is a NumPy array of shape (n, d) with preprocessed numerical features (e.g. scaled, encoded, etc.).
fit_ddc_coreset
S, w, info = fit_ddc_coreset(
X,
k,
*,
n0=None, # default: 20000 (backward compatible)
alpha=0.3,
gamma=1.0,
refine_iters=1,
reweight_full=True,
random_state=None,
# New parameters (v0.2.0):
mode="euclidean", # "euclidean" | "adaptive" | "auto"
preset="balanced", # "fast" | "balanced" | "robust" | "manual"
distance_cfg=None, # override distance config (if preset="manual")
pipeline_cfg=None, # override pipeline config (if preset="manual")
)
-
Parameters
X:(n, d)array-like, preprocessed data.k: number of representatives.n0: working sample size. IfNone(default), uses 20000. If>= n, uses all data.alpha: density–diversity trade-off (0 ≈ diversity,1 ≈ density).gamma: kernel scale multiplier (used in soft assignment).refine_iters: medoid refinement iterations (usually 1 is enough).reweight_full: ifTrue, reweights using the full dataset; else uses only the working sample.random_state: RNG seed.mode: Distance mode."euclidean"(default, backward compatible),"adaptive"(Mahalanobis), or"auto"(choose based on d).preset: Configuration preset."fast","balanced"(default),"robust", or"manual"(use*_cfgdicts).distance_cfg: Dict to override distance config (e.g.,{"m_neighbors": 64, "iterations": 2}).pipeline_cfg: Dict to override pipeline config (e.g.,{"dim_threshold_adaptive": 40}).
-
Returns
S:(k, d)representatives (always in original feature space).w:(k,)weights (w >= 0,sum(w) = 1).info: Dict with metadata including:"pipeline":{"mode", "preset", "adaptive", "pca_used", "d_original", "d_effective", "fallbacks"}"config": Resolved distance and pipeline configs"pca": PCA info if used- Standard fields:
"method", "k", "n", "n0", "working_indices", "selected_indices", ...
Recommended use:
Default choice when you do not yet know which strata or labels matter. Good for EDA, exploratory simulation, and early-stage modelling.
fit_random_coreset
S, w, info = fit_random_coreset(
X,
k,
n0=20000,
gamma=1.0,
reweight_full=True,
random_state=None,
)
- Samples
kpoints uniformly from a working sample (sizen0) and applies the same soft-weighting scheme as DDC.
Use case:
Baseline to compare against DDC and stratified; reflects what many teams do today (simple downsampling).
fit_stratified_coreset
S, w, info = fit_stratified_coreset(
X,
strata,
k,
n0=20000,
gamma=1.0,
reweight_full=True,
random_state=None,
)
-
Parameters
X:(n, d)data.strata: 1D array of lengthnwith stratum labels (e.g. product, region, class, risk band).- Other parameters analogous to
fit_random_coreset.
-
Internally:
- Computes stratum frequencies on the working sample.
- Allocates
k_greps per stratum ∝ frequency. - Samples uniformly inside each stratum.
- Applies the same soft-weighting scheme as DDC.
Use case:
When you know the relevant strata and must preserve their proportions (regulatory reporting, risk/actuarial slices, business segments).
fit_ddc_coreset_by_label (new in v0.2.0)
from dd_coresets import fit_ddc_coreset_by_label
S, w, info = fit_ddc_coreset_by_label(
X,
y,
k_total,
**ddc_kwargs, # same as fit_ddc_coreset (mode, preset, etc.)
)
-
Parameters
X:(n, d)data.y:(n,)class labels (integers).k_total: Total number of representatives.**ddc_kwargs: All parameters fromfit_ddc_coreset(mode, preset, alpha, etc.).
-
Internally:
- Computes class proportions from
y. - Allocates
k_creps per class ∝ proportion. - Runs
fit_ddc_coresetseparately per class. - Concatenates and renormalizes weights.
- Computes class proportions from
-
Returns
S:(k_total, d)representatives.w:(k_total,)weights (sum to 1).info: Dict with{"classes", "k_per_class", "n_per_class", "info_per_class", ...}.
Use case:
When you have class labels and want to preserve class proportions while still benefiting from density–diversity within each class. Better than stratified for preserving distributional structure within classes.
fit_kmedoids_coreset
S, w, info = fit_kmedoids_coreset(
X,
k,
n0=20000,
gamma=1.0,
reweight_full=True,
max_iters=10,
random_state=None,
)
-
Parameters
X:(n, d)data.k: number of medoids (representatives).n0: working sample size. IfNoneor>= n, uses all data.gamma: kernel scale multiplier for soft assignments.reweight_full: ifTrue, reweights using the full dataset; else uses only the working sample.max_iters: maximum iterations for PAM-like swap optimization.random_state: RNG seed.
