Framework for automatic classification and segmentation of hyperspectral images.
Project description
Hyperspectral Tissue Classification
This package is a framework for automated tissue classification and segmentation on medical hyperspectral imaging (HSI) data. It contains:
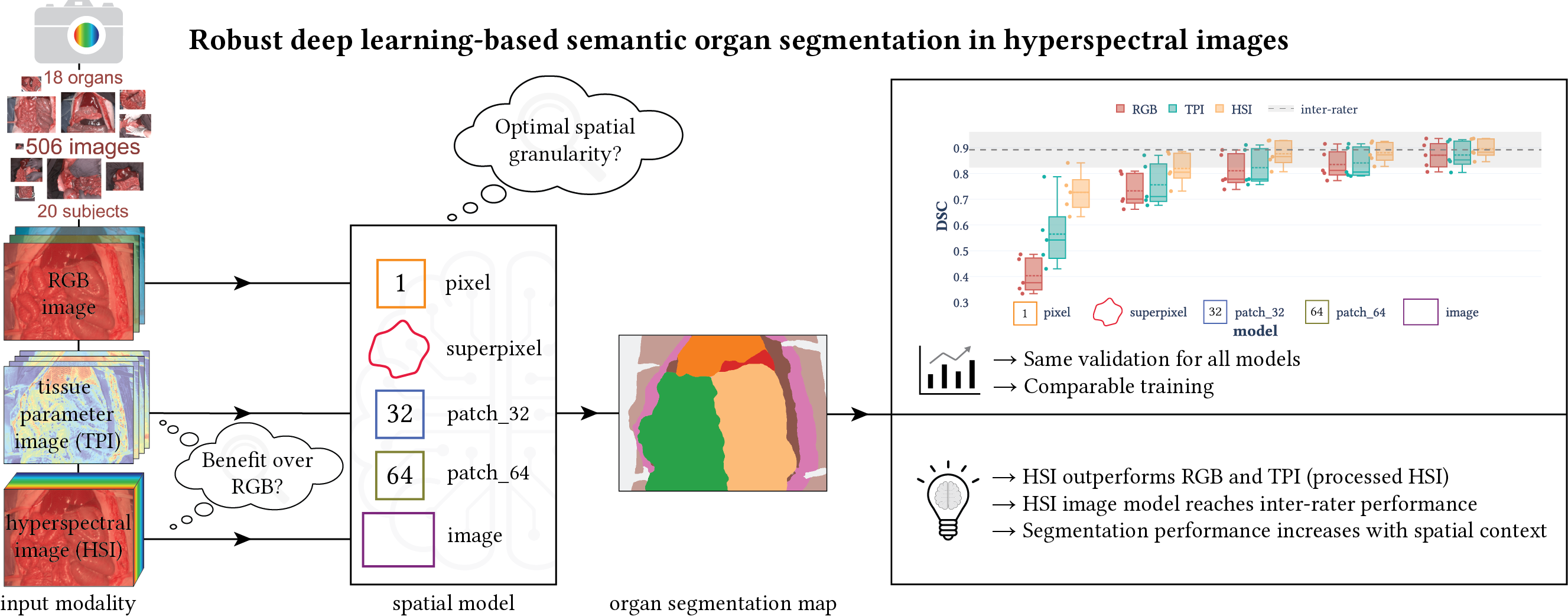
- The implementation of deep learning models to solve supervised classification and segmentation problems for a variety of different input spatial granularities (pixels, superpixels, patches and entire images, cf. figure below) and modalities (RGB data, raw and processed HSI data) from our paper “Robust deep learning-based semantic organ segmentation in hyperspectral images”. It is based on PyTorch and PyTorch Lightning.
- Corresponding pretrained models.
- A pipeline to efficiently load and process HSI data, to aggregate deep learning results and to validate and visualize findings.
- Presentation of several solutions to speed up the data loading process (see Pytorch Conference 2023 poster details below).
This framework is designed to work on HSI data from the Tivita cameras but you can adapt it to different HSI datasets as well. Potential applications include:
- Use our data loading and processing pipeline to easily access image and meta data for any work utilizing Tivita datasets.
- This repository is tightly coupled to work with the public HeiPorSPECTRAL dataset. If you already downloaded the data, you only need to perform the setup steps and then you can directly use the
htcframework to work on the data (cf. our tutorials). - Train your own networks and benefit from a pipeline offering e.g. efficient data loading, correct hierarchical aggregation of results and a set of helpful visualizations.
