FineST: Fine-grained Spatial Transcriptomic
Project description
This software package impletements FineST (Fine-grained Spatial Transcriptomic), which could identify super-resolved ligand-receptor interactions with spatial co-expression (i.e., spatial association) from a spot-level to a sub-spot level or single-cell level.
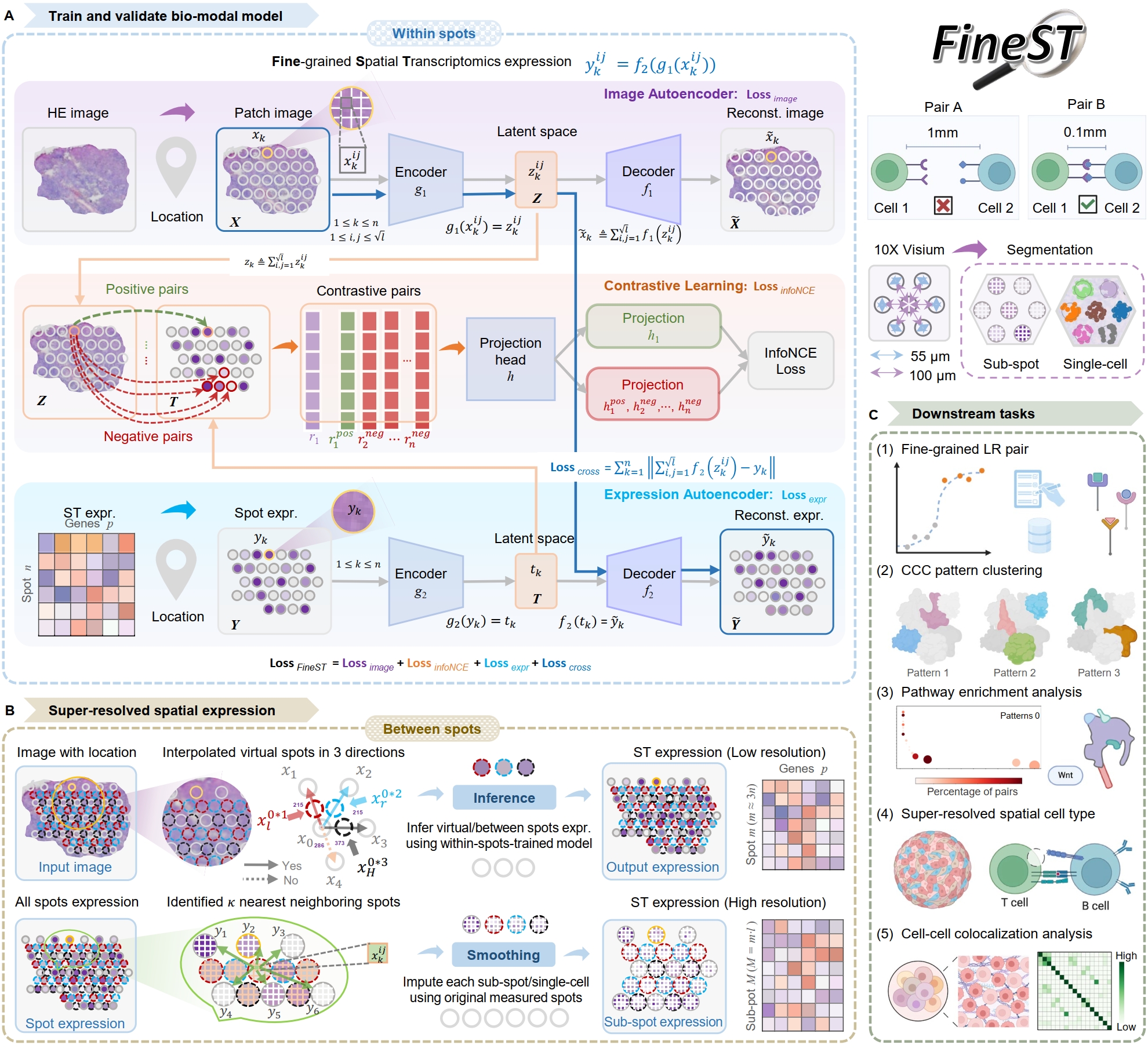
It comprises three components (Training-Imputation-Discovery) after HE image feature is extracted:
Step0: HE image feature extraction
Step1: Training FineST on the within spots
Step2: Super-resolution spatial RNA-seq imputation
Step3: Fine-grained LR pair and CCC pattern discovery
Installation
FineST is available through PyPI. To install, type the following command line and add -U for updates:
pip install -U FineSTAlternatively, install from this GitHub repository for latest (often development) version (time: < 1 min):
pip install -U git+https://github.com/StatBiomed/FineSTInstallation using Conda
$ git clone https://github.com/StatBiomed/FineST.git
$ conda create --name FineST python=3.8
$ conda activate FineST
$ cd FineST
$ pip install -r requirements.txtTypically installation is completed within a few minutes. Then install pytorch, refer to pytorch installation.