-
Internally:
- Uses k-means++ style initialization for medoids.
- Performs PAM-like swap optimization to minimize sum of distances to nearest medoid.
- Applies the same soft-weighting scheme as DDC.
Use case:
Clustering-based baseline that selects k real data points (medoids) minimizing within-cluster distances. Useful for comparison, but may struggle with non-convex structures.
Visual Examples
Example 1: 2D Multimodal Data (3 Gaussians + Ring)
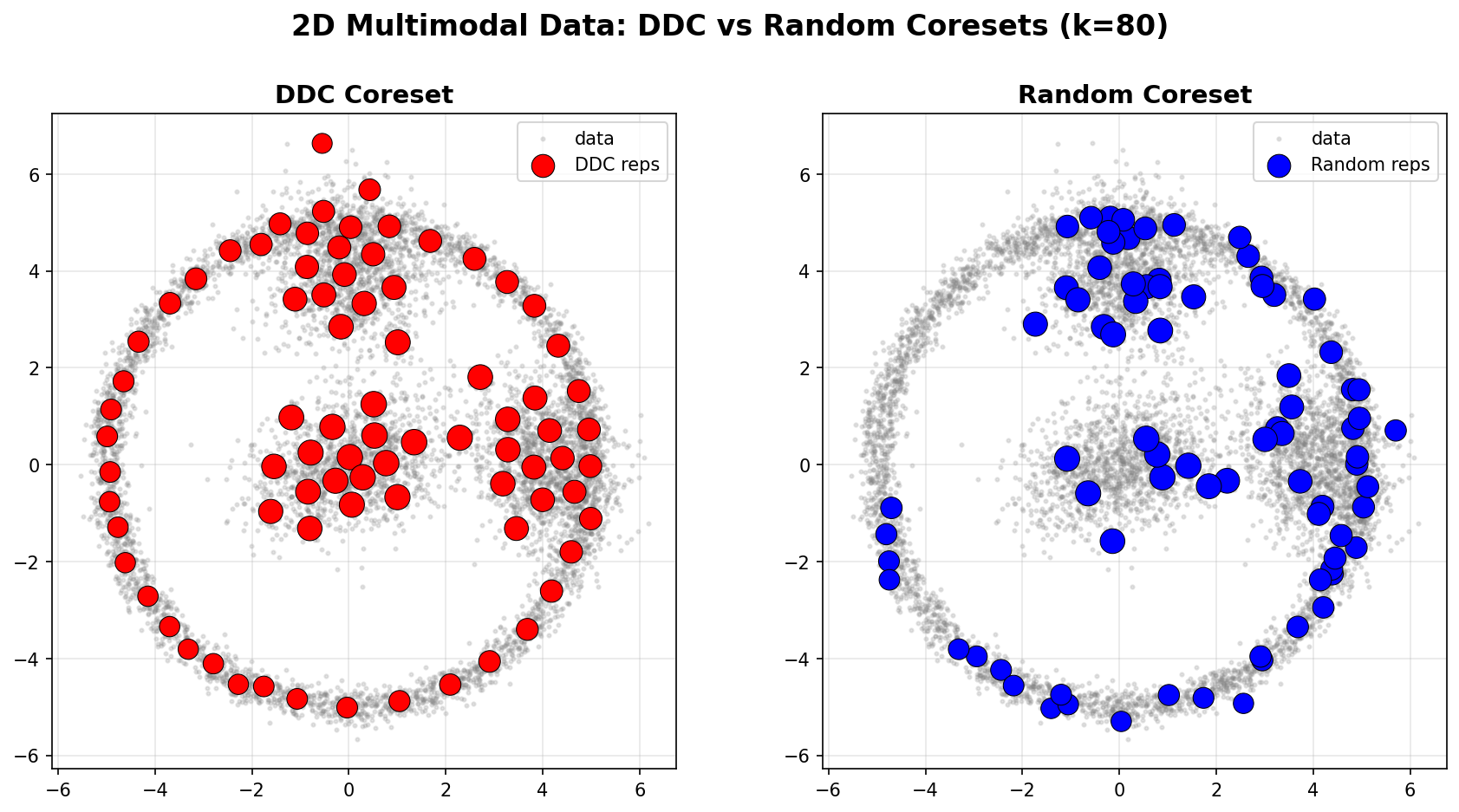
The following example demonstrates DDC on a 2D multimodal dataset (3 Gaussian blobs + a ring structure, n=8000). We compare DDC against random sampling with the same parameters (k=80, n0=None).