- Apply deep learning models for different spatial granularities and modalities on your own semantically annotated dataset.
- Use our pretrained models to initialize the weights for your own training.
- Use our pretrained models to generate predictions for your own data.
If you use the htc framework, please cite our paper “Robust deep learning-based semantic organ segmentation in hyperspectral images”.
Cite via BibTeX
@article{SEIDLITZ2022102488,
title = {Robust deep learning-based semantic organ segmentation in hyperspectral images},
journal = {Medical Image Analysis},
volume = {80},
pages = {102488},
year = {2022},
issn = {1361-8415},
doi = {10.1016/j.media.2022.102488},
url = {https://www.sciencedirect.com/science/article/pii/S1361841522001359},
author = {Silvia Seidlitz and Jan Sellner and Jan Odenthal and Berkin Özdemir and Alexander Studier-Fischer and Samuel Knödler and Leonardo Ayala and Tim J. Adler and Hannes G. Kenngott and Minu Tizabi and Martin Wagner and Felix Nickel and Beat P. Müller-Stich and Lena Maier-Hein}
}
Setup
Package Installation
This package can be installed via pip:
pip install imsy-htc
This installs all the required dependencies defined in requirements.txt. The requirements include PyTorch, so you may want to install it manually before installing the package in case you have specific needs (e.g. CUDA version).
⚠️ This framework was developed and tested using the Ubuntu 20.04+ Linux distribution. Despite we do provide wheels for Windows and macOS as well, they are not tested.
⚠️ Network training and inference was conducted using an RTX 3090 GPU with 24 GiB of memory. It should also work with GPUs which have less memory but you may have to adjust some settings (e.g. the batch size).
PyTorch Compatibility
We cannot provide wheels for all PyTorch versions. Hence, a version of imsy-htc may not work with all versions of PyTorch due to changes in the ABI. In the following table, we list the PyTorch versions which are compatible with the respective imsy-htc version.
imsy-htc |
torch |
|---|---|
| 0.0.9 | 1.13 |
| 0.0.10 | 1.13 |
| 0.0.11 | 2.0 |
| 0.0.12 | 2.0 |
| 0.0.13 | 2.1 |
| 0.0.14 | 2.1 |
However, we do not make explicit version constraints in the dependencies of the imsy-htc package because a future version of PyTorch may still work and we don't want to break the installation if it is not necessary.
💡 Please note that it is always possible to build the
imsy-htcpackage with your installed PyTorch version yourself (cf. Developer Installation).
Optional Dependencies (imsy-htc[extra])
Some requirements are considered optional (e.g. if they are only needed by certain scripts) and you will get an error message if they are needed but unavailable. You can install them via
pip install --extra-index-url https://read_package:CnzBrgDfKMWS4cxf-r31@git.dkfz.de/api/v4/projects/15/packages/pypi/simple imsy-htc[extra]
or by adding the following lines to your requirements.txt
--extra-index-url https://read_package:CnzBrgDfKMWS4cxf-r31@git.dkfz.de/api/v4/projects/15/packages/pypi/simple
imsy-htc[extra]
This installs the optional dependencies defined in requirements-extra.txt, including for example our Python wrapper for the challengeR toolkit.
Docker
We also provide a Docker setup for testing. As a prerequisite:
- Clone this repository
- Install Docker and the NVIDIA Container Toolkit
- Install the required dependencies to run the Docker startup script:
pip install python-dotenv
Make sure that your environment variables are available and then bash into the container
export PATH_Tivita_HeiPorSPECTRAL="/path/to/the/dataset"
python run_docker.py bash
You can now run any commands you like. All datasets you provided via an environment variable that starts with PATH_Tivita will be accessible in your container (you can also check the generated docker-compose.override.yml file for details). Please note that the Docker container is meant for small testing only and not for development. This is also reflected by the fact that per default all results are stored inside the container and hence will also be deleted after exiting the container. If you want to keep your results, let the environment variable PATH_HTC_DOCKER_RESULTS point to the directory where you want to store the results.
Developer Installation
If you want to make changes to the package code (which is highly welcome 😉), we recommend to install the htc package in editable mode in a separate conda environment:
# Set up the conda environment
# Note: By adding conda-forge to your default channels, you will get the latest patch releases for Python:
# conda config --add channels conda-forge
conda create --yes --name htc python=3.11
conda activate htc
# Install the htc package and its requirements
pip install -r requirements-dev.txt
pip install --no-use-pep517 -e .