$ conda install pytorch=1.7.1 torchvision torchaudio cudatoolkit=11.0 -c pytorchVerify the installation using the following command:
python
>>> import torch
>>> print(torch.__version__)
>>> print(torch.cuda.is_available())Tutorial:
For a tutorial, please see: https://github.com/StatBiomed/FineST/tree/main/tutorial/NPC_Train_Impute_demo.ipynb
Get Started for Visium or Visium HD data
Usage illustrations:
The source codes for reproducing the FineST analysis in this work are provided (see demo directory). All relevant materials involved in the reproducing codes are available from Google Drive.
For Visium, using a single slice of 10x Visium human nasopharyngeal carcinoma (NPC) data.
For Visium HD, using a single slice of 10x Visium HD human colorectal cancer (CRC) data with 16-um bin.
Step0: HE image feature extraction (for Visium)
Visium measures about 5k spots across the entire tissue area. The diameter of each individual spot is roughly 55 micrometers (um), while the center-to-center distance between two adjacent spots is about 100 um. In order to capture the gene expression profile across the whole tissue ASAP,
Firstly, interpolate between spots in horizontal and vertical directions, using Spot_interpolate.py.
python ./FineST/demo/Spot_interpolate.py \
--data_path ./Dataset/NPC/ \
--position_list tissue_positions_list.csv \
--dataset patient1with Input: tissue_positions_list.csv - Locations of within spots (n), and Output: _position_add_tissue.csv- Locations of between spots (m ~= 3n).
Then extracte the within spots HE image feature embeddings using Image_feature_extraction.py.
python ./FineST/demo/Image_feature_extraction.py \
--dataset AH_Patient1 \
--position ./Dataset/NPC/patient1/tissue_positions_list.csv \
--image ./Dataset/NPC/patient1/20210809-C-AH4199551.tif \
--scale_image False \
--method Virchow2 \
--output_path_img ./Dataset/NPC/HIPT/AH_Patient1_pth_112_14_image \
--output_path_pth ./Dataset/NPC/HIPT/AH_Patient1_pth_112_14 \
--patch_size 112 \
--logging_folder ./Logging/HIPT_AH_Patient1/Similarlly, extracte the between spots HE image feature embeddings using Image_feature_extraction.py.
python ./FineST/demo/Image_feature_extraction.py \
--dataset AH_Patient1 \
--position ./Dataset/NPC/patient1/patient1_position_add_tissue.csv \
--image ./Dataset/NPC/patient1/20210809-C-AH4199551.tif \
--scale_image False \
--method Virchow2 \
--output_path_img ./Dataset/NPC/HIPT/NEW_AH_Patient1_pth_112_14_image \
--output_path_pth ./Dataset/NPC/HIPT/NEW_AH_Patient1_pth_112_14 \
--patch_size 112 \
--logging_folder ./Logging/HIPT_AH_Patient1/The image segment execution time: 8.153s, the image feature extract time: 35.499s.
Input files:
20210809-C-AH4199551.tif: Raw histology image
patient1_position_add_tissue.csv: “Between spot” (Interpolated spots) locations
Output files:
NEW_AH_Patient1_pth_112_14_image: Segmeted “Between spot” histology image patches (.png)
NEW_AH_Patient1_pth_112_14: Extracted “Between spot” image feature embeddiings for each patche (.pth)
Step0: HE image feature extraction (for Visium HD)
Visium HD captures continuous squares without gaps, it measures the whole tissue area.
python ./FineST/demo/Image_feature_extraction.py \
--dataset HD_CRC_16um \
--position ./Dataset/CRC/square_016um/tissue_positions.parquet \
--image ./Dataset/CRC/square_016um/Visium_HD_Human_Colon_Cancer_tissue_image.btf \
--scale_image True \
--method Virchow2 \
--output_path_img ./Dataset/CRC/HIPT/HD_CRC_16um_pth_28_14_image \
--output_path_pth ./Dataset/CRC/HIPT/HD_CRC_16um_pth_28_14 \
--patch_size 28 \
--logging_folder ./Logging/HIPT_HD_CRC_16um/The image segment execution time: 62.491s, the image feature extract time: 1717.818s.