Spatial Coverage
Left (DDC): Representatives are strategically placed to cover:
- All three Gaussian modes (dense regions)
- The ring structure (diverse, low-density region)
- Points are weighted by their representativeness
Right (Random): Representatives are uniformly distributed, missing the ring structure and unevenly covering the modes.
Distributional Approximation
DDC Marginals:
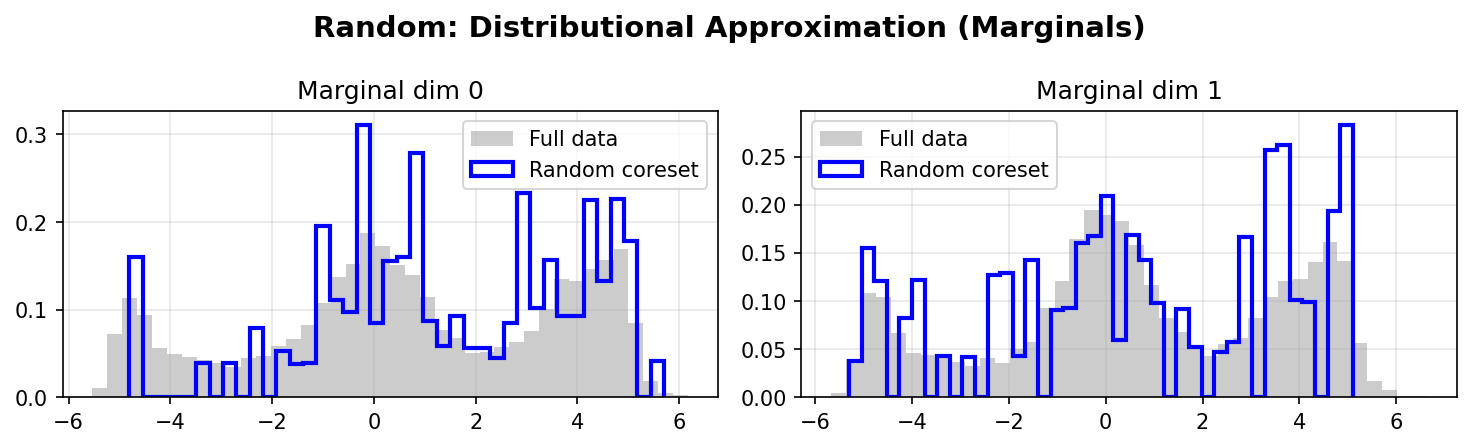
Random Marginals:
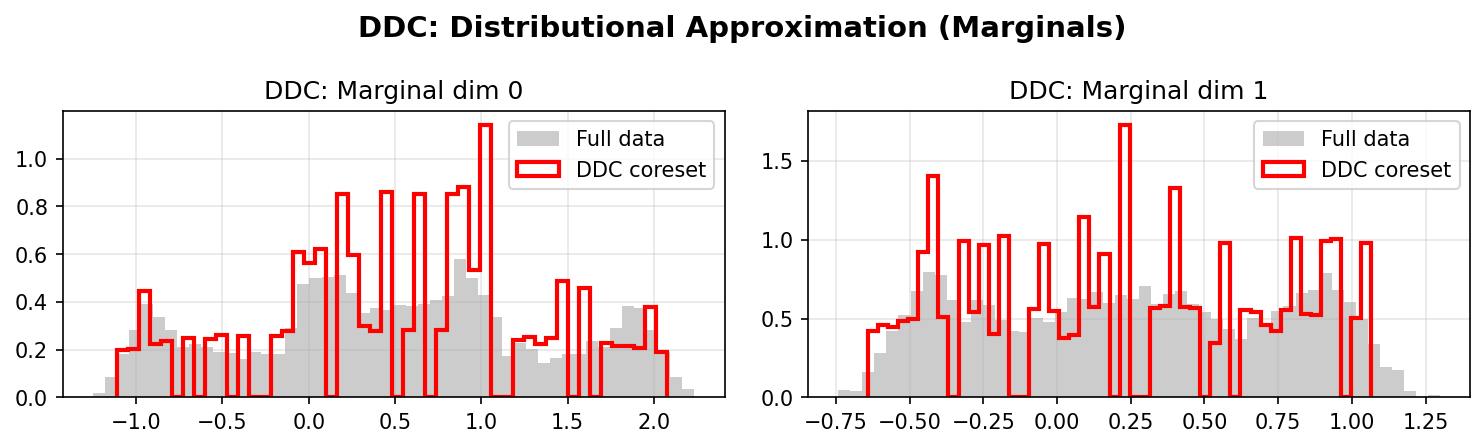
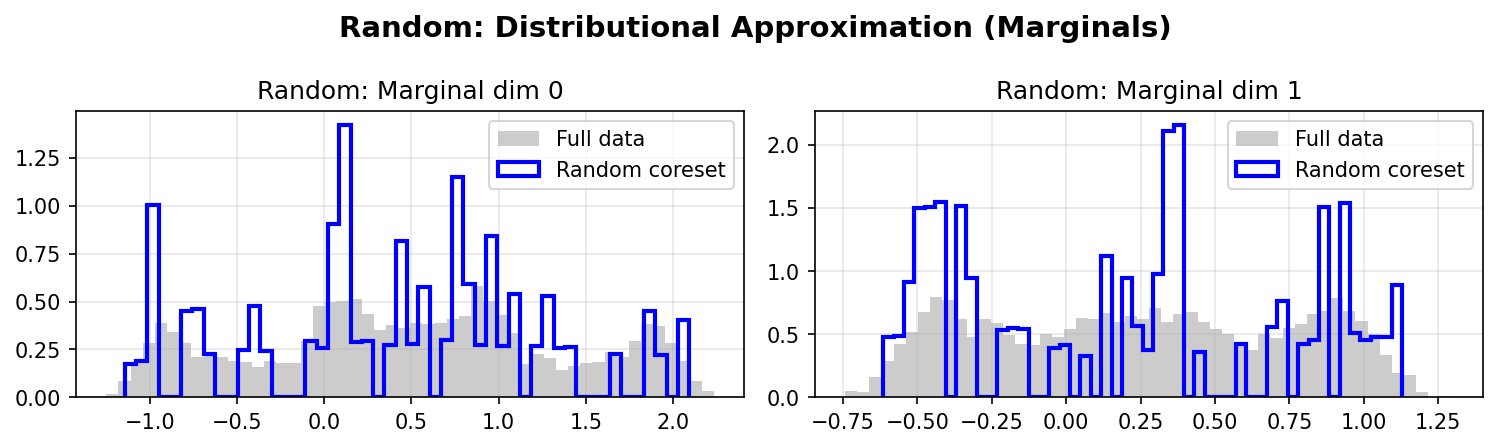
DDC better preserves the marginal distributions of the original data, especially in the tails and multimodal regions. The weighted coreset (red/blue lines) closely matches the full data distribution (gray histograms).
Quantitative Comparison
The following table compares DDC and Random coresets using standard distributional metrics:
| Method | Mean Error (L2) | Cov Error (Fro) | Corr Error (Fro) | W1 Mean | W1 Max | KS Mean | KS Max |
|---|---|---|---|---|---|---|---|
| DDC | 0.253 | 1.780 | 0.049 | 0.271 | 0.277 | 0.070 | 0.076 |
| Random | 0.797 | 2.486 | 0.080 | 0.515 | 0.806 | 0.098 | 0.138 |
Metrics explained:
- Mean Error (L2): L2 norm of the difference between full data mean and coreset weighted mean
- Cov Error (Fro): Frobenius norm of the difference between full data covariance and coreset weighted covariance
- Corr Error (Fro): Frobenius norm of the difference between correlation matrices
- W1 Mean/Max: Mean and maximum Wasserstein-1 distance across dimensions (lower is better)
- KS Mean/Max: Mean and maximum Kolmogorov-Smirnov statistic across dimensions (lower is better)
Key takeaway: DDC provides better spatial coverage and distributional fidelity than random sampling, especially when the data has multiple modes or complex geometries. Across all metrics, DDC achieves 2-3x lower error compared to random sampling.
