Identification of spatial domains in spatial transcriptomics by deep learning.
Project description
Identification of spatial domains in spatial transcriptomics by deep learning
Update
 May 28, 2025
May 28, 2025
(1) Updated the installation method for DeepST.
(2) Fixed some bugs.
Overview
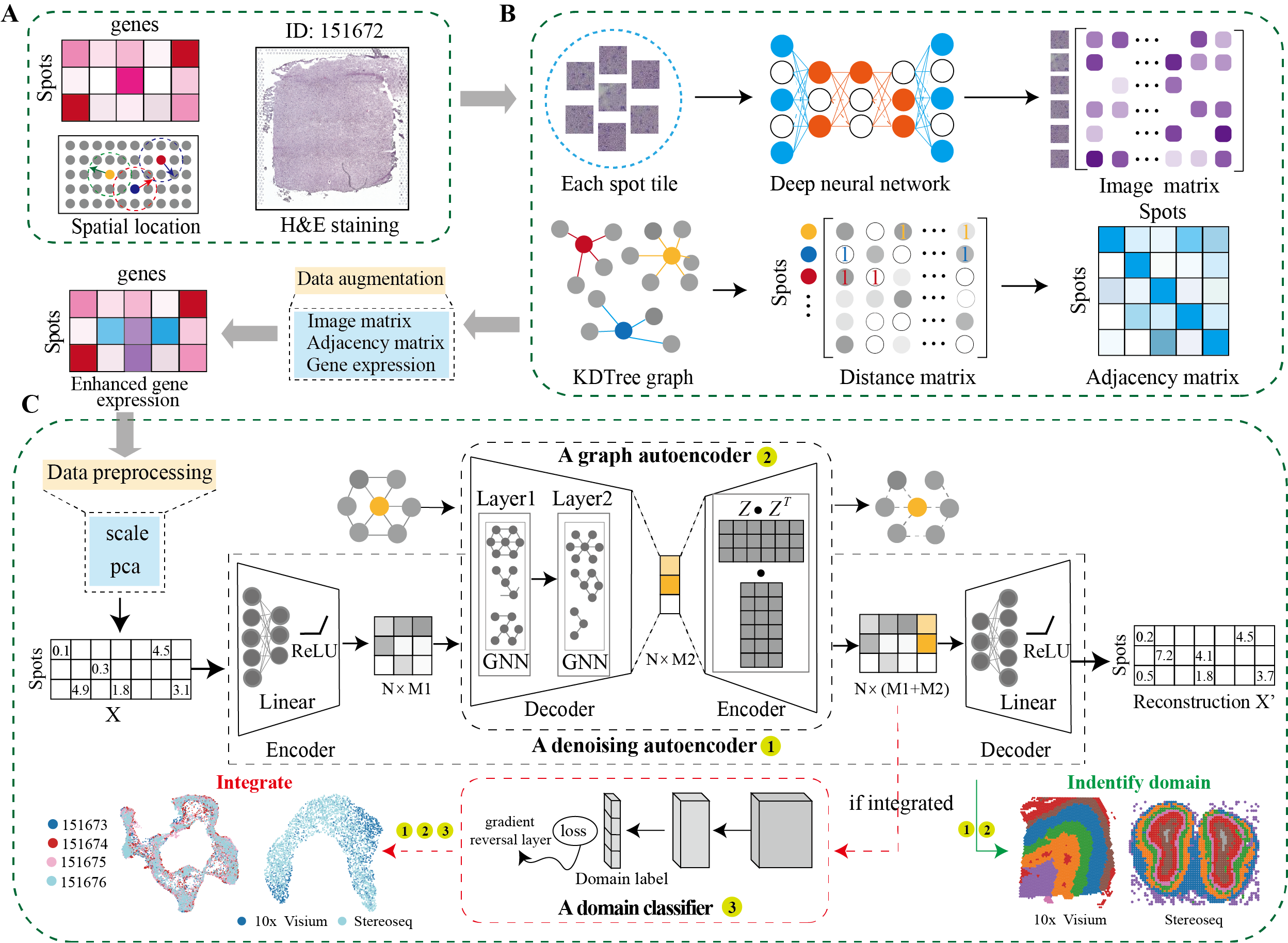
DeepST first uses H&E staining to extract tissue morphology information through a pre-trained deep learning model, and normalizes each spot’s gene expression according to the similarity of adjacent spots. DeepST further learns a spatial adjacency matrix on spatial location for the construction of graph convolutional network. DeepST uses a graph neural network autoencoder and a denoising autoencoder to jointly generate a latent representation of augmented ST data, while domain adversarial neural networks (DAN) are used to integrate ST data from multi-batches or different technologies. The output of DeepST can be applied to identify spatial domains, batch effect correction and downstream analysis.
How to install DeepST
To install DeepST, make sure you have PyTorch and PyG installed. For more details on dependencies, refer to the environment.yml file.
Step 1: Set Up Conda Environment
conda create -n deepst-env python=3.9
Step 2: Install PyTorch and PyG
Activate the environment and install PyTorch and PyG. Adjust the installation commands based on your CUDA version or choose the CPU version if necessary.