Input files:
Visium_HD_Human_Colon_Cancer_tissue_image.btf: Raw histology image (.btf Visium HD or .tif Visium)
tissue_positions.parquet: Spot/bin locations (.parquet Visium HD or .csv Visium)
Output files:
HD_CRC_16um_pth_28_14_image: Segmeted histology image patches (.png)
HD_CRC_16um_pth_28_14: Extracted image feature embeddiings for each patche (.pth)
Step1: Training FineST on the within spots
On Visium dataset, if trained weights (i.e. weight_save_path) have been obtained, just run the following command. Otherwise, if you want to re-train a model, just omit weight_save_path line.
python ./FineST/FineST/demo/FineST_train_infer.py \
--system_path '/mnt/lingyu/nfs_share2/Python/' \
--weight_path 'FineST/FineST_local/Finetune/' \
--parame_path 'FineST/FineST/parameter/parameters_NPC_P10125.json' \
--dataset_class 'Visium' \
--gene_selected 'CD70' \
--LRgene_path 'FineST/FineST/Dataset/LRgene/LRgene_CellChatDB_baseline.csv' \
--visium_path 'FineST/FineST/Dataset/NPC/patient1/tissue_positions_list.csv' \
--image_embed_path 'NPC/Data/stdata/ZhuoLiang/LLYtest/AH_Patient1_pth_112_14/' \
--spatial_pos_path 'FineST/FineST_local/Dataset/NPC/ContrastP1geneLR/position_order.csv' \
--reduced_mtx_path 'FineST/FineST_local/Dataset/NPC/ContrastP1geneLR/harmony_matrix.npy' \
--weight_save_path 'FineST/FineST_local/Finetune/20240125140443830148' \
--figure_save_path 'FineST/FineST_local/Dataset/NPC/Figures/'FineST_train_infer.py is used to train and evaluate the FineST model using Pearson Correlation, it outputs:
Average correlation of all spots: 0.8534651812923978
Average correlation of all genes: 0.8845136777311445
Input files:
parameters_NPC_P10125.json: The model parameters.
LRgene_CellChatDB_baseline.csv: The genes involved in Ligand or Receptor from CellChatDB.
tissue_positions_list.csv: It can be found in the spatial folder of 10x Visium outputs.
AH_Patient1_pth_112_14: Image feature folder from HIPT Image_feature_extraction.py.
position_order.csv: Ordered tissue positions list, according to image patches’ coordinates.
harmony_matrix.npy: Ordered gene expression matrix, according to image patches’ coordinates.
20240125140443830148: The trained weights. Just omit it if you want to newly train a model.
Output files:
Finetune: The logging results model.log and trained weights epoch_50.pt (.log and .pt)
Figures: The visualization plots, used to see whether the model trained well or not (.pdf)
Step2: Super-resolution spatial RNA-seq imputation
For sub-spot resolution
This step supposes that the trained weights (i.e. weight_save_path) have been obtained, just run the following.
python ./FineST/FineST/demo/High_resolution_imputation.py \
--system_path '/mnt/lingyu/nfs_share2/Python/' \
--weight_path 'FineST/FineST_local/Finetune/' \
--parame_path 'FineST/FineST/parameter/parameters_NPC_P10125.json' \
--dataset_class 'Visium' \
--gene_selected 'CD70' \
--LRgene_path 'FineST/FineST/Dataset/LRgene/LRgene_CellChatDB_baseline.csv' \
--visium_path 'FineST/FineST/Dataset/NPC/patient1/tissue_positions_list.csv' \
--imag_within_path 'NPC/Data/stdata/ZhuoLiang/LLYtest/AH_Patient1_pth_112_14/' \
--imag_betwen_path 'NPC/Data/stdata/ZhuoLiang/LLYtest/NEW_AH_Patient1_pth_112_14/' \
--spatial_pos_path 'FineST/FineST_local/Dataset/NPC/ContrastP1geneLR/position_order_all.csv' \
--weight_save_path 'FineST/FineST_local/Finetune/20240125140443830148' \
--figure_save_path 'FineST/FineST_local/Dataset/NPC/Figures/' \
--adata_all_supr_path 'FineST/FineST_local/Dataset/ImputData/patient1/patient1_adata_all.h5ad' \
--adata_all_spot_path 'FineST/FineST_local/Dataset/ImputData/patient1/patient1_adata_all_spot.h5ad'High_resolution_imputation.py is used to predict super-resolved gene expression based on the image segmentation (Geometric sub-spot level or Nuclei single-cell level).