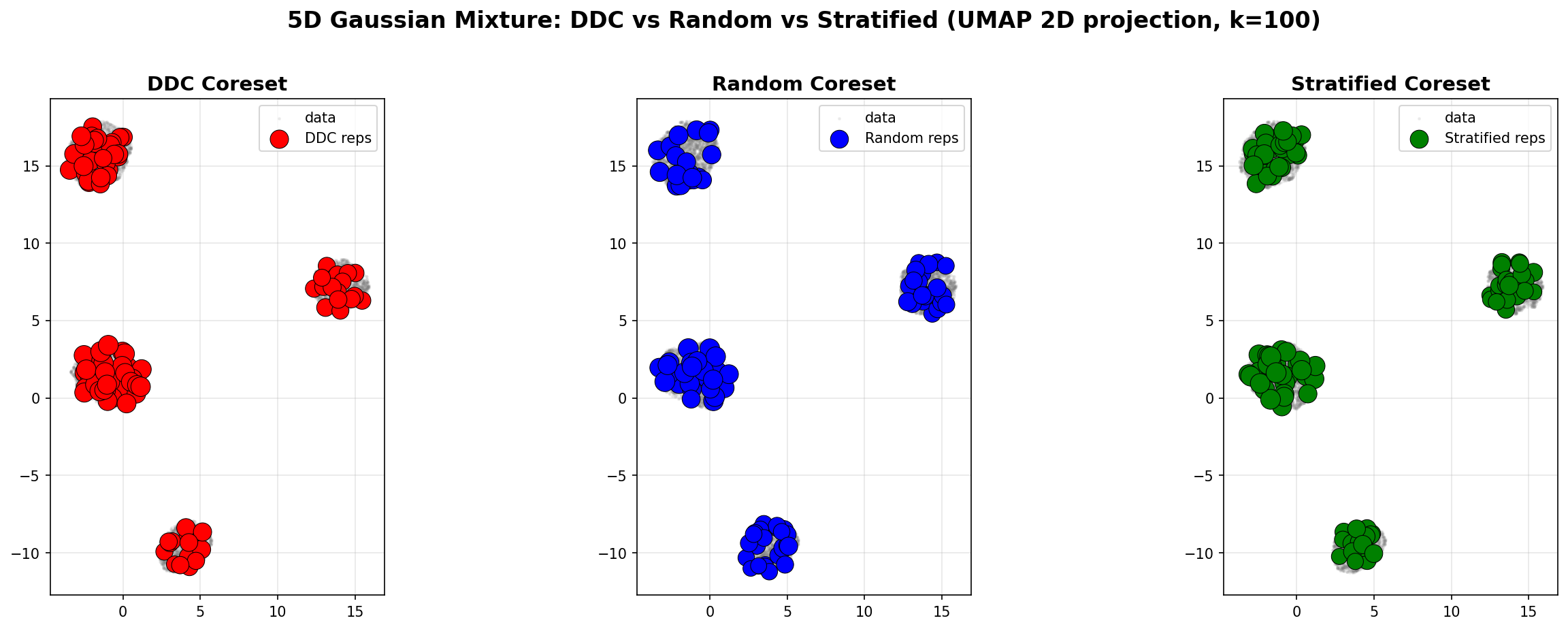
Example 2: 5D Gaussian Mixture
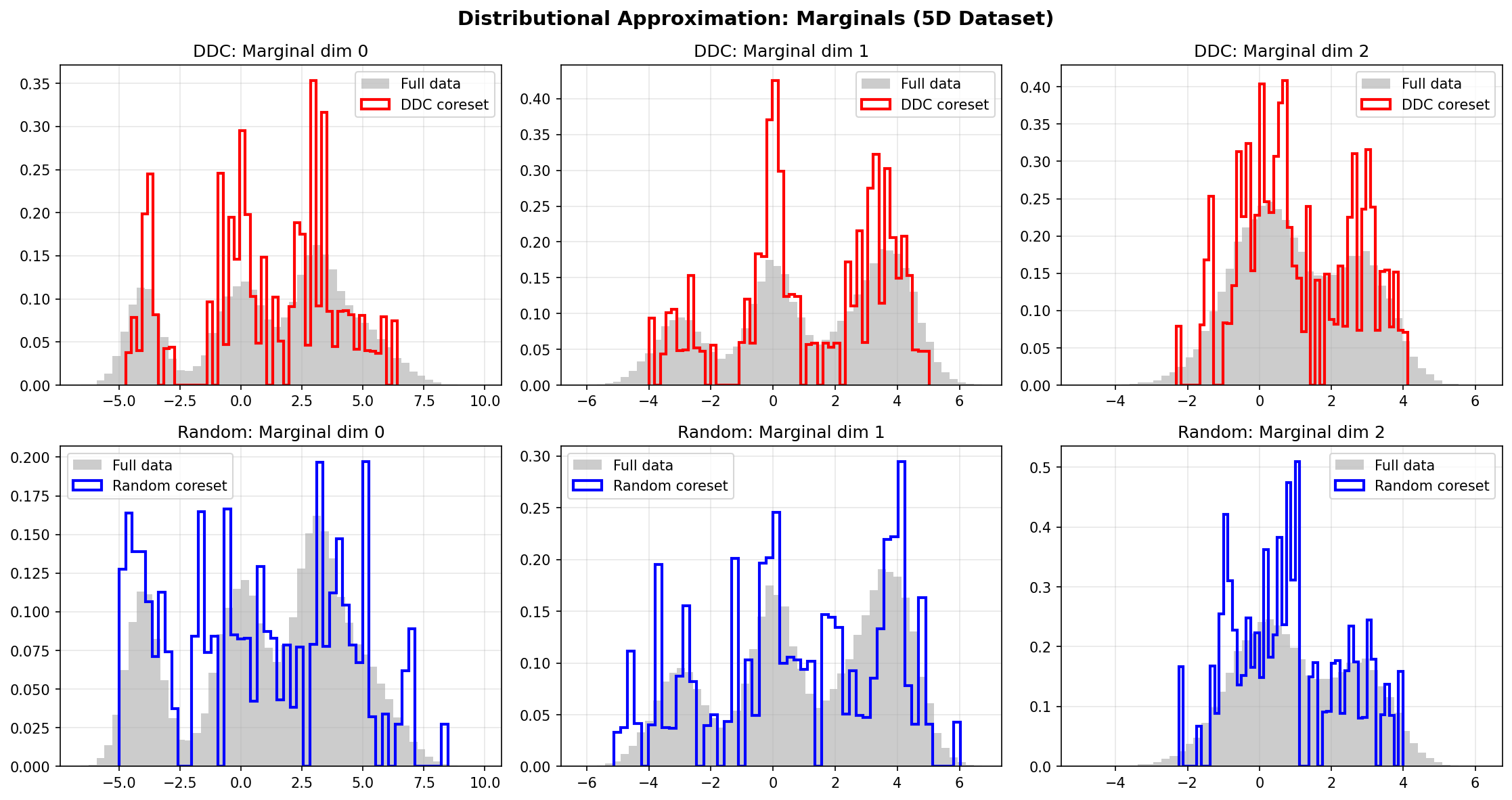
We also compare DDC, Random, and Stratified coresets on a 5D Gaussian mixture (4 components, n=50,000). The results are visualized using UMAP 2D projection and marginal distributions.
Spatial Coverage (UMAP 2D Projection)
Left (DDC): Representatives are distributed across all modes, capturing the mixture structure.
Middle (Random): Representatives are uniformly scattered, missing some modes.
Right (Stratified): Representatives respect component proportions, but may miss geometric structure.
Distributional Approximation
All three methods approximate the marginal distributions, with DDC and Stratified showing better fidelity to the full data distribution.
Quantitative Comparison
| Method | Mean Error (L2) | Cov Error (Fro) | Corr Error (Fro) | W1 Mean | W1 Max | KS Mean | KS Max |
|---|---|---|---|---|---|---|---|
| DDC | 0.174 | 4.197 | 0.530 | 0.251 | 0.418 | 0.073 | 0.090 |
