A Generic Particle+Grid Interface
Project description
GPGI
A Generic Particle + Grid data Interface
This small Python library implements fundamental grid deposition algorithms to analyse (rectilinear) grid + particle datasets, with an emphasize on performance. Core algorithms are implemented as Cython extensions.
Installation
python -m pip install --upgrade pip
python -m pip install gpgi
Supported applications
A rectilinear grid is defined as 1D arrays representing cell left edges in each directions. Note that the last point of such an array is interpreted as the right edge of the rightmost cell, so for instance, a 1D grid containing 100 cells is defined by 101 edges.
Particles are defined as points that live on the grid.
Deposition is the action of going from particule description to a grid description of a field. It is useful to analyze, compare and combine simulation data that exists in a combination of the two formalisms. This process is not reversible as it degrades information.
For instance, here's a simple overlay of a particle set (red dots) against a background that represents the deposited particle count.
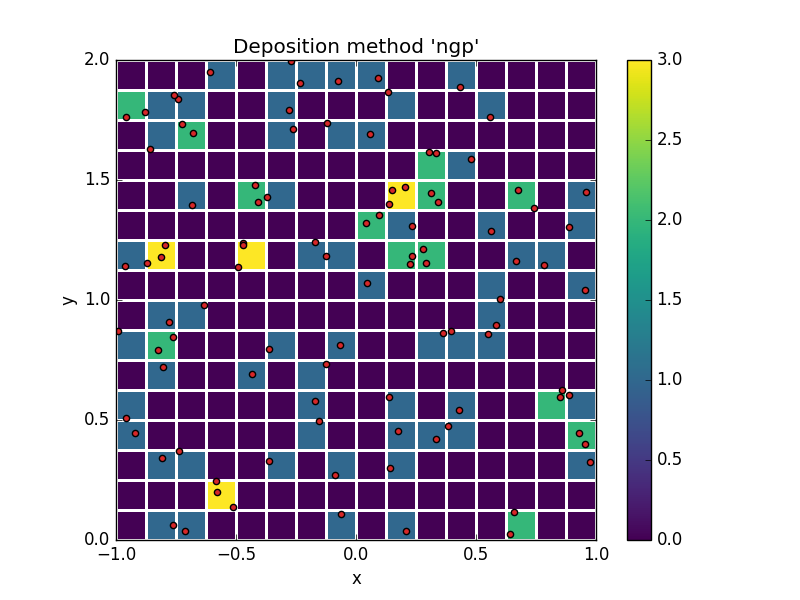
This example illustrates the simplest possible deposition method "Particle in Cell", in which each particle contributes only to the cell that contains it.
More refined methods are also available.
Supported deposition methods
| method name | abreviated name | order |
|---|---|---|
| Nearest Grid Point | NGP | 0 |
| Cloud in Cell | CIC | 1 |
| Triangular Shaped Cloud | TSC | 2 |
Supported geometries
| geometry name | axes order |
|---|---|
| cartesian | x, y, z |
| polar | radius, z, azimuth |
| cylindrical | radius, azimuth, z |
| spherical | radius, colatitude, azimuth |
| equatorial | radius, azimuth, latitude |
Time complexity
An important step in perfoming deposition is to associate particle indices to cell indices. This step is called "particle indexing". In directions where the grid is uniformly stepped (if any), indexing a particle is an O(1) operation. In the more general case, indexing is performed by bisection, which is a O(log(nx))) operation (where nx represents the number of cells in the direction of interest).
Usage
The API consists in a load function, which returns a Dataset object.
Load data
import numpy as np
import gpgi
nx = ny = 64
nparticles = 600_000
prng = np.random.RandomState(0)
ds = gpgi.load(
geometry="cartesian",
grid={
"cell_edges": {
"x": np.linspace(-1, 1, nx),
"y": np.linspace(-1, 1, ny),
},
},
particles={
"coordinates": {
"x": 2 * (prng.normal(0.5, 0.25, nparticles) % 1 - 0.5),
"y": 2 * (prng.normal(0.5, 0.25, nparticles) % 1 - 0.5),
},
"fields": {
"mass": np.ones(nparticles),
},
},
)
The Dataset object holds a grid and a particles attribute,
which both hold a fields attribute for accessing their data.
But more importantly, the Dataset has a deposit method to
translate particle fields to the grid formalism.