Before commiting any files, please run the static code checks locally:
git add .
pre-commit run --all-files
Environment Variables
This framework can be configured via environment variables. Most importantly, we need to know where your data is located (e.g. PATH_Tivita_HeiPorSPECTRAL) and where results should be stored (e.g. PATH_HTC_RESULTS). For a full list of possible environment variables, please have a look at the documentation of the Settings class.
💡 If you set an environment variable for a dataset path, it is important that the variable name matches the folder name (e.g. the variable name
PATH_Tivita_HeiPorSPECTRALmatches the dataset pathmy/path/HeiPorSPECTRALwith its folder nameHeiPorSPECTRAL, whereas the variable namePATH_Tivita_some_other_namedoes not match). Furthermore, the dataset path needs to point to a directory which contains adataand anintermediatessubfolder.
There are several options to set the environment variables. For example:
- You can specify a variable as part of your bash startup script
~/.bashrcor before running each command:PATH_HTC_RESULTS="~/htc/results" htc training --model image --config "models/image/configs/default"
However, this might get cumbersome or might not give you the flexibility you need. - Recommended if you cloned this repository (in contrast to simply installing it via pip): You can create a
.envfile in the repository root and add your variables, for example:export PATH_Tivita_HeiPorSPECTRAL=/mnt/nvme_4tb/HeiPorSPECTRAL export PATH_HTC_RESULTS=~/htc/results
- Recommended if you installed the package via pip: You can create user settings for this application. The location is OS-specific. For Linux the location might be at
~/.config/htc/variables.env. Please runhtc infoupon package installation to retrieve the exact location on your system. The content of the file is of the same format as of the.envabove.
After setting your environment variables, it is recommended to run htc info to check that your variables are correctly registered in the framework.
Tutorials
A series of tutorials can help you get started on the htc framework by guiding you through different usage scenarios.
💡 The tutorials make use of our public HSI dataset HeiPorSPECTRAL. If you want to directly run them, please download the dataset first and make it accessible via the environment variable
PATH_Tivita_HeiPorSPECTRALas described above.
- As a start, we recommend to take a look at this general notebook which showcases the basic functionalities of the
htcframework. Namely, it demonstrates the usage of theDataPathclass which is the entry point to load and process HSI data. For example, you will learn how to read HSI cubes, segmentation masks and meta data. Among others, you can use this information to calculate the median spectrum of an organ. - If you want to use our framework with your own dataset, it might be necessary to write a custom
DataPathclass so that you can load and process your images and annotations. We collected some tips on how this can be achieved. - You have some HSI data at hand and want to use one of our pretrained models to generate predictions? Then our prediction notebook has got you covered.
- You want to use our pretrained models to initialize the weights for your own training? See the section about pretrained models below for details.
- You want to use our framework to train a network? The network training notebook will show you how to achieve this on the example of a heart and lung segmentation network.
- If you are interested in our technical validation (e.g. because you want to compare your colorchecker images with ours) and need to create a mask to detect the different colorchecker fields, you might find our automatic colorchecker mask creation pipeline useful.
We do not have a separate documentation website for our framework yet. However, most of the functions and classes are documented so feel free to explore the source code or use your favorite IDE to display the documentation. If something does not become clear from the documentation, feel free to open an issue!
Pretrained Models
This framework gives you access to a variety of pretrained segmentation and classification models. The models will be automatically downloaded, provided you specify the model type (e.g. image) and the run folder (e.g. 2022-02-03_22-58-44_generated_default_model_comparison). It can then be used for example to create predictions on some data or as a baseline for your own training (see example below).
The following table lists all the models you can get:
💡 The modality
paramrefers to stacked tissue parameter images (named TPI in our paper “Robust deep learning-based semantic organ segmentation in hyperspectral images”). For the model typepatch, pretrained models are available for the patch sizes 64 x 64 and 32 x 32 pixels. The modality and patch size is not specified when loading a model as it is already characterized by specifying a certain run folder.
After successful installation of the htc package, you can use any of the pretrained models listed in the table. There are several ways to use them but the general principle is that models are always specified via their model and run_folder.