Input files:
parameters_NPC_P10125.json: The model parameters.
LRgene_CellChatDB_baseline.csv: The genes involved in Ligand or Receptor from CellChatDB.
tissue_positions_list.csv: It can be found in the spatial folder of 10x Visium outputs.
AH_Patient1_pth_112_14: Image feature of within-spots from Image_feature_extraction.py.
NEW_AH_Patient1_pth_112_14: Image feature of between-spots from Image_feature_extraction.py.
position_order_all.csv: Ordered tissue positions list, of both within spots and between spots.
20240125140443830148: The trained weights. Just omit it if you want to newly train a model.
Output files:
Finetune: The logging results model.log and trained weights epoch_50.pt (.log and .pt)
Figures: The visualization plots, used to see whether the model trained well or not (.pdf)
patient1_adata_all.h5ad: High-resolution gene expression, at sub-spot level (16x3x resolution).
patient1_adata_all_spot.h5ad: High-resolution gene expression, at spot level (3x resolution).
For single-cell resolution
Using sc Patient1 pth 16 16 i.e., the image feature of single-nuclei from Image_feature_extraction.py, just run the following.
python ./FineST/FineST/demo/High_resolution_imputation.py \
--system_path '/mnt/lingyu/nfs_share2/Python/' \
--weight_path 'FineST/FineST_local/Finetune/' \
--parame_path 'FineST/FineST/parameter/parameters_NPC_P10125.json' \
--dataset_class 'VisiumSC' \
--gene_selected 'CD70' \
--LRgene_path 'FineST/FineST/Dataset/LRgene/LRgene_CellChatDB_baseline.csv' \
--visium_path 'FineST/FineST/Dataset/NPC/patient1/tissue_positions_list.csv' \
--imag_within_path 'NPC/Data/stdata/ZhuoLiang/LLYtest/AH_Patient1_pth_112_14/' \
--image_embed_path_sc 'NPC/Data/stdata/ZhuoLiang/LLYtest/sc_Patient1_pth_16_16/' \
--spatial_pos_path_sc 'FineST/FineST_local/Dataset/NPC/ContrastP1geneLR/position_order_sc.csv' \
--adata_super_path_sc 'FineST/FineST_local/Dataset/ImputData/patient1/patient1_adata_all_sc.h5ad' \
--weight_save_path 'FineST/FineST_local/Finetune/20240125140443830148' \
--figure_save_path 'FineST/FineST_local/Dataset/NPC/Figures/'Step3: Fine-grained LR pair and CCC pattern discovery
This step is based on SpatialDM and SparseAEH (developed by our Lab).
SpatialDM: for significant fine-grained ligand-receptor pair selection.
SparseAEH: for fastly cell-cell communication pattern discovery, 1000 times speedup to SpatialDE.
Detailed Manual
The full manual is at FineST tutorial for installation, tutorials and examples.
Spot interpolation for Visium datasets.
Step1 and Step2 Train FineST and impute super-resolved spatial RNA-seq.
Step3 Fine-grained LR pair and CCC pattern discovery.
Downstream analysis Cell type deconvolution, ROI region cropping, cell-cell colocalization.
Performance evaluation of FineST vs (TESLA and iSTAR).
Inference comparison of FineST vs iStar (only LR genes).
Contact Information
Please contact Lingyu Li (lingyuli@hku.hk) or Yuanhua Huang (yuanhua@hku.hk) if any enquiry.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file FineST-0.1.2.tar.gz.
File metadata
- Download URL: FineST-0.1.2.tar.gz
- Upload date:
- Size: 73.0 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.8.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1946dc8437abea8507f720f230662b91cfeaf64f2d9c159e42b13b6e7e4ba151
|
|
| MD5 |
48985039ffe4b8f745863a7f916386ec
|
|
| BLAKE2b-256 |
f69d98a4dd58fba5fad87d097fb1ff13352b8d7ca6776422b112efa5b16cfc3e
|
File details
Details for the file FineST-0.1.2-py3-none-any.whl.
File metadata
- Download URL: FineST-0.1.2-py3-none-any.whl
- Upload date:
- Size: 138.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.8.2
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7c08667b9240e08b728d4a6cd9fcf89608b2c3fff4f8d6ba7681e0e964a77f44
|
|
| MD5 |
9f0a0564998ad428e7f214be0553872e
|
|
| BLAKE2b-256 |
87a47fc865cee39c6558f8d65eb12025d05ff74b3bb694e9b71e23368af30912
|