| Random | 0.694 | 4.104 | 0.462 | 0.349 | 0.644 | 0.112 | 0.137 |
| Stratified | 0.315 | 2.708 | 0.246 | 0.213 | 0.361 | 0.063 | 0.080 |
Observations:
- DDC excels in mean approximation and Wasserstein distances (best W1 metrics).
- Stratified performs best on covariance and correlation (benefits from known component structure).
- Random shows highest errors across most metrics, confirming the value of structured sampling.
Takeaway: When component labels are available, Stratified can outperform DDC on moment-based metrics. However, DDC provides the best unsupervised performance and excels at distributional metrics (Wasserstein, KS).
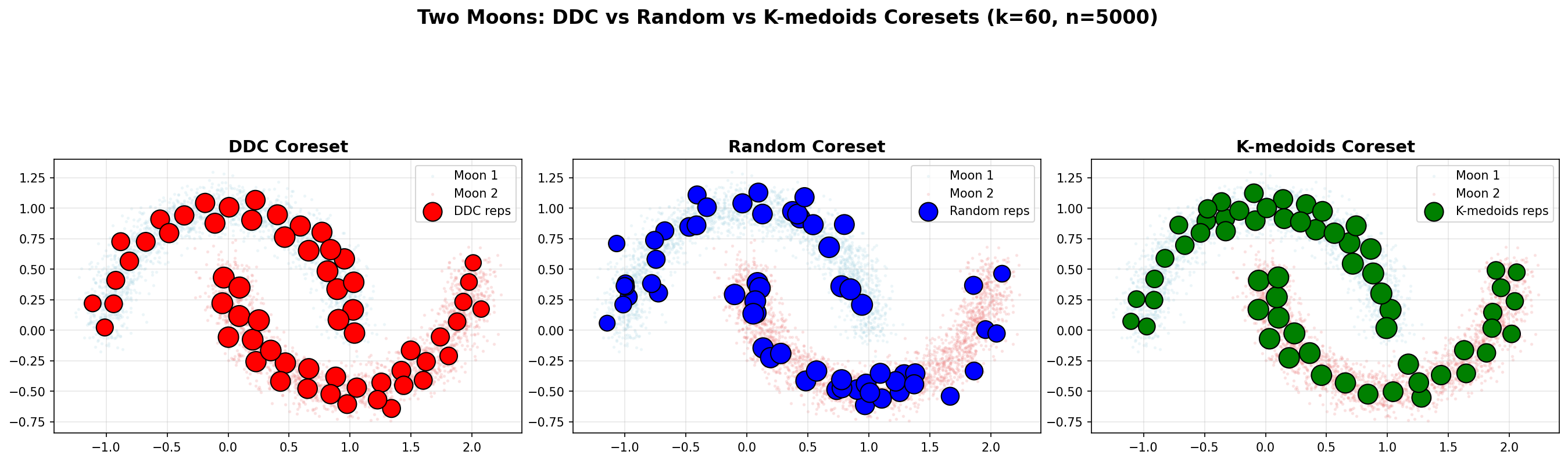
Example 3: Two Moons (Non-Convex Structure)
The Two Moons dataset demonstrates DDC's ability to handle non-convex structures. It consists of two interleaving half-circles (n=5000), creating a challenging geometry where random sampling and k-medoids often fail to connect both arcs.
