Create visual representations of BLASTn alignments
Project description

🧬 Create visual representations of BLASTn alignments
HomologyViz is a Python-based web app for visualizing pairwise BLASTn alignments between DNA sequences with gene annotations and customizable color maps. It allows you to explore homologous regions and gene structures interactively, with publication-ready output.
HomologyViz reads GenBank files (.gb), uses the CDS features to plot genes as arrows, and performs local BLASTn alignments between sequences. Homology regions are colored by identity percentage using a flexible, user-selectable color scale.
You may optionally customize gene colors directly in the GenBank file by adding a /Color tag to a CDS feature. For example, /Color="#00ff00" will render that gene in green. You can also edit gene colors interactively using the web interface.
HomologyViz is ideal for researchers and students with little or no coding experience. It runs locally and requires no cloud deployment or server hosting.
🚀 Features
- Interactive plots of aligned DNA sequences
- Gene annotations rendered as directional arrows
- Homology regions colored by % identity using customizable color scales
- Click-to-select genes or regions and change their color
- Exportable visualizations (SVG, PNG, PDF, HTML)
- Built with Dash and Plotly
🖼 Example
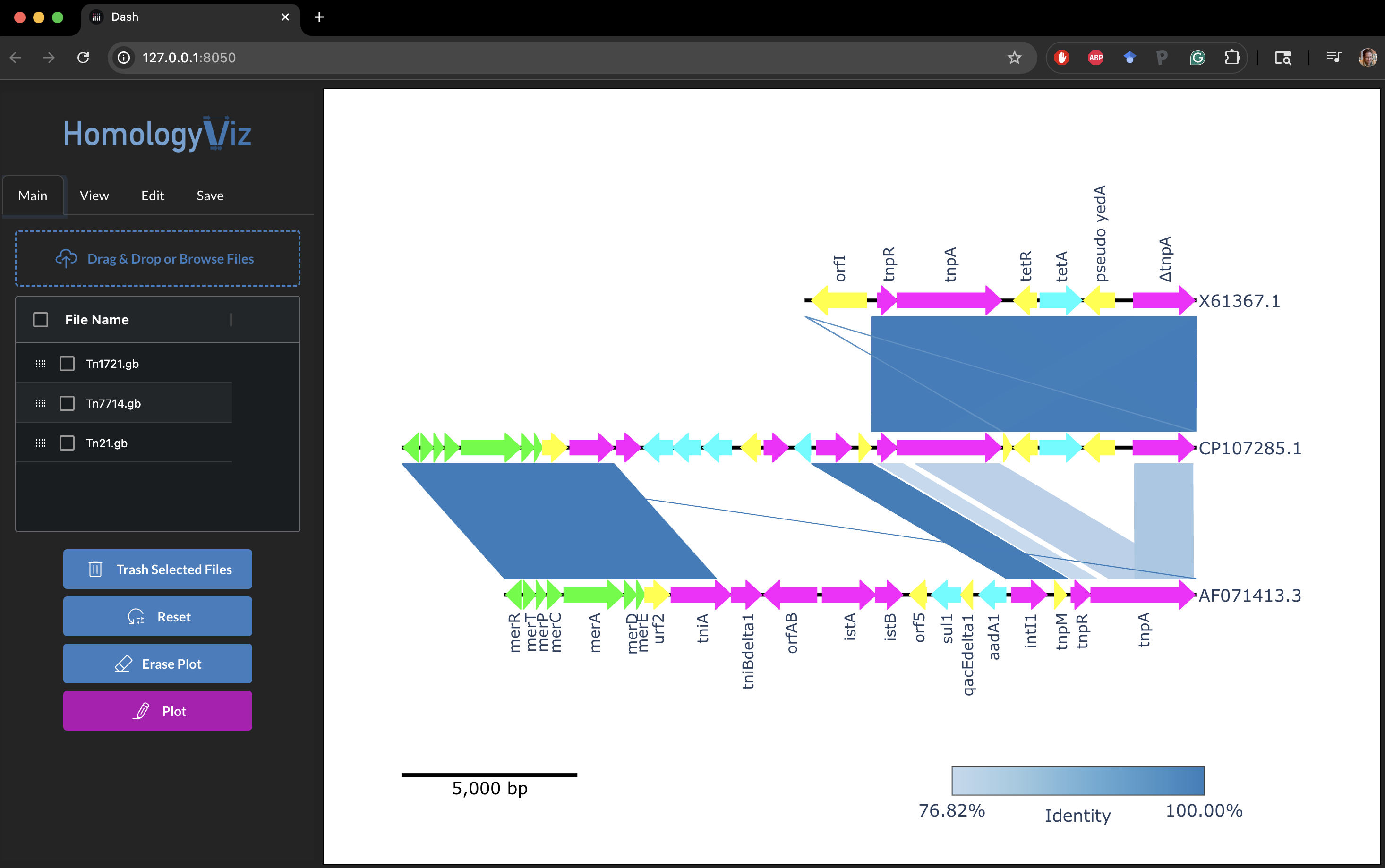
✅ Requirements
HomologyViz has been tested in Google Chrome on macOS, but should work across all modern browsers and platforms.
📦 Installation
You can install HomologyViz using pip:
pip install homologyviz
Or clone this repository:
git clone https://github.com/yourusername/homologyviz.git
cd homologyviz
pip install -e .
🧪 Usage
Once installed, launch the app by typing:
homologyviz
📚 Documentation
You can find the full documentation for HomologyViz at:
👉 https://homologyviz.readthedocs.io
👤 Author
Created by Iván Muñoz Gutiérrez, PhD
University of California, Irvine
School of Biological Sciences
Github: @ivanmugu
🙏 Acknowledgments
- Built on Dash, Plotly, and Biopython
- Inspired by easyfig: Sullivan et al (2011) Bioinformatics 27(7):1009-1010
📄 License
BSD 3-Clause License
📝 Notes
I am developing HomologyViz in my free time, so if you find a bug, it may take me some time to fix it. However, I will fix the problems as soon as possible. Also, if you have any suggestions, let me know, and I will try to implement them.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file homologyviz-0.1.33.tar.gz.
File metadata
- Download URL: homologyviz-0.1.33.tar.gz
- Upload date:
- Size: 479.9 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.11.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
580f2360ecbc07dde376df02647130bc1bf240824560f39ecd114ac69faabcf0
|
|
| MD5 |
1a8b2effe341534439495812be6f9bab
|
|
| BLAKE2b-256 |
251805f65bcdc98bf846910292b9c9430b5d2daa08e740a6e4d5fe37f476606e
|
File details
Details for the file homologyviz-0.1.33-py3-none-any.whl.
File metadata
- Download URL: homologyviz-0.1.33-py3-none-any.whl
- Upload date:
- Size: 422.5 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.1.0 CPython/3.11.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
b72463db3325d7de374b568d0ee23f1bb2f8f6e52423e84148cd493f0f8f4650
|
|
| MD5 |
4cae227aa7643824703782188f22dde3
|
|
| BLAKE2b-256 |
d4b47d2426833a27633a5e535dacf30bbc5b22e9e9ac36fc9bd1af72a3eda891
|















