jarvis-tools: an open-source software package for data-driven atomistic materials design. https://jarvis.nist.gov/
Project description
















JARVIS-Tools
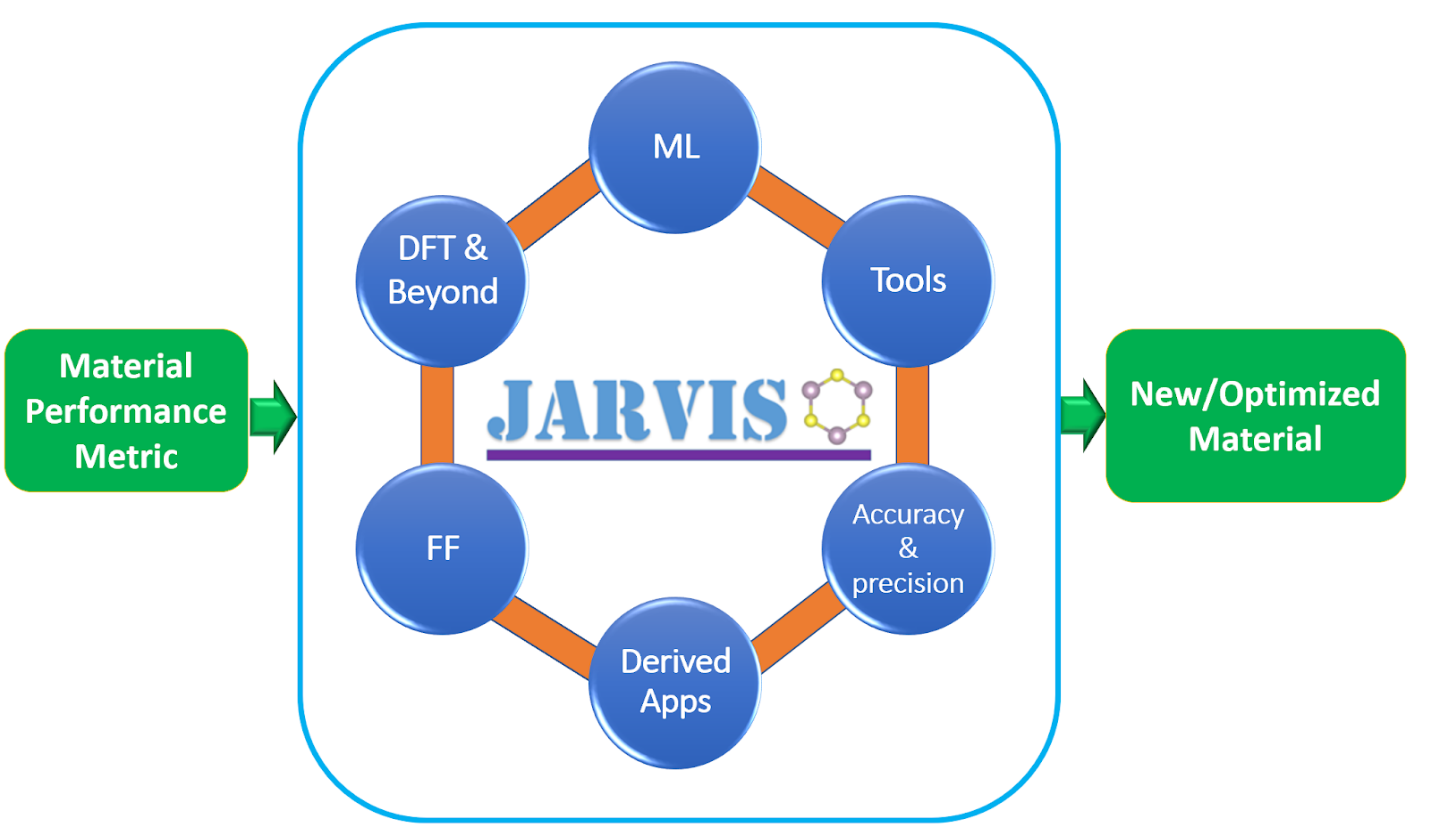
The JARVIS-Tools is an open-access software package for atomistic data-driven materials desgin. JARVIS-Tools can be used for a) setting up calculations, b) analysis and informatics, c) plotting, d) database development and e) web-page development.
JARVIS-Tools empowers NIST-JARVIS (Joint Automated Repository for Various Integrated Simulations) repository which is an integrated framework for computational science using density functional theory, classical force-field/molecular dynamics and machine-learning. The NIST-JARVIS official website is: https://jarvis.nist.gov . This project is a part of the Materials Genome Initiative (MGI) at NIST (https://mgi.nist.gov/).
For more details, checkout our latest article: The joint automated repository for various integrated simulations (JARVIS) for data-driven materials design and YouTube videos
Documentation
Capabilities
Software workflow tasks for preprcessing, executing and post-processing: VASP, Quantum Espresso, Wien2k BoltzTrap, Wannier90, LAMMPS, Scikit-learn, TensorFlow, LightGBM, Qiskit, Tequila, Pennylane, DGL, PyTorch.
