An Evolutionary-scale Model (ESM) for protein function prediction from amino acid sequences using the Gene Ontology (GO).
Project description
ESMC Protein Function Predictor
An Evolutionary-scale Model (ESM) for protein function prediction from amino acid sequences using the Gene Ontology (GO). Based on the ESM Cambrian Transformer architecture, pre-trained on UniRef, MGnify, and the Joint Genome Institute's database and fine-tuned on the AmiGO Boost protein function dataset, this protein language model predicts the GO subgraph for a particular protein sequence - giving you insight into the molecular function, biological process, and location of the activity inside the cell.
Key Features
-
Sequence-to-function prediction — Predicts Molecular Function, Biological Process, and Cellular Component ontologies directly from raw amino acid sequences, eliminating the need for homology searches, structural data, or multiple sequence alignments.
-
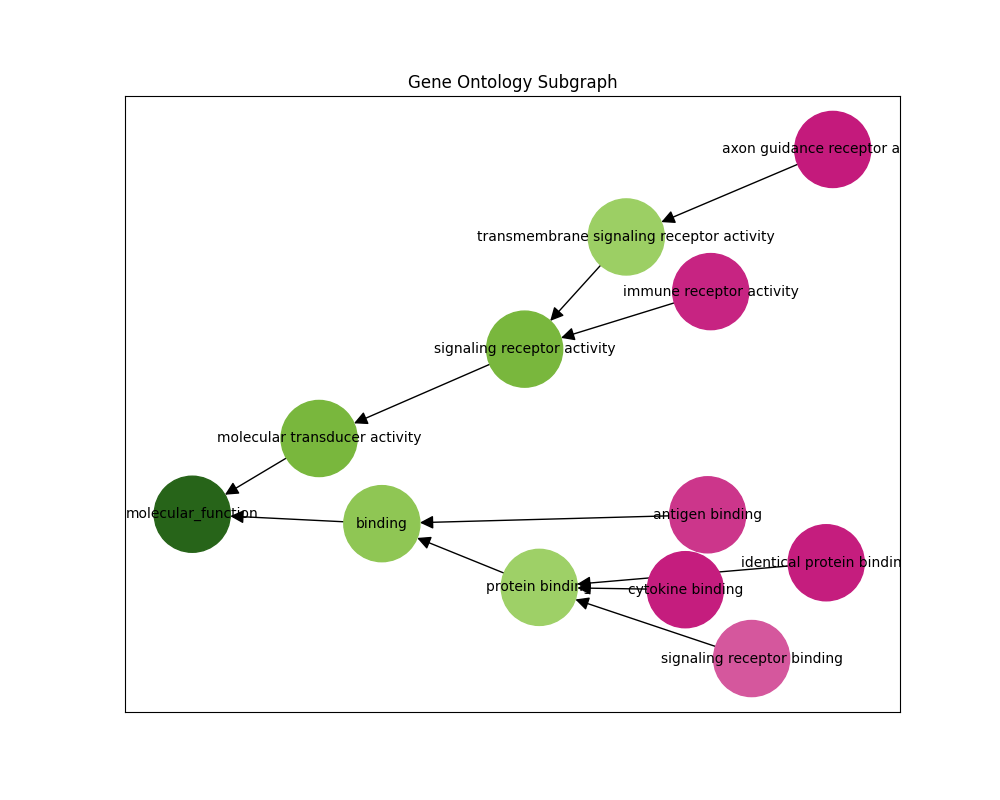
Hierarchy-aware GO subgraph reconstruction — Outputs a full GO directed acyclic graph (DAG) ensuring predictions respect the ontology structure rather than treating each term as an independent binary label.
-
Efficient inference at scale — Supports weight quantization and quantization-aware training (QAT), enabling memory-efficient, high-throughput screening of large sequence datasets without accuracy loss.
What are GO terms?
"The Gene Ontology (GO) is a concept hierarchy that describes the biological function of genes and gene products at different levels of abstraction (Ashburner et al., 2000). It is a good model to describe the multi-faceted nature of protein function."
"GO is a directed acyclic graph. The nodes in this graph are functional descriptors (terms or classes) connected by relational ties between them (is_a, part_of, etc.). For example, terms 'protein binding activity' and 'binding activity' are related by an is_a relationship; however, the edge in the graph is often reversed to point from binding towards protein binding. This graph contains three subgraphs (subontologies): Molecular Function (MF), Biological Process (BP), and Cellular Component (CC), defined by their root nodes. Biologically, each subgraph represent a different aspect of the protein's function: what it does on a molecular level (MF), which biological processes it participates in (BP) and where in the cell it is located (CC)."
