Make graphical representations of BLASTn alignments
Project description



Make a graphical representation of BLASTn alignments
Homology Visualization (HomologyViz) uses GenBank files (.gb) to align the sequences and plot the genes. HomologyViz uses the information from the CDS features section to plot the genes. To customize the colors for plotting genes, you can add a Color tag in the CDS features with a color in hexadecimal. For example, add the tag /Color="#00ff00" to show a green gene. Or, you can edit the colors interactively in the plot.
HomologyViz is an easy-to-use option for people with little coding knowledge. The program uses Dash for an interactive and web-friendly experience. HomologyViz is a flexible app that allows you to export your graph in different formats and sizes for publication quality.
Requirements
- blastn must be installed locally and in the path
HomologyViz has been tested in Chrome using macOS.
Installation
First, create a virtual environment with conda or venv. Then, install
homologyviz using pip as follows:
pip install homologyviz
Usage
To run the app type:
homologyviz
Output:
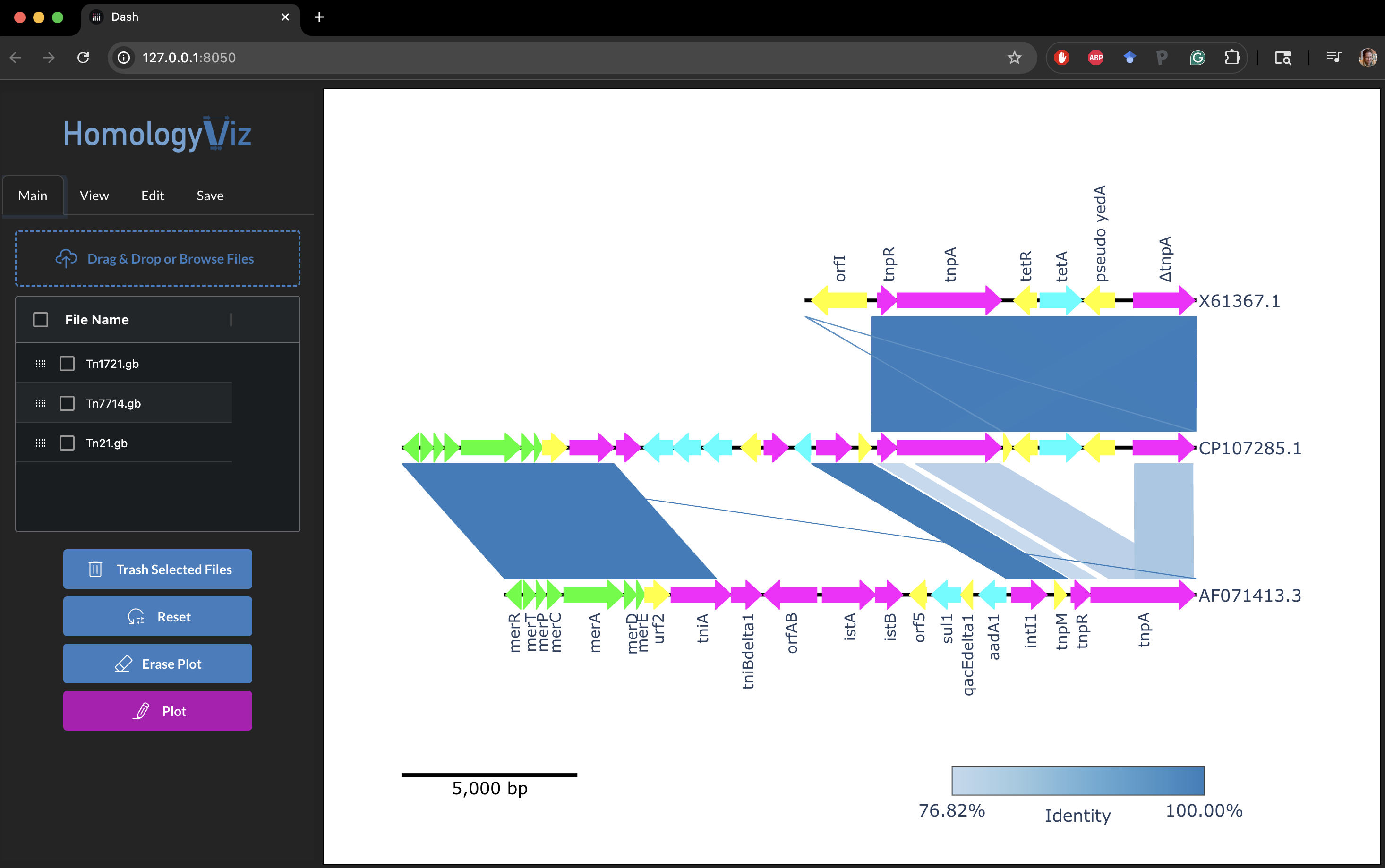
Credits
Inspired by easyfig: Sullivan et al (2011) Bioinformatics 27(7):1009-1010
License
BSD 3-Clause License
Notes
I am developing HomologyViz in my free time, so if you find a bug, it may take me some time to fix it. However, I will fix the problems as soon as possible. Also, if you have any suggestions, let me know, and I will try to implement them.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file homologyviz-0.0.20-py3-none-any.whl.
File metadata
- Download URL: homologyviz-0.0.20-py3-none-any.whl
- Upload date:
- Size: 254.6 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/5.1.1 CPython/3.11.3
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
badd26bb1f03819415e3cb8db5a5e58aeae7c47d12d5cd803a7230b7805a5ef9
|
|
| MD5 |
d3f1625f4c71b571da324005eb7e5428
|
|
| BLAKE2b-256 |
276b0aa9615e384e1aa0f62183f10682f9bbcdb80162d078d2e164f9ff9331c8
|