Deposit Particle fields on the grid
particle_mass = ds.deposit("mass", method="nearest_grid_point") # or "ngp" for shorts
Visualize
In this example we'll use matplotlib for rendering, but note that matplotlib is not a dependency to gpgi
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
ax.set(aspect=1, xlabel="x", ylabel="y")
im = ax.pcolormesh(
"x",
"y",
particle_mass.T,
data=ds.grid.cell_edges,
cmap="viridis",
)
fig.colorbar(im, ax=ax)
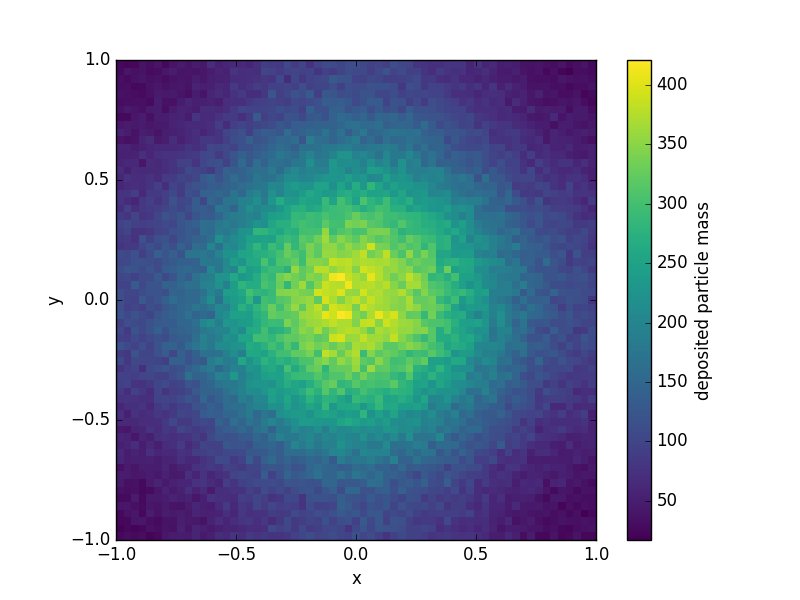
The example script given here takes about a second (top to bottom).
Supplying arbitrary metadata
new in gpgi 0.4.0
Dataset objects have a special attribute metadata which is a dictionary with string keys.
This attribute is meant to hold any special metadata that may be relevant for labelling or processing (e.g. simulation time, author, ...).
Metadata can be supplied at load time as
ds = gpgi.load(
geometry="cartesian",
grid=...,
particles=...,
metadata={"simulation_time": 12.5, "author": "Clément Robert"}
)
Boundary conditions
new in gpgi 0.5.0
With CIC and TSC deposition, particles contribute to cells neighbouring the one
that contains them. For particles that live in the outermost layer of the
domain, this means some of their contribution is lost. This behaviour
corresponds to the default 'open' boundary condition, but gpgi has builtin
support for more conservative boundary conditions.
Boundary conditions can selected per field, per axis and per side. Builtin recipes all perform linear combinations of ghost layers (same-side and opposite side) and active domain layers (same-side and opposite side), and replace the same-side active layer with the result.
User-selected boundary conditions take the form of an optional argument to
Dataset.deposit, as dictionnary with keys being axes names, and values being
2-tuples of boundary conditions names (for left and right side respectively).
For instance, here's how one would require periodic boundary conditions on all axes:
ds.deposit(
"mass",
method="cic",
boundaries={
"x": ("periodic", "periodic"),
"y": ("periodic", "periodic"),
}
)
Unspecified axes will use the default 'open' boundary.
Builtin recipes
| boundary conditions | description | conservative ? |
|---|---|---|
| open (default) | no special treatment | no |
| periodic | add opposite ghost layer to the active domain | yes |
| wall | add same-side ghost layer to the active domain | yes |
| antisymmetric | substract same-side ghost layer from the active domain | no |
Define custom recipes
gpgi's boundary recipes can be customized. Let's illustrate this feature with a simple example.
Say we want to fix the value of the deposited field in some outer layer.
This is done by defining a new function on the user side:
def ones(
same_side_active_layer,
same_side_ghost_layer,
opposite_side_active_layer,
opposite_side_ghost_layer,
weight_same_side_active_layer,
weight_same_side_ghost_layer,
weight_opposite_side_active_layer,
weight_opposite_side_ghost_layer,
side,
metadata,
):
return 1.0
where all first eight arguments are numpy.ndarray objects with the same shape (which includes ghost padding !),
to which the return value must be broadcastable, side can only be either
"left" or "right", and metadata is the special Dataset.metadata
attribute. Not all arguments need be used in the body of the function, but this
signature is required.