- General Installation Command
conda activate deepst-env
pip install torch==2.1.0 torchvision==0.16.0 torchaudio==2.1.0 --index-url https://download.pytorch.org/whl/cu118
pip install pyg_lib==0.3.1+pt21cu118 torch_scatter torch_sparse torch_cluster torch_spline_conv -f https://data.pyg.org/whl/torch-2.1.0+cu118.html
- Tips for selecting the correct CUDA version
- Run the following command to verify CUDA version:
nvcc --version- Alternatively, use:
nvidia-smi - Modify installation commands based on CUDA
- For CUDA 12.1
pip install torch==2.1.0 torchvision==0.16.0 torchaudio==2.1.0 --index-url https://download.pytorch.org/whl/cu121 pip install pyg_lib==0.3.1+pt21cu121 torch_scatter torch_sparse torch_cluster torch_spline_conv -f https://data.pyg.org/whl/torch-2.1.0+cu121.html - For CPU-only
pip install torch==2.1.0 torchvision==0.16.0 torchaudio==2.1.0 --index-url https://download.pytorch.org/whl/cpu pip install pyg_lib==0.3.1+pt21cpu torch_scatter torch_sparse torch_cluster torch_spline_conv -f https://data.pyg.org/whl/torch-2.1.0+cpu.html
- For CUDA 12.1
Step 3: Install dirac from shell
pip install sodeepst
Step 4: Import DIRAC in your jupyter notebooks or/and scripts
import sodeepst as dt
Quick Start
-
DeepST on DLPFC from 10x Visium.
import os
from DeepST import run
import matplotlib.pyplot as plt
from pathlib import Path
import scanpy as sc
data_path = "../data/DLPFC" #### to your path
data_name = '151673' #### project name
save_path = "../Results" #### save path
n_domains = 7 ###### the number of spatial domains.
deepen = run(save_path = save_path,
task = "Identify_Domain", #### DeepST includes two tasks, one is "Identify_Domain" and the other is "Integration"
pre_epochs = 800, #### choose the number of training
epochs = 1000, #### choose the number of training
use_gpu = True)
###### Read in 10x Visium data, or user can read in themselves.
adata = deepen._get_adata(platform="Visium", data_path=data_path, data_name=data_name)
###### Segment the Morphological Image
adata = deepen._get_image_crop(adata, data_name=data_name)
###### Data augmentation. spatial_type includes three kinds of "KDTree", "BallTree" and "LinearRegress", among which "LinearRegress"
###### is only applicable to 10x visium and the remaining omics selects the other two.
###### "use_morphological" defines whether to use morphological images.
adata = deepen._get_augment(adata, spatial_type="LinearRegress", use_morphological=True)
###### Build graphs. "distType" includes "KDTree", "BallTree", "kneighbors_graph", "Radius", etc., see adj.py
graph_dict = deepen._get_graph(adata.obsm["spatial"], distType = "BallTree")
###### Enhanced data preprocessing
data = deepen._data_process(adata, pca_n_comps = 200)
###### Training models
deepst_embed = deepen._fit(
data = data,
graph_dict = graph_dict,)
###### DeepST outputs
adata.obsm["DeepST_embed"] = deepst_embed
###### Define the number of space domains, and the model can also be customized. If it is a model custom priori = False.
adata = deepen._get_cluster_data(adata, n_domains=n_domains, priori = True)
###### Spatial localization map of the spatial domain
sc.pl.spatial(adata, color='DeepST_refine_domain', frameon = False, spot_size=150)
plt.savefig(os.path.join(save_path, f'{data_name}_domains.pdf'), bbox_inches='tight', dpi=300)
-
DeepST integrates data from mutil-batches or different technologies.
import os
from DeepST import run
import matplotlib.pyplot as plt
from pathlib import Path
import scanpy as sc
data_path = "../data/DLPFC"
data_name_list = ['151673', '151674', '151675', '151676']
save_path = "../Results"
n_domains = 7
deepen = run(save_path = save_path,
task = "Integration",
pre_epochs = 800,
epochs = 1000,
use_gpu = True,
)
###### Generate an augmented list of multiple datasets
augement_data_list = []
graph_list = []
for i in range(len(data_name_list)):
adata = deepen._get_adata(platform="Visium", data_path=data_path, data_name=data_name_list[i])
adata = deepen._get_image_crop(adata, data_name=data_name_list[i])
adata = deepen._get_augment(adata, spatial_type="LinearRegress")
graph_dict = deepen._get_graph(adata.obsm["spatial"], distType = "KDTree")
augement_data_list.append(adata)
graph_list.append(graph_dict)
######## Synthetic Datasets and Graphs
multiple_adata, multiple_graph = deepen._get_multiple_adata(adata_list = augement_data_list, data_name_list = data_name_list, graph_list = graph_list)
###### Enhanced data preprocessing
data = deepen._data_process(multiple_adata, pca_n_comps = 200)
deepst_embed = deepen._fit(
data = data,
graph_dict = multiple_graph,
domains = multiple_adata.obs["batch"].values, ##### Input to Domain Adversarial Model
n_domains = len(data_name_list))
multiple_adata.obsm["DeepST_embed"] = deepst_embed
multiple_adata = deepen._get_cluster_data(multiple_adata, n_domains=n_domains, priori = True)
sc.pp.neighbors(multiple_adata, use_rep='DeepST_embed')
sc.tl.umap(multiple_adata)
sc.pl.umap(multiple_adata, color=["DeepST_refine_domain","batch_name"])
plt.savefig(os.path.join(save_path, f'{"_".join(data_name_list)}_umap.pdf'), bbox_inches='tight', dpi=300)
for data_name in data_name_list:
adata = multiple_adata[multiple_adata.obs["batch_name"]==data_name]
sc.pl.spatial(adata, color='DeepST_refine_domain', frameon = False, spot_size=150)
plt.savefig(os.path.join(save_path, f'{data_name}_domains.pdf'), bbox_inches='tight', dpi=300)
-
DeepST works on other spatial omics data.