Several examples: Notebooks and test scripts to explain the package.
Several analysis tools: Atomic structure, Electronic structure, Spacegroup, Diffraction, 2D materials and other vdW bonded systems, Mechanical, Optoelectronic, Topological, Solar-cell, Thermoelectric, Piezoelectric, Dielectric, STM, Phonon, Dark matter, Wannier tight binding models, Point defects, Heterostructures, Magnetic ordering, Images, Spectrum etc.
Database upload and download: Download JARVIS databases such as JARVIS-DFT, FF, ML, WannierTB, Solar, STM and also external databases such as Materials project, OQMD, AFLOW etc.
Access raw input/output files: Download input/ouput files for JARVIS-databases to enhance reproducibility.
Train machine learning models: Use different descriptors, graphs and datasets for training machine learning models.
HPC clusters: Torque/PBS and SLURM.
Available datasets: Summary of several datasets .
Installation
>>> pip install -U jarvis-tools
or
>>> conda install -c conda-forge jarvis-tools
For detailed instructions, please see Installation instructions
Example function
>>> from jarvis.core.atoms import Atoms
>>> box = [[2.715, 2.715, 0], [0, 2.715, 2.715], [2.715, 0, 2.715]]
>>> coords = [[0, 0, 0], [0.25, 0.25, 0.25]]
>>> elements = ["Si", "Si"]
>>> Si = Atoms(lattice_mat=box, coords=coords, elements=elements)
>>> density = round(Si.density,2)
>>> print (density)
2.33
>>>
>>> from jarvis.db.figshare import data
>>> dft_3d = data(dataset='dft_3d')
>>> print (len(dft_3d))
48527
>>> from jarvis.io.vasp.inputs import Poscar
>>> for i in dft_3d:
... atoms = Atoms.from_dict(i['atoms'])
... poscar = Poscar(atoms)
... jid = i['jid']
... filename = 'POSCAR-'+jid+'.vasp'
... poscar.write_file(filename)
>>> dft_2d = data(dataset='dft_2d')
>>> print (len(dft_2d))
1070
>>> for i in dft_2d:
... atoms = Atoms.from_dict(i['atoms'])
... poscar = Poscar(atoms)
... jid = i['jid']
... filename = 'POSCAR-'+jid+'.vasp'
... poscar.write_file(filename)
>>> # Example to parse DOS data from JARVIS-DFT webpages
>>> from jarvis.db.webpages import Webpage
>>> from jarvis.core.spectrum import Spectrum
>>> import numpy as np
>>> new_dist=np.arange(-5, 10, 0.05)
>>> all_atoms = []
>>> all_dos_up = []
>>> all_jids = []
>>> for ii,i in enumerate(dft_3d):
all_jids.append(i['jid'])
... try:
... w = Webpage(jid=i['jid'])
... edos_data = w.get_dft_electron_dos()
... ens = np.array(edos_data['edos_energies'].strip("'").split(','),dtype='float')
... tot_dos_up = np.array(edos_data['total_edos_up'].strip("'").split(','),dtype='float')
... s = Spectrum(x=ens,y=tot_dos_up)
... interp = s.get_interpolated_values(new_dist=new_dist)
... atoms=Atoms.from_dict(i['atoms'])
... ase_atoms=atoms.ase_converter()
... all_dos_up.append(interp)
... all_atoms.append(atoms)
... all_jids.append(i['jid'])
... filename=i['jid']+'.cif'
... atoms.write_cif(filename)
... break
... except Exception as exp :
... print (exp,i['jid'])
... pass
Find more examples at
Citing
Please cite the following if you happen to use JARVIS-Tools for a publication.
https://www.nature.com/articles/s41524-020-00440-1
Choudhary, K. et al. The joint automated repository for various integrated simulations (JARVIS) for data-driven materials design. npj Computational Materials, 6(1), 1-13 (2020).
References
Please see Publications related to JARVIS-Tools
Correspondence
Please report bugs as Github issues (https://github.com/usnistgov/jarvis/issues) or email to kamal.choudhary@nist.gov.
Funding support
NIST-MGI (https://www.nist.gov/mgi).