Spatial Coverage
Left (DDC): Representatives are distributed along both arcs, maintaining connectivity and covering the non-convex structure.
Middle (Random): Representatives are scattered uniformly, potentially missing connections between the two moons and creating gaps.
Right (K-medoids): Representatives cluster around local centers, but may miss the connectivity between arcs due to the clustering objective.
Distributional Approximation
DDC Marginals:
Random Marginals:
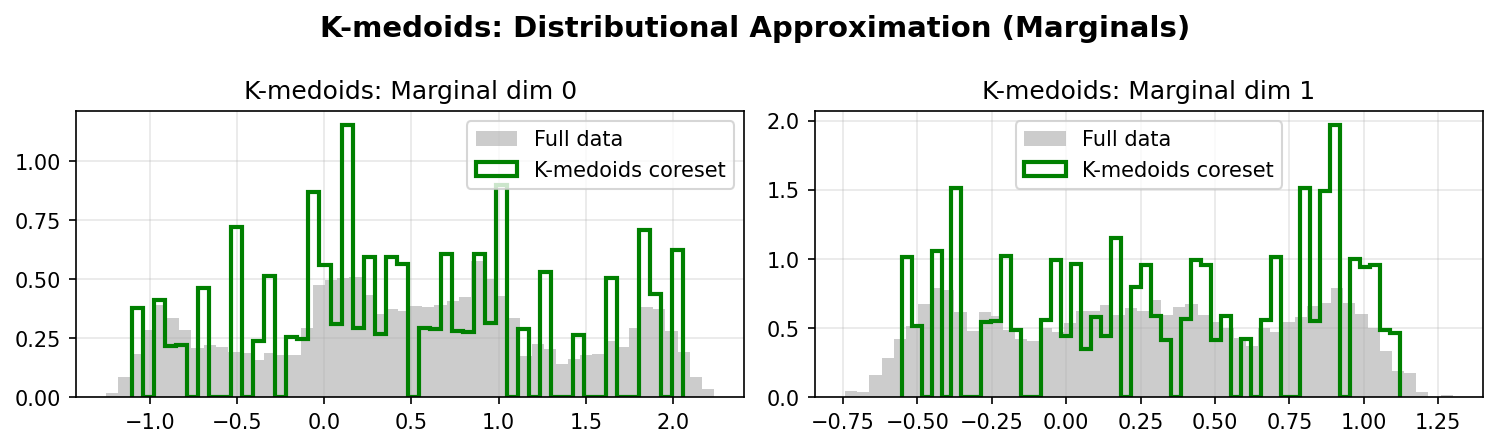
K-medoids Marginals:
Quantitative Comparison
| Method | Mean Error (L2) | Cov Error (Fro) | Corr Error (Fro) | W1 Mean | W1 Max | KS Mean | KS Max |
|---|---|---|---|---|---|---|---|
| DDC | 0.069 | 0.144 | 0.006 | 0.062 | 0.094 | 0.075 | 0.081 |
| Random | 0.100 | 0.109 | 0.069 | 0.087 | 0.102 | 0.117 | 0.132 |
| K-medoids | 0.103 | 0.077 | 0.004 | 0.091 | 0.112 | 0.092 | 0.112 |
Key observations:
- DDC achieves 1.4x lower mean error than Random and 1.5x lower than K-medoids.
- DDC shows 11.5x better correlation preservation than Random and 1.5x better than K-medoids.
- DDC demonstrates superior Wasserstein and KS metrics, indicating better distributional fidelity.
- K-medoids struggles with non-convex structures, as its clustering objective focuses on minimizing within-cluster distances rather than preserving global geometry.
Takeaway: DDC excels at preserving geometric structure in non-convex datasets, outperforming both random sampling and k-medoids. This makes it particularly valuable for complex manifolds and multimodal distributions.
