Detect and extract individual panels from scientific figures using YOLOv12
Project description
figpanel
Detect and extract individual panels from scientific figures using YOLOv12.
Scientific papers often combine multiple plots into a single composite figure (Figure 1a, 1b, 1c...). figpanel automatically detects these sub-panels, reads their labels via OCR, and extracts each one as a separate image.
Quick Start
pip install figpanel
import figpanel
# Detect subplots and captions (bounding boxes + confidence scores)
results = figpanel.detect("figure.png")
# {'subplots': [(x1,y1,x2,y2,conf), ...], 'captions': [...]}
# Visualize detections on the image
figpanel.visualize("figure.png", save="annotated.png")
For the full pipeline (detect + OCR + match + crop + deduplicate):
pip install figpanel[full]
# Extract individual panels
panels = figpanel.extract("figure.png", "output/")
for panel in panels:
print(f"Panel {panel.label}: bbox={panel.bbox}, confidence={panel.confidence:.2f}")
panel.image.show()
Examples
 |
 |
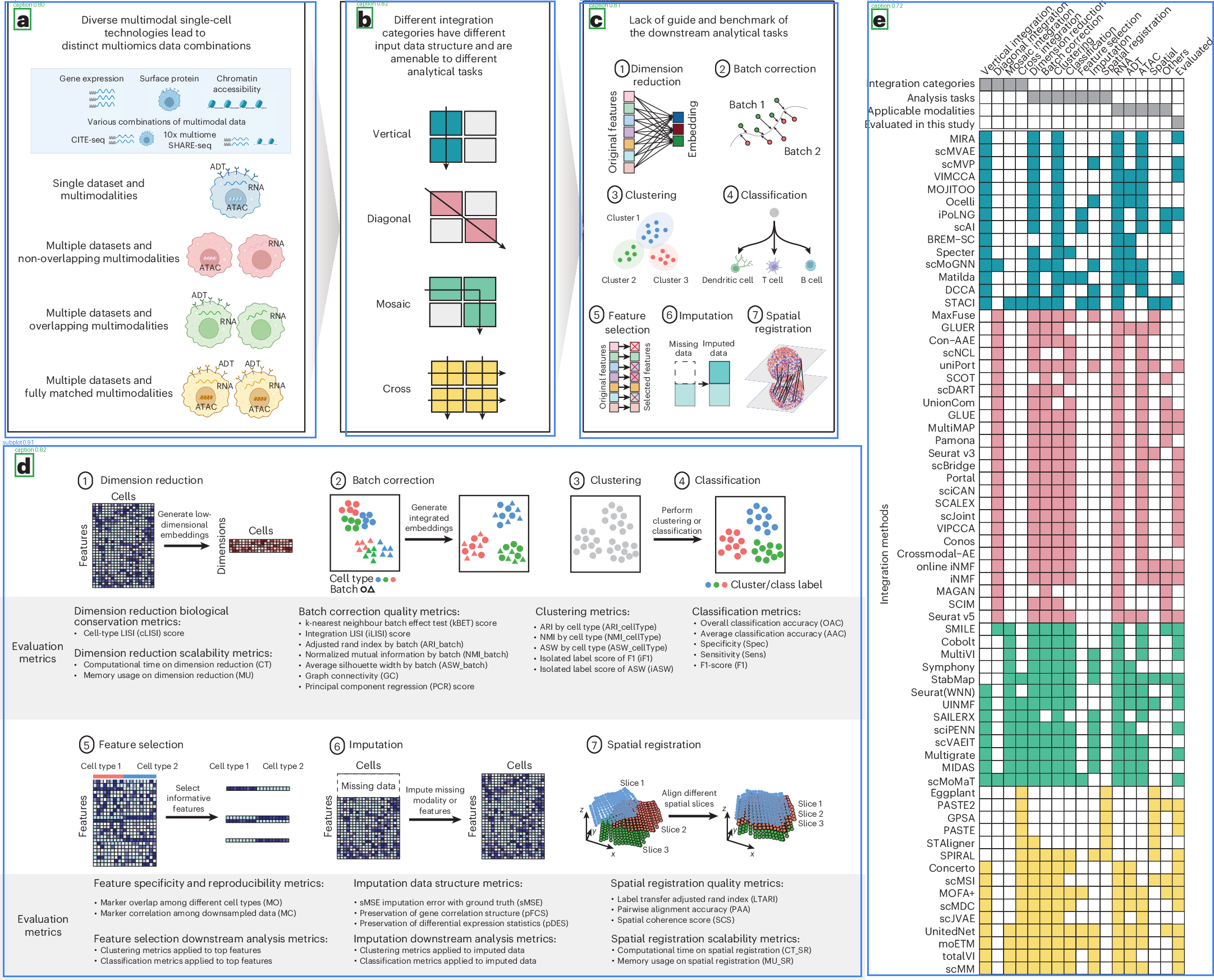 |
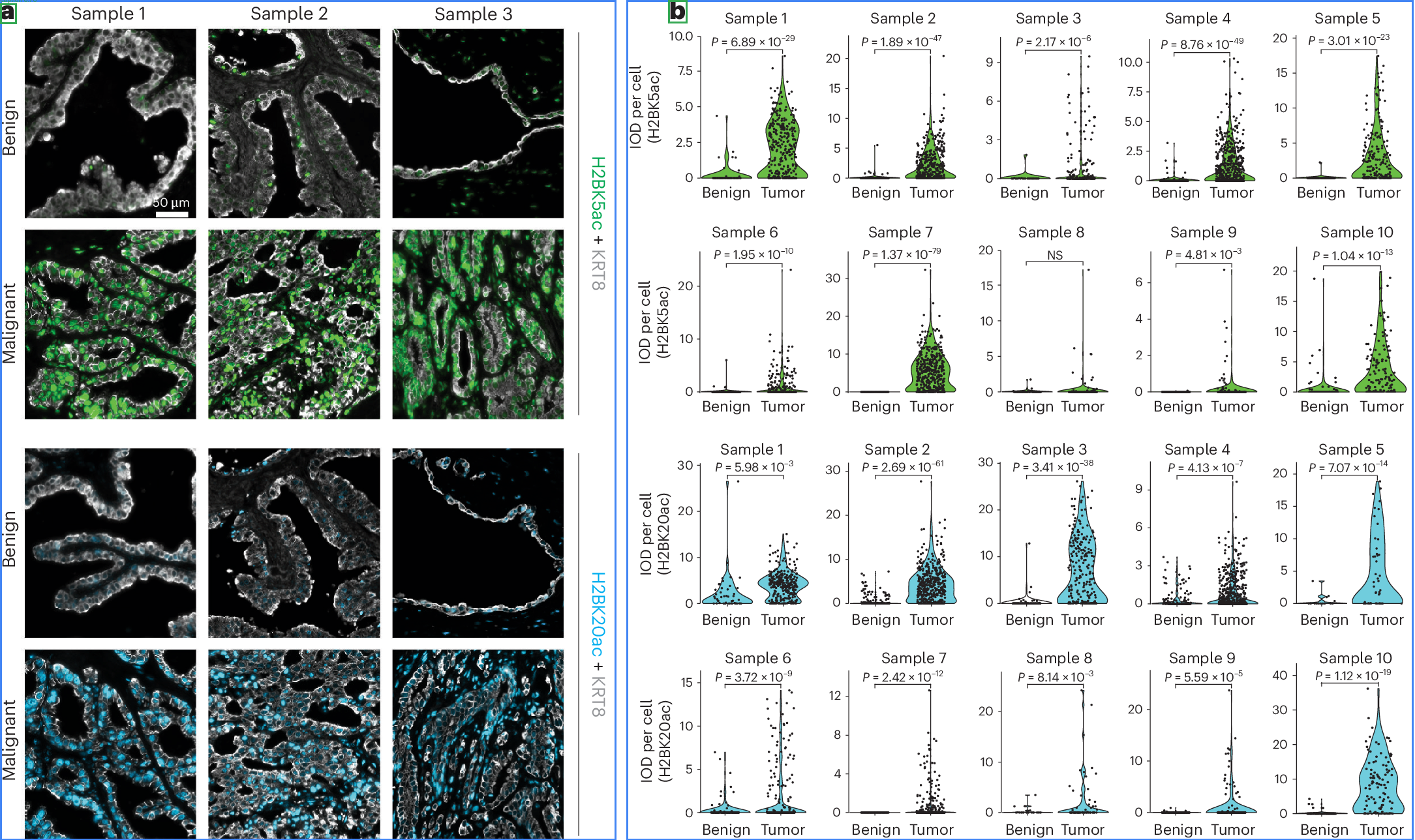
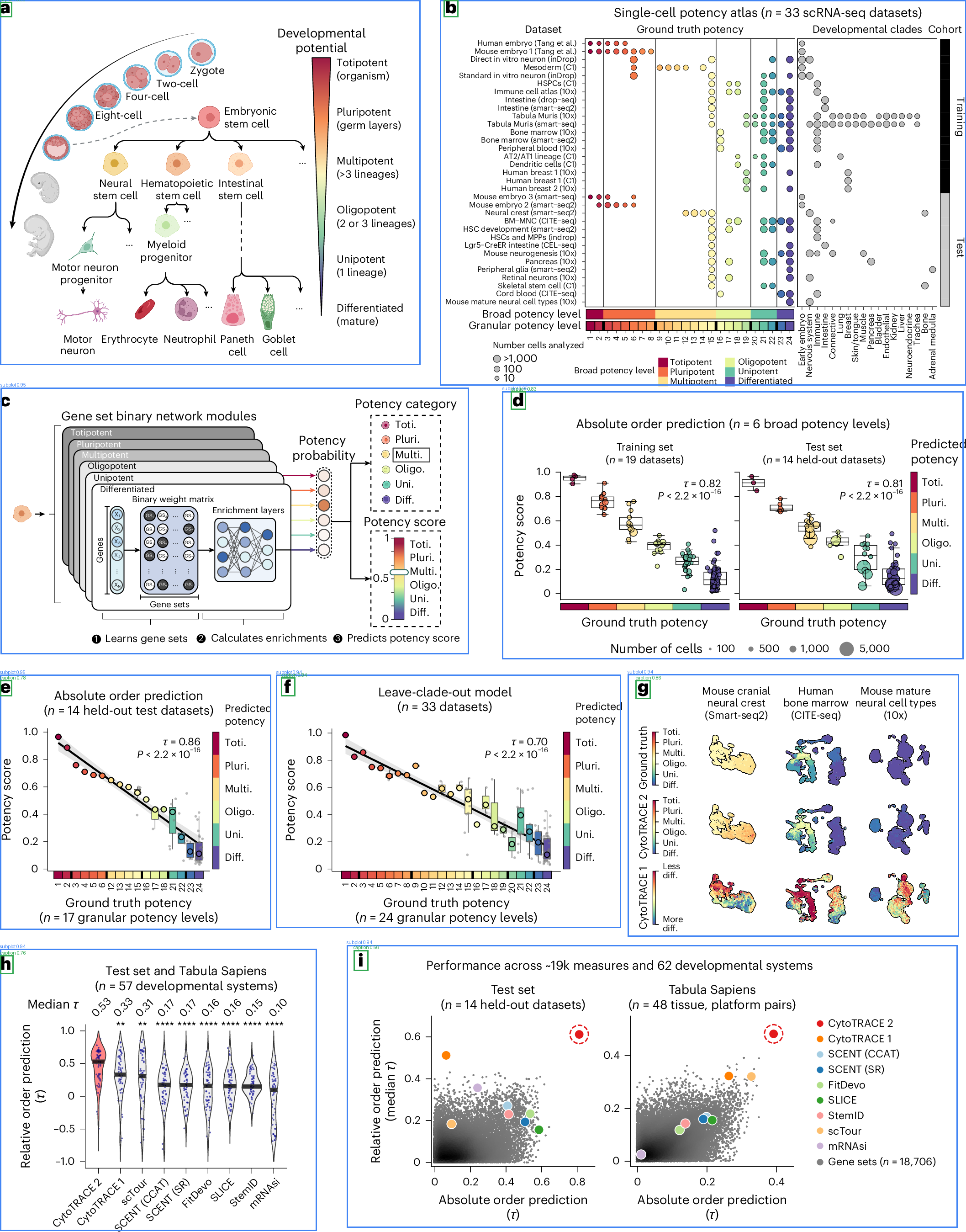
Blue boxes = detected subplots, Green boxes = detected captions. Images from open-access papers (CC BY 4.0).