Code of conduct
Please see Code of conduct
Module structure
jarvis/ ├── ai │ ├── descriptors │ │ ├── cfid.py │ │ ├── coulomb.py │ ├── gcn │ ├── pkgs │ │ ├── lgbm │ │ │ ├── classification.py │ │ │ └── regression.py │ │ ├── sklearn │ │ │ ├── classification.py │ │ │ ├── hyper_params.py │ │ │ └── regression.py │ │ └── utils.py │ ├── uncertainty │ │ └── lgbm_quantile_uncertainty.py ├── analysis │ ├── darkmatter │ │ └── metrics.py │ ├── defects │ │ ├── surface.py │ │ └── vacancy.py │ ├── diffraction │ │ └── xrd.py │ ├── elastic │ │ └── tensor.py │ ├── interface │ │ └── zur.py │ ├── magnetism │ │ └── magmom_setup.py │ ├── periodic │ │ └── ptable.py │ ├── phonon │ │ ├── force_constants.py │ │ └── ir.py │ ├── solarefficiency │ │ └── solar.py │ ├── stm │ │ └── tersoff_hamann.py │ ├── structure │ │ ├── neighbors.py │ │ ├── spacegroup.py │ ├── thermodynamics │ │ ├── energetics.py │ ├── topological │ │ └── spillage.py ├── core │ ├── atoms.py │ ├── composition.py │ ├── graphs.py │ ├── image.py │ ├── kpoints.py │ ├── lattice.py │ ├── pdb_atoms.py │ ├── specie.py │ ├── spectrum.py │ └── utils.py ├── db │ ├── figshare.py │ ├── jsonutils.py │ ├── lammps_to_xml.py │ ├── restapi.py │ ├── vasp_to_xml.py │ └── webpages.py ├── examples │ ├── lammps │ │ ├── jff_test.py │ │ ├── Al03.eam.alloy_nist.tgz │ ├── vasp │ │ ├── dft_test.py │ │ ├── SiOptb88.tgz ├── io │ ├── boltztrap │ │ ├── inputs.py │ │ └── outputs.py │ ├── calphad │ │ └── write_decorated_poscar.py │ ├── lammps │ │ ├── inputs.py │ │ └── outputs.py │ ├── pennylane │ │ ├── inputs.py │ ├── phonopy │ │ ├── fcmat2hr.py │ │ ├── inputs.py │ │ └── outputs.py │ ├── qe │ │ ├── inputs.py │ │ └── outputs.py │ ├── qiskit │ │ ├── inputs.py │ ├── tequile │ │ ├── inputs.py │ ├── vasp │ │ ├── inputs.py │ │ └── outputs.py │ ├── wannier │ │ ├── inputs.py │ │ └── outputs.py │ ├── wanniertools │ │ ├── inputs.py │ │ └── outputs.py │ ├── wien2k │ │ ├── inputs.py │ │ ├── outputs.py ├── tasks │ ├── boltztrap │ │ └── run.py │ ├── lammps │ │ ├── templates │ │ └── lammps.py │ ├── phonopy │ │ └── run.py │ ├── vasp │ │ └── vasp.py │ ├── queue_jobs.py ├── tests │ ├── testfiles │ │ ├── ai │ │ ├── analysis │ │ │ ├── darkmatter │ │ │ ├── defects │ │ │ ├── elastic │ │ │ ├── interface │ │ │ ├── magnetism │ │ │ ├── periodic │ │ │ ├── phonon │ │ │ ├── solar │ │ │ ├── stm │ │ │ ├── structure │ │ │ ├── thermodynamics │ │ │ ├── topological │ │ ├── core │ │ ├── db │ │ ├── io │ │ │ ├── boltztrap │ │ │ ├── calphad │ │ │ ├── lammps │ │ │ ├── pennylane │ │ │ ├── phonopy │ │ │ ├── qiskit │ │ │ ├── qe │ │ │ ├── tequila │ │ │ ├── vasp │ │ │ ├── wannier │ │ │ ├── wanniertools │ │ │ ├── wien2k │ │ ├── tasks │ │ │ ├── test_lammps.py │ │ │ └── test_vasp.py └── README.rst
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file jarvis-tools-2021.10.3.tar.gz.
File metadata
- Download URL: jarvis-tools-2021.10.3.tar.gz
- Upload date:
- Size: 878.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.1 importlib_metadata/4.0.1 pkginfo/1.7.0 requests/2.25.1 requests-toolbelt/0.9.1 tqdm/4.60.0 CPython/3.8.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
5bf6b54df811eeab0ed4fbc1ae90b06205d90639ef34215eeb3468237f2d2908
|
|
| MD5 |
a69eb7edd67b0a1dcde9bc7a00605cb4
|
|
| BLAKE2b-256 |
fcdcbe4a6ff3ca0b960cbe30f1395a02a18ba5c2f6e769aa4f3b3d0021169462
|
File details
Details for the file jarvis_tools-2021.10.3-py2.py3-none-any.whl.
File metadata
- Download URL: jarvis_tools-2021.10.3-py2.py3-none-any.whl
- Upload date:
- Size: 947.7 kB
- Tags: Python 2, Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/3.4.1 importlib_metadata/4.0.1 pkginfo/1.7.0 requests/2.25.1 requests-toolbelt/0.9.1 tqdm/4.60.0 CPython/3.8.8
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
8c5445e261daa18e98320824691cfce60978177854fef17c1f5c7c1c7aa851d9
|
|
| MD5 |
4c9db8121edc1656abd7b966978aa7d5
|
|
| BLAKE2b-256 |
a3fcb84daf32dfe031f7a9361ffe02e9faf3c3d4fa93e87424a1066d213b15ed
|