Option 1: Use the models in your own training pipeline
Every model class listed in the table has a static method pretrained_model() which you can use to create a model instance and initialize it with the pretrained weights. The model object will be an instance of torch.nn.Module. The function has examples for all the different model types but as a teaser consider the following example which loads the pretrained image HSI network:
import torch
from htc import ModelImage, Normalization
run_folder = "2022-02-03_22-58-44_generated_default_model_comparison" # HSI model
model = ModelImage.pretrained_model(model="image", run_folder=run_folder, n_channels=100, n_classes=19)
input_data = torch.randn(1, 100, 480, 640) # NCHW
input_data = Normalization(channel_dim=1)(input_data) # Model expects L1 normalized input
model(input_data).shape
# torch.Size([1, 19, 480, 640])
💡 Please note that when initializing the weights as in this example, the segmentation head is initialized randomly. Meaningful predictions on your own data can thus not be expected out of the box, but you will have to train the model on your data first.
Option 2: Use the models to create predictions for your data
The models can be used to predict segmentation masks for your data. The segmentation models automatically sample from your input image according to the selected model spatial granularity (e.g. by creating patches) and the output is always a segmentation mask for an entire image. The set of output classes is determined by the training configuration, e.g. 18 organ classes + background for our semantic models. The CreatingPredictions notebook shows how to create predictions and how to map the network output to meaningful label names.
Option 3: Use the models to train a network with the htc package
If you are using the htc framework to train your networks, you only need to define the model in your configuration:
{
"model": {
"pretrained_model": {
"model": "image",
"run_folder": "2022-02-03_22-58-44_generated_default_model_comparison",
}
}
}
This is very similar to option 1 but may be more convenient if you train with the htc framework.
CLI
There is a common command line interface for many scripts in this repository. More precisely, every script which is prefixed with run_NAME.py can also be run via htc NAME from any directory. For more details, just type htc.
Papers
This repository contains code to reproduce our publications listed below:
📝 Robust deep learning-based semantic organ segmentation in hyperspectral images
In this paper, we tackled fully automatic organ segmentation and compared deep learning models on different spatial granularities (e.g. patch vs. image) and modalities (e.g. HSI vs. RGB). Furthermore, we studied the required amount of training data and the generalization capabilities of our models across subjects. The pretrained networks are related to this paper. You can find the notebooks to generate the paper figures in paper/MIA2021 (the folder also includes a reproducibility document) and the models in htc/models. For each model, there are three configuration files, namely default, default_rgb and default_parameters, which correspond to the HSI, RGB and TPI modality, respectively. You can also download the NSD thresholds which we used for the NSD metric (cf. Fig. 12).
📂 The dataset for this paper is not publicly available.
Cite via BibTeX
@article{seidlitz_seg_2022,