From CAFA 5 Protein Function Prediction
V1 Pretrained Models
The following pretrained models are available on HuggingFace Hub and require the esmc-protein-function library version 1.x.x for inference. All V1 models have been optimized with quantization-aware post-training.
| Name | Embedding Dimensions | Encoder Layers | Context Length | Total Parameters |
|---|---|---|---|---|
| andrewdalpino/ESMC-Protein-Function-V1-300M | 960 | 30 | 2048 | 397M |
V0 Pretrained Models
The following pretrained models are available on HuggingFace Hub and require the esmc_function_classifier library version 0.1.x for inference.
| Name | Embedding Dimensions | Encoder Layers | Context Length | Total Parameters |
|---|---|---|---|---|
| andrewdalpino/ESMC-Protein-Function-V0-300M | 960 | 30 | 2048 | 361M |
| andrewdalpino/ESMC-Protein-Function-V0-300M-QAT | 960 | 30 | 2048 | 361M |
| andrewdalpino/ESMC-Protein-Function-V0-600M | 1152 | 36 | 2048 | 644M |
| andrewdalpino/ESMC-Protein-Function-V0-600M-QAT | 1152 | 36 | 2048 | 644M |
Basic Pretrained Example
First, install the esmc-protein-function package using pip.
pip install esmc-protein-function
Then, we'll load the model weights from HuggingFace Hub by calling the from_pretrained() method. We'll also need the ESM tokenizer from the esm library. Then, tokenize the sequence and query the model like in the example below.
import torch
from esm.tokenization import EsmSequenceTokenizer
from esmc_protein_function.model import ESMCProteinFunction
model_name = "andrewdalpino/ESMC-Protein-Function-V1-300M"
sequence = "MPPKGHKKTADGDFRPVNSAGNTIQAKQKYSIDDLLYPKSTIKNLAKETLPDDAIISKDALTAIQRAATLFVSYMASHGNASAEAGGRKKIT"
top_p = 0.5
tokenizer = EsmSequenceTokenizer()
model = ESMCProteinFunction.from_pretrained(model_name)
out = tokenizer(sequence, max_length=2048, truncation=True)
input_ids = torch.tensor(out["input_ids"], dtype=torch.int32)
go_term_probabilities = model.predict_terms(
input_ids, top_p=top_p
)
Predict GO Subgraph
You can also output the gene-ontology (GO) networkx subgraph for a given sequence like in the example below. You'll need an up-to-date gene ontology database that you can import using the obonet package.
pip install obonet
Then, load the GO DAG and call the predict_all_subgraphs() method like in the example below.
import networkx as nx
import obonet
# Visit https://geneontology.org/docs/download-ontology/ to download.
go_db_path = "./dataset/go-basic.obo"
graph = obonet.read_obo(go_db_path)
model.load_gene_ontology(graph)
subgraph, go_term_probabilities = model.predict_all_subgraphs(
input_ids, top_p=top_p
)
json = nx.node_link_data(subgraph)
print(json)
Quantized Model
To quantize the model weights using int8 call the quantize_weights() method. Any model can be quantized, but we recommend one that has been quantization-aware trained (QAT) for the best performance. The group_size argument controls the granularity at which quantization scales are computed.
model.quantize_weights(group_size=64)
References
- T. Hayes, et al. Simulating 500 million years of evolution with a language model, 2024.
- M. Ashburner, et al. Gene Ontology: tool for the unification of biology, 2000.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file esmc_protein_function-1.0.0.tar.gz.
File metadata
- Download URL: esmc_protein_function-1.0.0.tar.gz
- Upload date:
- Size: 17.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
05e460a7ffe5c3a8ca325bf8cb2d3440f3d099bcc2049bcc086d295028aeba92
|
|
| MD5 |
35af9052891ebf57ba12d0259b06900c
|
|
| BLAKE2b-256 |
52dc2178322698a638d0b638b65254d68df356c162fc8fdf98ee152583cd23d6
|
File details
Details for the file esmc_protein_function-1.0.0-py3-none-any.whl.
File metadata
- Download URL: esmc_protein_function-1.0.0-py3-none-any.whl
- Upload date:
- Size: 12.4 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.12.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6015fe208471f21988e389da99b4dc03686534920051324a8a4023050e705db9
|
|
| MD5 |
4604ed92877fefe511b2bc77bbcc976b
|
|
| BLAKE2b-256 |
a65ff631680b5e6ce7f9e22a3bf2fe62301501518b8960210d6821e63002c7d2
|