The method must then be registered as a boundary condition recipe as
ds.boundary_recipes.register("ones", ones)
where the associated key (here "ones") is arbitrary. The recipe can now be
used exactly as builtin ones, and all of them can be mixed arbitrarily.
ds.deposit(
"mass",
method="cic",
boundaries={
"x": ("ones", "wall"),
"y": ("periodic", "periodic"),
}
)
Note that all first eight arguments in a boundary recipe function should
represent an extensive physical quantity (as opposed to intensive). When
depositing an intensive quantity u, a weight field w should be supplied
(see next section), in which case, the first four arguments represent u*w and
the following four represent w, so that u can still be obtained within the
function as a ratio if needed.
Weight fields
new in gpgi 0.7.0
Intrisically, deposition algorithms assume that the deposited field u
represents an extensive physical quantity (like mass or momentum).
In order to deposit a intensive quantity (like velocity or temperature), one must provide an appropriate weight field w.
Let u' and w' be the equivalent on-grid descriptions to u and w. They are obtained as
w'(x) = Σ w(i) c(i,x)
u'(x) = (1/w'(x)) Σ u(i) w(i) c(i,x)
where x is the spatial position, i is a particle index, and c(i,x) are
geometric coefficients associated with the deposition method.
Boundary recipes may be associated to the weight field with the
weight_field_boundaries argument.
Call help(ds.deposit) for more detail.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distributions
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file gpgi-0.7.2.tar.gz.
File metadata
- Download URL: gpgi-0.7.2.tar.gz
- Upload date:
- Size: 359.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
63a4c42aae45c2a0fd7eeb1f0a627568fa0fcf5f4788ddc7cf142efc8d299d5f
|
|
| MD5 |
8f2d2e8627c320749538accef36681a1
|
|
| BLAKE2b-256 |
6446889736c09ba0546450f4214e8226836e74573ce8dd10efcec48fbb7cdd88
|
File details
Details for the file gpgi-0.7.2-cp311-cp311-win_amd64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp311-cp311-win_amd64.whl
- Upload date:
- Size: 201.2 kB
- Tags: CPython 3.11, Windows x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dd634472a342878115a4a90c3576142e2aa130a34254f57d6cf12f92a128d994
|
|
| MD5 |
baadc860ebf11c7f8cac80a5a0defc3e
|
|
| BLAKE2b-256 |
5ec6f96a273f5614f828f47bccb877d0264b839fbc676e599d2b4203e350f31d
|
File details
Details for the file gpgi-0.7.2-cp311-cp311-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp311-cp311-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 1.3 MB
- Tags: CPython 3.11, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9fa68b0bd7b76247102362630d28af5d513a5c24ca9b94e9f61610b2f2b4c5bf
|
|
| MD5 |
53f0b33d562be6506cb3f32447799c0e
|
|
| BLAKE2b-256 |
4884820e91ffb6f62a636b36f999df7f31c045b60d8a9ba67189bd133de00518
|
File details
Details for the file gpgi-0.7.2-cp311-cp311-macosx_10_9_x86_64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp311-cp311-macosx_10_9_x86_64.whl
- Upload date:
- Size: 239.0 kB
- Tags: CPython 3.11, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1a3b810cab1ab3e765ab8bd438214f8430f12a3307ebf269a03be3ce90654fef
|
|
| MD5 |
896b527c3bed49c42a4809a3bc0588b1
|
|
| BLAKE2b-256 |
3c3f85b3eba50d7be6812736553616369da4e650590d5c5cb4e26c28d2e48e6f
|
File details
Details for the file gpgi-0.7.2-cp310-cp310-win_amd64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp310-cp310-win_amd64.whl
- Upload date:
- Size: 203.1 kB
- Tags: CPython 3.10, Windows x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
cd1ccf0ce6e9983f2729eaf02f26709d2ceb4fe99a185d918d7e052bd4aad345
|
|
| MD5 |
7951044c312e3c30b7ec03bdb4502dd3
|
|
| BLAKE2b-256 |
427e1b76b28c353cf5fdd2d9e32915ba13404a47072a61cadb3a3aecd999a706