author = {Seidlitz, Silvia and Sellner, Jan and Odenthal, Jan and Özdemir, Berkin and Studier-Fischer, Alexander and Knödler, Samuel and Ayala, Leonardo and Adler, Tim J. and Kenngott, Hannes G. and Tizabi, Minu and Wagner, Martin and Nickel, Felix and Müller-Stich, Beat P. and Maier-Hein, Lena},
language = {en},
date = {2022-08},
issn = {1361-8415},
journaltitle = {Medical Image Analysis},
keywords = {Hyperspectral imaging,Surgical data science,Deep learning,Open surgery,Organ segmentation,Semantic scene segmentation},
pages = {102488},
title = {Robust deep learning-based semantic organ segmentation in hyperspectral images},
volume = {80},
}
📝 Semantic segmentation of surgical hyperspectral images under geometric domain shifts
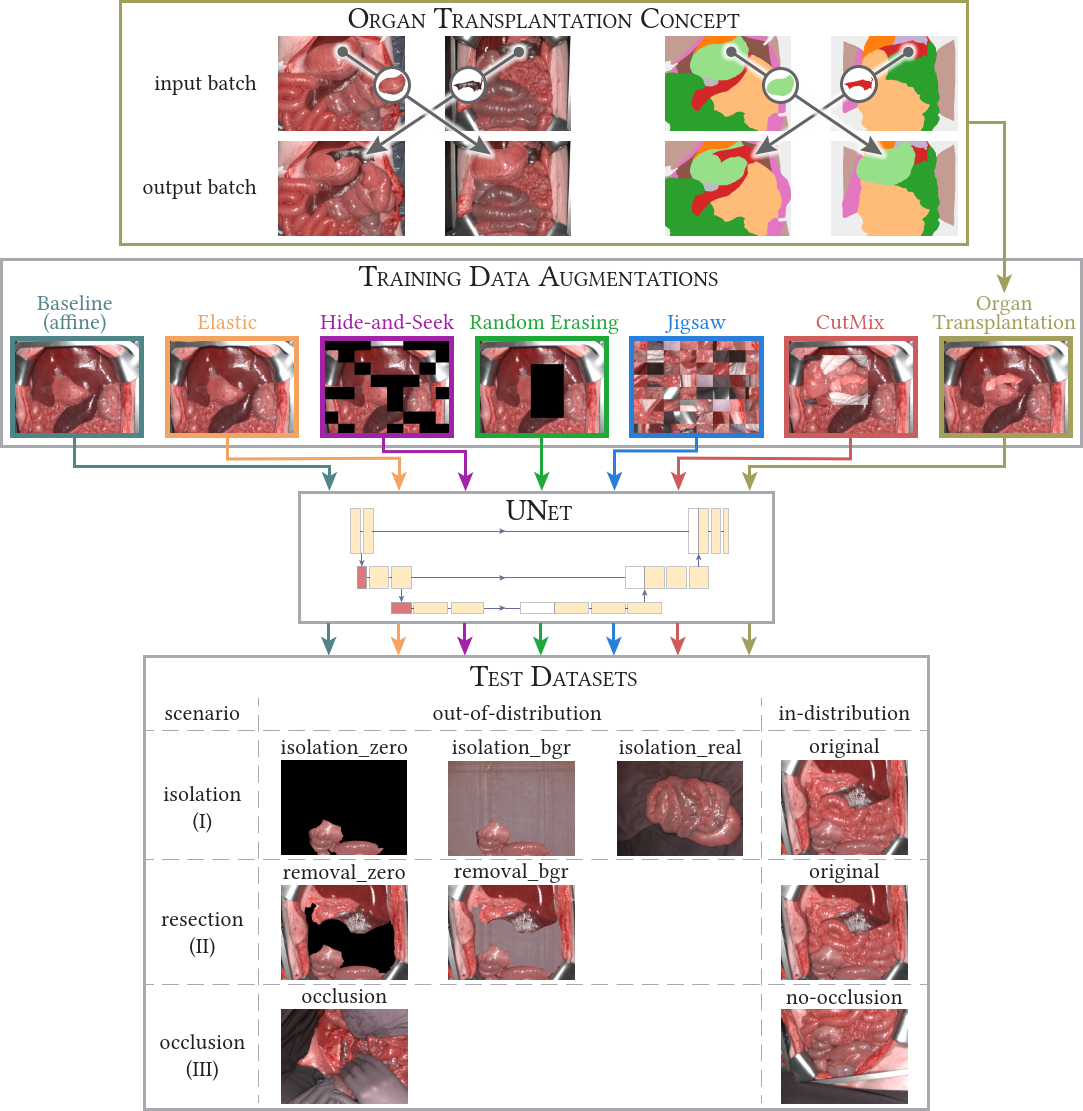
This MICCAI2023 paper is the direct successor of our MIA2021 paper. We analyzed how well our networks perform under geometrical domain shifts which commonly occur in real-world open surgeries (e.g. situs occlusions). The effect is drastic (drop of Dice similarity coefficient by 45 %) but the good news is that performance on par with in-distribution data can be achieved with our simple, model-independent solution (augmentation method). You can find all the code in htc/context and paper figures as well as reproducibility instructions in paper/MICCAI2023. Pretrained models are available for our organ transplantation networks with HSI and RGB modalities.
💡 If you are only interested in our data augmentation method, you can also head over to Kornia where this augmentation is implemented for generic use cases (including 2D and 3D data). You will find it under the name
RandomTransplantation.
📂 The dataset for this paper is not publicly available.
Cite via BibTeX
@inproceedings{sellner_context_2023,
author = {Sellner, Jan and Seidlitz, Silvia and Studier-Fischer, Alexander and Motta, Alessandro and Özdemir, Berkin and Müller-Stich, Beat Peter and Nickel, Felix and Maier-Hein, Lena},
editor = {Greenspan, Hayit and Madabhushi, Anant and Mousavi, Parvin and Salcudean, Septimiu and Duncan, James and Syeda-Mahmood, Tanveer and Taylor, Russell},
location = {Cham},
publisher = {Springer Nature Switzerland},
booktitle = {Medical Image Computing and Computer Assisted Intervention -- MICCAI 2023},
date = {2023},
isbn = {978-3-031-43996-4},
pages = {618--627},
title = {Semantic Segmentation of Surgical Hyperspectral Images Under Geometric Domain Shifts},
}
📝 Dealing with I/O bottlenecks in high-throughput model training
The poster was presented at the PyTorch Conference 2023 and presents several solutions to improve data loading for faster network training. This originated from our MICCAI2023 paper, where we load huge amount of data while using a relatively small network resulting in GPU idle times when the GPU has to wait for the CPU to deliver new data. This requested the need for fast data loading strategies so that the CPU delivers data in-time for the GPU. The solutions include (1) efficient data storage via Blosc compression, (2) appropriate precision settings, (3) GPU instead of CPU augmentations using the Kornia library and (4) a fixed shared pinned memory buffer for efficient data transfer to the GPU. For the last part, you will find the relevant code to create the buffer in this repository as part of the SharedMemoryDatasetMixin class (_add_tensor_shared() method).