Features
- YOLOv12 Detection - Trained on 5,000+ annotated scientific figures to detect subplots and caption labels
- Automatic Model Download - Model weights are hosted on HuggingFace and downloaded automatically on first use
- OCR Caption Reading - Reads single-character panel labels (a, b, c...) using Tesseract
- Smart Matching - Greedy nearest-neighbor algorithm matches captions to their corresponding subplots
- Deduplication - Removes duplicate panels using perceptual hashing (pHash) and ORB feature matching
- Two-Tier Install - Base install for detection only;
[full]adds OCR, matching, and deduplication
Model Details
| Property | Value |
|---|---|
| Architecture | YOLOv12 |
| Classes | subplot (panel region), caption (label character) |
| Training Data | 5,000+ annotated figures from open-access journals |
| Model Size | ~18 MB |
| Input | Any image (PNG, JPEG, etc.) |
| Hosted On | HuggingFace |
Installation Options
# Detection only (ultralytics + huggingface_hub)
pip install figpanel
# Full pipeline (+ pytesseract, imagehash, opencv)
pip install figpanel[full]
Note: The
[full]install requires Tesseract OCR to be installed on your system.
API Reference
figpanel.detect(image_path, conf=0.25, iou=0.5)
Run YOLO detection on a scientific figure.
Returns: dict with "subplots" and "captions" keys, each containing a list of (x1, y1, x2, y2, confidence) tuples.
figpanel.extract(image_path, output_dir=None, *, conf=0.25, iou=0.5, padding=4, dedup_thresh=0.3, prefix=None)
Full pipeline: detect, OCR, match, crop, and deduplicate panels.
Returns: list[Panel] where each Panel has .label, .bbox, .confidence, and .image attributes.
Requires: pip install figpanel[full]
figpanel.visualize(image_path, save=None, *, conf=0.25, iou=0.5, line_width=3)
Draw detection boxes on the image. Subplots in blue, captions in green.
Returns: Annotated PIL.Image.
Use Cases
- Literature Mining - Automatically decompose composite figures for indexing and search
- Dataset Creation - Build plot-type datasets from scientific papers at scale
- Accessibility - Extract individual panels for screen readers and alternative text
- Research Tools - Integrate panel detection into paper analysis pipelines
- Quality Control - Validate figure composition in manuscript preparation
About Plottie
figpanel is built and maintained by Plottie - the scientific plot discovery platform. Plottie helps researchers explore, collect, and find inspiration from high-quality scientific plots across open-access literature.
Visit plottie.art to browse thousands of curated scientific figures.
Citation
If you use figpanel in your research, please cite:
@software{figpanel,
title = {figpanel: Scientific Figure Panel Detector},
author = {Plottie},
url = {https://github.com/Plottie/figpanel},
version = {0.1.0},
year = {2026},
note = {Built by Plottie (https://plottie.art)}
}
License
This project is licensed under the AGPL-3.0 license. The YOLOv12 model weights inherit this license from Ultralytics.
Academic and research use is unaffected. If you integrate figpanel into a network service, you must make your source code available under AGPL-3.0.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file figpanel-0.1.0.tar.gz.
File metadata
- Download URL: figpanel-0.1.0.tar.gz
- Upload date:
- Size: 7.0 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
ad3d96231ee51c16308be1ff5be9e9b4ab7c6526a7c0d58a83d7f6ed1d219810
|
|
| MD5 |
2d13416aa28deee161d65e7685a74dd7
|
|
| BLAKE2b-256 |
2082b8e6b42e582588a05302ce074bb96b1b5de5b9e17bfbffdd353be09bacda
|
File details
Details for the file figpanel-0.1.0-py3-none-any.whl.
File metadata
- Download URL: figpanel-0.1.0-py3-none-any.whl
- Upload date:
- Size: 23.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.12.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
19750ff6f6bd274a5a7c102a744b990eff402f00b4cd8c31af18890fc12ae173
|
|
| MD5 |
e99947ea6fc4108521d3936e61075f32
|
|
| BLAKE2b-256 |
63b0b72eb2fb8cd41e8386967ee29667d42bcbcd7d174e61411eeea5c440022a
|
