|
File details
Details for the file gpgi-0.7.2-cp310-cp310-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp310-cp310-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 1.2 MB
- Tags: CPython 3.10, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dafded8e6c14fa03439384e77cf629b993ba8f3171032c2a642d8fc1d7ffbfb9
|
|
| MD5 |
11f10d978361f6210ef75886dc2673c5
|
|
| BLAKE2b-256 |
d944a7e2756808d1b2a72773e654cd53d274d1e6c598b70d60d411de088ddb95
|
File details
Details for the file gpgi-0.7.2-cp310-cp310-macosx_10_9_x86_64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp310-cp310-macosx_10_9_x86_64.whl
- Upload date:
- Size: 242.9 kB
- Tags: CPython 3.10, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
966f5af3180665e883628873f8204928fcc8c04ffa2f2204885a8d362dc1ba00
|
|
| MD5 |
2ddf8e29f5fea905d4ca0d18fed0ba87
|
|
| BLAKE2b-256 |
b0b3968d2529f831844c0a38f50533bdcfd52d7324a31e83daf60d2a250edaf7
|
File details
Details for the file gpgi-0.7.2-cp39-cp39-win_amd64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp39-cp39-win_amd64.whl
- Upload date:
- Size: 204.8 kB
- Tags: CPython 3.9, Windows x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
beb4117649e05f6fc04789cf464a3b6edb4cc23cf5e9577a19be790af088af7f
|
|
| MD5 |
b27daeffc95c56e676ed6a8fa7e752af
|
|
| BLAKE2b-256 |
79a896ef2ca2f28924bb9215d16be1705bbd0f9b39bb9ac7bb1bb342bece2d01
|
File details
Details for the file gpgi-0.7.2-cp39-cp39-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp39-cp39-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 1.3 MB
- Tags: CPython 3.9, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3c72728758bf4eddd5588163dffdbe6b2b442bdef811f2012604ebf7a0b2cc95
|
|
| MD5 |
b942988a213c796bd36b60ed94043d71
|
|
| BLAKE2b-256 |
277de50549d60ca69310767437c5ceb94fc6df2fc22e89f8c525f46526c0f5e0
|
File details
Details for the file gpgi-0.7.2-cp39-cp39-macosx_10_9_x86_64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp39-cp39-macosx_10_9_x86_64.whl
- Upload date:
- Size: 241.5 kB
- Tags: CPython 3.9, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
05ea088534a9d1cc7c38367ff42c8ff54bffd340140e607a2969136974ad5be5
|
|
| MD5 |
104c3267eab23545045ec24715127caf
|
|
| BLAKE2b-256 |
3e4e37d611d6c8ccf7f77d14df5a9edc07bc0f5b5ba97cae999dde72764046fa
|
File details
Details for the file gpgi-0.7.2-cp38-cp38-win_amd64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp38-cp38-win_amd64.whl
- Upload date:
- Size: 205.2 kB
- Tags: CPython 3.8, Windows x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
89f4960f90f96d74098df34972fbb243db6549421d0b88ff45d596902fb79d21
|
|
| MD5 |
dddc66a4d137ae8354aef1ba637febab
|
|
| BLAKE2b-256 |
49d06e173b892e75b54ed6269f9a057c79b394bcdc8201bc2bccd72b1f248efc
|
File details
Details for the file gpgi-0.7.2-cp38-cp38-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp38-cp38-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 1.3 MB
- Tags: CPython 3.8, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c3637d30213f2d352c36cfd3cb1c6873c973adf50474b2b3c0488059a074af87
|
|
| MD5 |
7476fd6ef8d9348c83098bc3c13598fb
|
|
| BLAKE2b-256 |
05f11af177eeec9d08b6e403c27cb926e4c1e1eeaa4b5f58261cbe97972df2e7
|
File details
Details for the file gpgi-0.7.2-cp38-cp38-macosx_10_9_x86_64.whl.
File metadata
- Download URL: gpgi-0.7.2-cp38-cp38-macosx_10_9_x86_64.whl
- Upload date:
- Size: 238.4 kB
- Tags: CPython 3.8, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.1 CPython/3.9.15
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
4026be3aff8d97fc591c8a162770083c528cad1a09a1acf9f90265587fc95c15
|
|
| MD5 |
a54320cd7fa8998f9897463c2cedff73
|
|
| BLAKE2b-256 |
95151be705089856a5ddd77ea41390d26d5c7b263bf2dd44bdca246f4e3a23b9
|