You can find the code to generate the results figures of the poster in paper/PyTorchConf2023 including reproducibility instructions.
📂 The dataset for this poster is not publicly available.
Cite via BibTeX
@misc{sellner_benchmarking_2023,
author = {Sellner, Jan and Seidlitz, Silvia and Maier-Hein, Lena},
language = {en},
url = {https://e130-hyperspectal-tissue-classification.s3.dkfz.de/figures/PyTorchConference_Poster.pdf},
date = {2023-10-16},
title = {Dealing with I/O bottlenecks in high-throughput model training},
}
📝 Spectral organ fingerprints for machine learning-based intraoperative tissue classification with hyperspectral imaging in a porcine model
In this paper, we trained a classification model based on median spectra from HSI data. You can find the model code in htc/tissue_atlas and the confusion matrix figure of the paper in paper/NatureReports2021 (including a reproducibility document).
📂 The dataset for this paper is not fully publicly available, but a subset of the data is available through the public HeiPorSPECTRAL dataset.
Cite via BibTeX
@article{studierfischer_atlas_2022,
author = {Studier-Fischer, Alexander and Seidlitz, Silvia and Sellner, Jan and Özdemir, Berkin and Wiesenfarth, Manuel and Ayala, Leonardo and Odenthal, Jan and Knödler, Samuel and Kowalewski, Karl Friedrich and Haney, Caelan Max and Camplisson, Isabella and Dietrich, Maximilian and Schmidt, Karsten and Salg, Gabriel Alexander and Kenngott, Hannes Götz and Adler, Tim Julian and Schreck, Nicholas and Kopp-Schneider, Annette and Maier-Hein, Klaus and Maier-Hein, Lena and Müller-Stich, Beat Peter and Nickel, Felix},
url = {https://doi.org/10.1038/s41598-022-15040-w},
date = {2022-06-30},
doi = {10.1038/s41598-022-15040-w},
issn = {2045-2322},
journaltitle = {Scientific Reports},
number = {1},
pages = {11028},
title = {Spectral organ fingerprints for machine learning-based intraoperative tissue classification with hyperspectral imaging in a porcine model},
volume = {12},
}
📝 HeiPorSPECTRAL - the Heidelberg Porcine HyperSPECTRAL Imaging Dataset of 20 Physiological Organs
This paper introduces the HeiPorSPECTRAL dataset containing 5756 hyperspectral images from 11 subjects. We are using these images in our tutorials. You can find the visualization notebook for the paper figures in paper/NatureData2023 (the folder also includes a reproducibility document) and the remaining code in htc/tissue_atlas_open.
If you want to learn more about the HeiPorSPECTRAL dataset (e.g. the underlying data structure) or you stumbled upon a file and want to know how to read it, you might find this notebook with low-level details helpful.
📂 The dataset for this paper is publicly available.
Cite via BibTeX
@article{studierfischer_open_2023,
author = {Studier-Fischer, Alexander and Seidlitz, Silvia and Sellner, Jan and Bressan, Marc and Özdemir, Berkin and Ayala, Leonardo and Odenthal, Jan and Knoedler, Samuel and Kowalewski, Karl-Friedrich and Haney, Caelan Max and Salg, Gabriel and Dietrich, Maximilian and Kenngott, Hannes and Gockel, Ines and Hackert, Thilo and Müller-Stich, Beat Peter and Maier-Hein, Lena and Nickel, Felix},
url = {https://doi.org/10.1038/s41597-023-02315-8},
date = {2023-06-24},
doi = {10.1038/s41597-023-02315-8},
issn = {2052-4463},
journaltitle = {Scientific Data},
number = {1},
pages = {414},
title = {HeiPorSPECTRAL - the Heidelberg Porcine HyperSPECTRAL Imaging Dataset of 20 Physiological Organs},
volume = {10},
}
📝 Künstliche Intelligenz und hyperspektrale Bildgebung zur bildgestützten Assistenz in der minimal-invasiven Chirurgie
This paper presents several applications of intraoperative HSI, including our organ segmentation and classification work. You can find the code generating our figure for this paper at paper/Chirurg2022.
📂 The sample image used here is contained in the dataset from our paper “Robust deep learning-based semantic organ segmentation in hyperspectral images” and hence not publicly available.
Cite via BibTeX
@article{chalopin_chirurgie_2022,
author = {Chalopin, Claire and Nickel, Felix and Pfahl, Annekatrin and Köhler, Hannes and Maktabi, Marianne and Thieme, René and Sucher, Robert and Jansen-Winkeln, Boris and Studier-Fischer, Alexander and Seidlitz, Silvia and Maier-Hein, Lena and Neumuth, Thomas and Melzer, Andreas and Müller-Stich, Beat Peter and Gockel, Ines},
url = {https://doi.org/10.1007/s00104-022-01677-w},
date = {2022-10-01},
doi = {10.1007/s00104-022-01677-w},
issn = {2731-698X},
journaltitle = {Die Chirurgie},
number = {10},
pages = {940--947},
title = {Künstliche Intelligenz und hyperspektrale Bildgebung zur bildgestützten Assistenz in der minimal-invasiven Chirurgie},
volume = {93},
}
Funding
This project has received funding from the European Research Council (ERC) under the European Unions Horizon 2020 research and innovation programme (NEURAL SPICING, grant agreement No. 101002198) and was supported by the German Cancer Research Center (DKFZ) and the Helmholtz Association under the joint research school HIDSS4Health (Helmholtz Information and Data Science School for Health). It further received funding from the Surgical Oncology Program of the National Center for Tumor Diseases (NCT) Heidelberg.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distributions
Built Distributions
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file imsy_htc-0.0.14-cp311-cp311-win_amd64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp311-cp311-win_amd64.whl
- Upload date:
- Size: 2.0 MB
- Tags: CPython 3.11, Windows x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
d03aa73df3e189aae602684213a05bb3b6932232d0de89ab70b5d15396ecb310
|
|
| MD5 |
0671d61a069d2b5ca2196214e65ba034
|
|
| BLAKE2b-256 |
d503a59fc0c903a989a7a2c1ed5c67d6cd7dbe6686ed9432567547c847f66d66
|
File details
Details for the file imsy_htc-0.0.14-cp311-cp311-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp311-cp311-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 13.3 MB
- Tags: CPython 3.11, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
dce8a614ae90c9e91866b94035480a1210d5d6f611fe7c625b26eb3a4d7e110d
|
|
| MD5 |
61ae6de5575b85f10c0ba0bad8425fac
|
|
| BLAKE2b-256 |
291ee966524fa5731df1d097a1cbc142142cfa6db308915d5af9cc43fb8459e6
|
File details
Details for the file imsy_htc-0.0.14-cp311-cp311-macosx_11_0_arm64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp311-cp311-macosx_11_0_arm64.whl
- Upload date:
- Size: 2.0 MB
- Tags: CPython 3.11, macOS 11.0+ ARM64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
1502b7f404859349878dab227d27196838925e1c7beda5cce53b067ce320cf19
|
|
| MD5 |
addddca7d3d3d9c026c57f778f05428d
|
|
| BLAKE2b-256 |
7ed5cbebcf8344ff40df1fc0f1180018e46d48f0181880c7a3caef93de72471f
|
File details
Details for the file imsy_htc-0.0.14-cp311-cp311-macosx_10_9_x86_64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp311-cp311-macosx_10_9_x86_64.whl
- Upload date:
- Size: 2.0 MB
- Tags: CPython 3.11, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
01fecbe5e3309ce47017fe3069c2faa869e59d530499951a93c7964d79edee26
|
|
| MD5 |
3e03c36c85afa7007c7d05749b28b326
|
|
| BLAKE2b-256 |
d4915aa874b075d425d72dc863439f7094cc48bba509eee75549373edc59d412
|
File details
Details for the file imsy_htc-0.0.14-cp310-cp310-win_amd64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp310-cp310-win_amd64.whl
- Upload date:
- Size: 2.0 MB
- Tags: CPython 3.10, Windows x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
