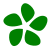
Analyze antibody repertoires and discover new V genes
Project description



IgDiscover
IgDiscover analyzes antibody repertoires and discovers new V genes from high-throughput sequencing reads. Heavy chains, kappa and lambda light chains are supported (to discover VH, VK and VL genes).
IgDiscover is the result of a collaboration between the Gunilla Karlsson Hedestam group at the Department of Microbiology, Tumor and Cell Biology at Karolinska Institutet, Sweden and the Bioinformatics Long-Term Support facility at Science for Life Laboratory (SciLifeLab), Sweden.
If you use IgDiscover, please cite:
Corcoran, Martin M. and Phad, Ganesh E. and Bernat, Néstor Vázquez and Stahl-Hennig, Christiane and Sumida, Noriyuki and Persson, Mats A.A. and Martin, Marcel and Karlsson Hedestam, Gunilla B.Production of individualized V gene databases reveals high levels of immunoglobulin genetic diversity.Nature Communications 7:13642 (2016)
Links
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file igdiscover-0.15.1.tar.gz.
File metadata
- Download URL: igdiscover-0.15.1.tar.gz
- Upload date:
- Size: 253.3 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.0 CPython/3.9.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
534f0e81b71acabe24577f0989832c61e69b73e0e562eb147b1af38c4c9bf725
|
|
| MD5 |
cb3cd57b87b9ee8e1b35ff71d72791fc
|
|
| BLAKE2b-256 |
7b98383bc2105c3945e1908175338bd9792f0ef627f4128301013e6ee51d5753
|
File details
Details for the file igdiscover-0.15.1-py3-none-any.whl.
File metadata
- Download URL: igdiscover-0.15.1-py3-none-any.whl
- Upload date:
- Size: 109.8 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.0 CPython/3.9.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7bb27ae586d8113ffa04388ebafe341cdcb007c072a6522312d71420b4044edd
|
|
| MD5 |
2ea07726de507a29eed56b78f0d2f879
|
|
| BLAKE2b-256 |
6470796a2bddb7eb30edcb13d92d3a245360fff1a250d250400ed689020ae4dd
|