import os
from DeepST import run
import matplotlib.pyplot as plt
from pathlib import Path
import scanpy as sc
data_path = "../data"
data_name = 'Stereoseq'
save_path = "../Results"
n_domains = 15
deepen = run(save_path = save_path,
task = "Identify_Domain",
pre_epochs = 800,
epochs = 1000,
use_gpu = True)
###### Read in other spatial data, or user can read in themselves. Including original expression
###### information and spatial location information, where the location information is saved in .obsm["spatial"]
adata = deepen._get_adata(platform="Stereoseq", data_path=data_path, data_name=data_name)
###### Data augmentation. spatial_type includes three kinds of "KDTree", "BallTree" and "LinearRegress", among which "LinearRegress"
###### is only applicable to 10x visium and the remaining omics selects the other two.
###### "use_morphological" defines whether to use morphological images.
adata = deepen._get_augment(adata, spatial_type="BallTree", use_morphological=False)
###### Build graphs. "distType" includes "KDTree", "BallTree", "kneighbors_graph", "Radius", etc., see adj.py
graph_dict = deepen._get_graph(adata.obsm["spatial"], distType = "BallTree")
###### Enhanced data preprocessing
data = deepen._data_process(adata, pca_n_comps = 200)
###### Training models
deepst_embed = deepen._fit(
data = data,
graph_dict = graph_dict,)
###### DeepST outputs
adata.obsm["DeepST_embed"] = deepst_embed
###### Define the number of space domains, and the model can also be customized. If it is a model custom priori = False.
adata = deepen._get_cluster_data(adata, n_domains=n_domains, priori = True)
###### Spatial localization map of the spatial domain
sc.pl.spatial(adata, color='DeepST_refine_domain', frameon = False, spot_size=150)
plt.savefig(os.path.join(save_path, f'{data_name}_domains.pdf'), bbox_inches='tight', dpi=300)
Compared tools
Tools that are compared include:
Download data
| Platform | Tissue | SampleID |
|---|---|---|
| 10x Visium | Human dorsolateral pre-frontal cortex (DLPFC) | 151507, 151508, 151509, 151510, 151669, 151670, 151671, 151672, 151673, 151674, 151675, 151676 |
| 10x Visium | Mouse brain section | Coronal, Sagittal-Anterior, Sagittal-Posterior |
| 10x Visium | Human breast cancer | Invasive Ductal Carcinoma breast, Ductal Carcinoma In Situ & Invasive Carcinoma |
| Stereo-Seq | Mouse olfactory bulb | Olfactory bulb |
| Slide-seq | Mouse hippocampus | Coronal |
| MERFISH | Mouse brain slice | Hypothalamic preoptic region |
Spatial transcriptomics data of other platforms can be downloaded https://www.spatialomics.org/SpatialDB/
Contact
Feel free to submit an issue or contact us at xuchang0214@163.com for problems about the packages.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file deepstkit-2.0.2.tar.gz.
File metadata
- Download URL: deepstkit-2.0.2.tar.gz
- Upload date:
- Size: 29.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
bfeb47116e340375c1da90ab293c4219078c8d1edf0536bfa9cfa256d3a32e3c
|
|
| MD5 |
6e7f9448242f2fe4ede44be769ff08b9
|
|
| BLAKE2b-256 |
db1cfb8e6538772acc72e6bd4007df61f17208802c5c661753cc0b54a10241e8
|
File details
Details for the file deepstkit-2.0.2-py3-none-any.whl.
File metadata
- Download URL: deepstkit-2.0.2-py3-none-any.whl
- Upload date:
- Size: 30.1 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
150b9f0ca6772876f5b2ded73c578f467ac366fdcb4659951a887f2c6bf1a30f
|
|
| MD5 |
ad8d14a0f84b4badeb09f2b2aeebfd92
|
|
| BLAKE2b-256 |
f3d14bda3e1d77a5337952b94f05737a68cc4828a3a79cf46ca3e3fcda8517d8
|