c0b0cf3ecb2e93b94973764f39ce9add2eed99bf10e401bec4c06167973a8328
|
|
| MD5 |
84f8db6c5d1fde0de3251e7ca024dfb8
|
|
| BLAKE2b-256 |
628bb28d0f7294794a850b7e63d50196566568b2d0e40a5cc54da5064c8bcfe8
|
File details
Details for the file imsy_htc-0.0.14-cp310-cp310-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp310-cp310-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 13.2 MB
- Tags: CPython 3.10, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
57255fd26de52c290436f752fb316e3f85d6d214e3a2c5777f277451aae7b887
|
|
| MD5 |
09b5676b2f54736790c830c9fd7a2c13
|
|
| BLAKE2b-256 |
4b9a7653b031ef0db70dadecb44941fc7926d3c29149e12dfbdcfed3d59f2910
|
File details
Details for the file imsy_htc-0.0.14-cp310-cp310-macosx_11_0_arm64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp310-cp310-macosx_11_0_arm64.whl
- Upload date:
- Size: 2.0 MB
- Tags: CPython 3.10, macOS 11.0+ ARM64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
467dd05d62b6f353129e2b99ab3ec9b7104f8148c9f124249f40d4b077cf6917
|
|
| MD5 |
652e595faf9d91b070eaf487ac221603
|
|
| BLAKE2b-256 |
025a83ec198d955b27eed9d55c95e58dd0369bbd686c207e1f4868acfef03ecb
|
File details
Details for the file imsy_htc-0.0.14-cp310-cp310-macosx_10_9_x86_64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp310-cp310-macosx_10_9_x86_64.whl
- Upload date:
- Size: 2.0 MB
- Tags: CPython 3.10, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
6953bbe2e3b132f94078dd0b615e4a41b4cd83e68abc9d92366d8064cd1f8a5f
|
|
| MD5 |
5caf98c69ac00ac9f879bc5d82d17ce6
|
|
| BLAKE2b-256 |
bced8aa79494dab3699aae0e637e59bb78ddc07183d2efa644e8299d7669b970
|
File details
Details for the file imsy_htc-0.0.14-cp39-cp39-win_amd64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp39-cp39-win_amd64.whl
- Upload date:
- Size: 2.0 MB
- Tags: CPython 3.9, Windows x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
9c9f4108be633611df406eb41e1f08a3f168b9a9fc38e225994403920d770325
|
|
| MD5 |
bdf5d657c0a84182c15be4da9146d867
|
|
| BLAKE2b-256 |
df90a87c55347c027519abc4bf059d46bf1414c1ddd8329419361997601a47b1
|
File details
Details for the file imsy_htc-0.0.14-cp39-cp39-manylinux_2_17_x86_64.manylinux2014_x86_64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp39-cp39-manylinux_2_17_x86_64.manylinux2014_x86_64.whl
- Upload date:
- Size: 13.2 MB
- Tags: CPython 3.9, manylinux: glibc 2.17+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8f5bcd94de00fa0928945920251ec60aecdf48701ee633a94681ab763a422657
|
|
| MD5 |
6458670c84e8e9b3150f2e3c54264bfb
|
|
| BLAKE2b-256 |
f7359c787ca2ba1cdbe6d1353ba2dddf68f2f8016296ac19f8f5f5ba52cb261f
|
File details
Details for the file imsy_htc-0.0.14-cp39-cp39-macosx_11_0_arm64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp39-cp39-macosx_11_0_arm64.whl
- Upload date:
- Size: 2.0 MB
- Tags: CPython 3.9, macOS 11.0+ ARM64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
27aa6db341fcea69f1b178d70e9b216b88e44542789132250abb20ee232af099
|
|
| MD5 |
50788c1c6813a203c92c63616a157cec
|
|
| BLAKE2b-256 |
a40c1e3abd557d5bb3a066c1e8b023f4c8a465b7c0f7fa97ee9029b12f72274c
|
File details
Details for the file imsy_htc-0.0.14-cp39-cp39-macosx_10_9_x86_64.whl.
File metadata
- Download URL: imsy_htc-0.0.14-cp39-cp39-macosx_10_9_x86_64.whl
- Upload date:
- Size: 2.0 MB
- Tags: CPython 3.9, macOS 10.9+ x86-64
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/4.0.2 CPython/3.11.7
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
86c2b230c61b43b6b7687e307701525452ba265d93cca22f5879c1b3d54d9dc3
|
|
| MD5 |
79ecde978aa7772c60cecbdb1154f07c
|
|
| BLAKE2b-256 |
08b03c07f63b0ecbd2fdba677fa1fa9eae6a252b234a5e67035274471a012f78
|