Experiments
The repo includes three example scripts under experiments/:
-
synthetic_ddc_vs_baselines.py
5D Gaussian mixture (4 components, n=50,000):- DDC vs Random vs Stratified comparison,
- UMAP 2D visualization,
- metrics: mean/cov/corr errors, Wasserstein-1 marginals, KS.
-
multimodal_2d_ring_ddc.py
2D example (3 Gaussians + ring, n=8,000):- visual comparison DDC vs Random,
- shows how DDC covers multiple modes and a ring structure.
-
two_moons_ddc.py
2D Two Moons (non-convex structure, n=5,000):- demonstrates DDC's ability to handle non-convex geometries,
- DDC vs Random vs K-medoids comparison with quantitative metrics.
Run:
python experiments/synthetic_ddc_vs_baselines.py
python experiments/multimodal_2d_ring_ddc.py
python experiments/two_moons_ddc.py
When to Use What?
-
DDC (
fit_ddc_coreset):
Default in low-knowledge regimes (no clear strata yet). Better than random sampling for a fixedk. Best for unsupervised distributional approximation. -
Stratified (
fit_stratified_coreset):
Preferred in high-knowledge regimes (well-defined strata aligned with the task, e.g. risk bands, products), especially whenkis large enough. Can outperform DDC on moment-based metrics when strata are known. -
Random (
fit_random_coreset):
Baseline and sanity check; still useful when you want the simplest possible comparison. Reflects what many teams do today (simple downsampling). -
K-medoids (
fit_kmedoids_coreset):
Clustering-based baseline; useful for comparison but may struggle with non-convex geometries. Minimizes within-cluster distances rather than distributional fidelity.
License
MIT License. See LICENSE for details.
Citation
If you use dd-coresets in your research, please cite:
@software{dd-coresets,
title = {dd-coresets: Density–Diversity Coresets for Dataset Summarization},
author = {Azevedo, Carlos R. B.},
url = {https://github.com/crbazevedo/dd-coresets},
version = {0.1.3},
year = {2025}
}
Project details
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file dd_coresets-0.2.0.tar.gz.
File metadata
- Download URL: dd_coresets-0.2.0.tar.gz
- Upload date:
- Size: 32.6 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
2c71d56c58a348d6b4fea30eb43f4ad49c808870ce7177ea3b06e09c2effec02
|
|
| MD5 |
bc1754c77a74dc88c01b1da5626997b7
|
|
| BLAKE2b-256 |
169678dd9e1adc4be4509fc8a946e351de09c73af8590c15aaf8202395c8b410
|
Provenance
The following attestation bundles were made for dd_coresets-0.2.0.tar.gz:
Publisher:
publish-pypi.yml on crbazevedo/dd-coresets
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
dd_coresets-0.2.0.tar.gz -
Subject digest:
2c71d56c58a348d6b4fea30eb43f4ad49c808870ce7177ea3b06e09c2effec02 - Sigstore transparency entry: 701709630
- Sigstore integration time:
-
Permalink:
crbazevedo/dd-coresets@383ba3f0fe94248d9d09ecba15b1471fc4981179 -
Branch / Tag:
refs/tags/v0.2.0 - Owner: https://github.com/crbazevedo
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-pypi.yml@383ba3f0fe94248d9d09ecba15b1471fc4981179 -
Trigger Event:
release
-
Statement type:
File details
Details for the file dd_coresets-0.2.0-py3-none-any.whl.
File metadata
- Download URL: dd_coresets-0.2.0-py3-none-any.whl
- Upload date:
- Size: 19.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1189e3160b14eacc14796000eac6652d0a06e2918aba526915241b3c579525f5
|
|
| MD5 |
0c291611823d96a75e32b6c07f8fd72d
|
|
| BLAKE2b-256 |
8c570e0b1cbbdee9c4b3fc822b3e3ee13e0d5212fbcbac6ca3e3379e90ff6142
|
Provenance
The following attestation bundles were made for dd_coresets-0.2.0-py3-none-any.whl:
Publisher:
publish-pypi.yml on crbazevedo/dd-coresets
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
dd_coresets-0.2.0-py3-none-any.whl -
Subject digest:
1189e3160b14eacc14796000eac6652d0a06e2918aba526915241b3c579525f5 - Sigstore transparency entry: 701709638
- Sigstore integration time:
-
Permalink:
crbazevedo/dd-coresets@383ba3f0fe94248d9d09ecba15b1471fc4981179 -
Branch / Tag:
refs/tags/v0.2.0 - Owner: https://github.com/crbazevedo
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish-pypi.yml@383ba3f0fe94248d9d09ecba15b1471fc4981179 -
Trigger Event:
release
-
Statement type: