jbioseqtools
Project description
JBioSeqTools - python library
JBioSeqTools is the Python library for gene sequence downloading, optimization, structure prediction, vector building, and visualization

Author: Jakub Kubiś
Polish Academy of Sciences
Description
Used databases:
Table of contents
- seq_tools - part of the library containing sequence optimization, structure prediction, and visualization
1.1. Import part of library
1.2. Loading metadata
1.3. Downloading reference sequences with gene name
1.4. Downloading sequences with accession numbers
1.5. Creating FASTA format *.FASTA
1.6. Loading FASTA format from file *.FASTA
1.7. Writing to FASTA format *.FASTA
1.8. Conducting Multiple Alignments Analysis (MUSCLE) form FASTA
1.9. Display alignment plot
1.10. Decoding alignment file
1.11. Writing to ALIGN format *.ALIGN
1.12. Extracting consensus parts of alignments
1.13. Passing sequence from an external source
1.14. Clearing junk characters from the sequence
1.15. Checking that the sequence is coding protein (CDS)
1.16. Checking that all bases in sequence are contained in the UPAC code
1.17. Reversing the DNA / RNA sequence: 5' --> 3' and 3' --> 5'
1.18. Complementing the DNA / RNA second strand sequence
1.19. Changing DNA to RNA sequence
1.20. Changing RNA to DNA sequence
1.21. Changing DNA or RNA sequence to amino acid / protein sequence
1.22. Prediction of RNA / DNA sequence secondary structure
1.23. Prediction of RNAi on the provided sequence
1.24. Correcting of RNAi_data for complementarity to the loop sequence
1.25. Correcting of RNAi_data for complementarity to the additional external sequence
1.26. Codon optimization
1.27. Checking susceptibility to restriction enzymes
1.28. Checking and removing susceptibility to restriction enzymes
1.29. Sequence comparison (Interval Fold Change Estimation) - vector_build - part of the library containing building plasmid vectors with optimization elements from seq_tools
2.1. Import part of library
2.2. Creating vector plasmid
2.2.1 Creating expression of the plasmid vector
2.2.2 Creating RNAi / RNAi + expression of the plasmid vector
2.2.3 Creating plasmid vector of in-vitro transcription of mRNA
2.2.4 Creating plasmid vector of in-vitro transcription of RNAi
2.3. Creating sequence for synthesis de novo
2.3.5 Creating sequence prediction of mRNA for de novo synthesis
2.3.6 Creating sequence prediction of RNAi for de novo synthesisi
2.4. Creating vector plasmid from FASTA - display existing or custom editing FASTA file
2.4.1 Loading fasta from the file
2.4.2 Converting the FASTA string to the data frame
2.4.3 Decoding FASTA information
2.4.4 Creating graph of the plasmid vector
2.4.5 Writing FASTA format of the plasmid vector
2.4.6 Converting FASTA format to GeneBank format
2.4.7 Writing GeneBank format of the plasmid vector
Installation
In command line write:
pip install JBioSeqTools
- During the first library loading additional requirements will be installed (BLAST, MUSCLE) and metadata downloaded.
Usage
1. seq_tools - part of the library containing sequence optimization, structure prediction, and visualization
1.1. Import part of library
from jbst import seq_tools as st
1.2. Loading metadata
metadata = st.load_metadata()
This function loads the metadata from library repository, which includes such elements like: 'codons', 'vectors', 'linkers', 'regulators', 'fluorescent_tag', 'backbone', 'promoters', 'restriction', 'polya_seq', 'selection_markers', 'rnai', 'capacity', 'utr5', 'utr3'.
These elements are necessary for algorithms, as well as, the database of regulatory genetic sequences for users to project vectors and other genetic therapeutics.
Args:
Some metadata is not necessary for algorithms of plasmid vector creation, such as:
linkers (bool) - linkers (sequences between coding sequences) and their description, Default: True
loops (bool) - loops (sequences for creating shRNA / siRNA) and their description, Default: True
regulators (bool) - regulators (regulatory sequence elements for expression enhancement) and their description, Default: True
fluorescent_tag (bool) - fluorescent_tag (sequences coding fluorescent proteins) and their description, Default: True
promoters (bool) - promoters (previously described promoter sequences for providing coding and non-coding sequence transcription) and their description, Default: True
polya (bool) - signal of polyadenylation sequences and their description, Default: True
marker (bool) - selection marker sequences for bacterial selection by resistance to antibiotics and their description, Default: True
utr5 (bool) - 5`UTR sequences and their description, Default: True
utr3 (bool) - 3`UTR sequences and their description, Default: True
If the user uses external sources for providing these sequences and names, it can be set to False and these metadata will not load and reduce RAM usage.
Returns:
dict: Dictionary with the crucial set of metadata
1.3. Downloading reference sequences with gene name
data_dict = st.get_sequences_gene(gene_name, species = 'human', max_results = 20)
This function gets sequences from NCBI database based on gene name & species
Args:
gene_name (str) - name of searching gene in the HGNC nomenclature
species (str) - specie for which the gene sequence is searching (human / mouse / rat / both* / both2* / multi* / other**). Default: 'human'
*both - gene sequences for Mus musculus and Homo sapiens
*both2 - gene sequences for Rattus norvegicus and Homo sapiens
*multi - gene sequences for Mus musculus, Rattus norvegicus, and Homo sapiens
**other - the user can provide any species in Latin language via binomial nomenclature eg. Bos taurus, Zea mays, Sus scrofa, Danio rerio, Oryza sativa ...
max_results (int) - number of maximal amount of results for the provided gene and species. Default: 20
input_dict (dict) - dictionary of metadata provided by the user
Returns:
dict: Dictionary including all sequence variants, their names, features, and IDs of provided gene
1.4. Downloading sequences with accession numbers
data_dict = st.get_sequences_accesion(accesion_list)
This function gets sequences from NCBI database based on accession numbers
Args:
accesion_list (list) - accession numbers or number of searching sequences inside list eg. ['X81403.1', 'KJ890665.1']; ['KJ890665.1']
Returns:
dict: Dictionary including all sequences from the provided query, their names, features, and IDs
1.5. Creating FASTA format *.FASTA
fasta_string = st.generate_fasta_string(data_dict)
This function trnasform dictionaries from get_sequences_accesion() or get_sequences_gene() into FASTA format.
Args:
data_dict (dict) - dictionaries from get_sequences_accesion() or get_sequences_gene()
Returns:
txt: FASTA format of input dictionary
1.6. Loading FASTA format from file *.FASTA
fasta_string = st.load_fasta(path)
This function finds and removes restriction places inside the sequence.
Args:
path (str) - path to the FASTA file *.FASTA
Returns:
str: Loaded FASTA file to the string object
1.7. Writing to FASTA format *.FASTA
st.write_fasta(fasta_string, path = None, name = 'fasta_file')
This function saves into FASTA *.fasta
Args:
fasta_string (str/FASTA) - sequences provided in FASTA format from generate_fasta_string() or loaded from external sources
path (str | None) - the path to save. If None save it to the current working directory. Default: None
name (str) - the name of the saving file. Default: 'fasta_file'
1.8. Conducting Multiple Alignments Analysis (MUSCLE) form FASTA
alignment_file = st.MuscleMultipleSequenceAlignment(fasta_string, output = None, gapopen = 10, gapextend = 0.5)
This function conducts alignments of sequences provided in FASTA format
Args:
fasta_string (str\FASTA) - sequences provided in FASTA format from generate_fasta_string() or loaded from external sources
gapopen (int | float) - description under the link provided below. Default: 10
gapextend (int | float) - description under the link provided below. Default: 0.5
output (str | None) - path to the TMP alignment file. If None then the current working directory and the TMP file are removed. Default: None
More information in the source code of the primary function authors:
- https://www.drive5.com/muscle/
Returns:
txt: FASTA format of input dictionary
1.9. Display alignment plot
alignment_plot = st.DisplayAlignment(alignment_file, color_scheme="Taylor", wrap_length=80, show_grid=True, show_consensus=True)
alignment_plot.savefig("alignment_plot.svg")
This function makes the graphical presentation of the alignments
Args:
alignment_file (Bio.Align.MultipleSeqAlignment class) - the output file from MuscleMultipleSequenceAlignment()
color_scheme (str) - color palette for plotting. Default: "Taylor"
wrap_length (int) - max number of nucleotides shown in a row on the graph. Default: 80
show_grid (bool) - show the grid on the graph. Default: True
show_consensus (bool) - highlight the consensus sequences. Default: True
More information in the source code of the primary function authors:
- https://pypi.org/project/pyMSAviz/
Returns:
graph: The graphical presentation of sequences alignments
Example of the alignment graph:
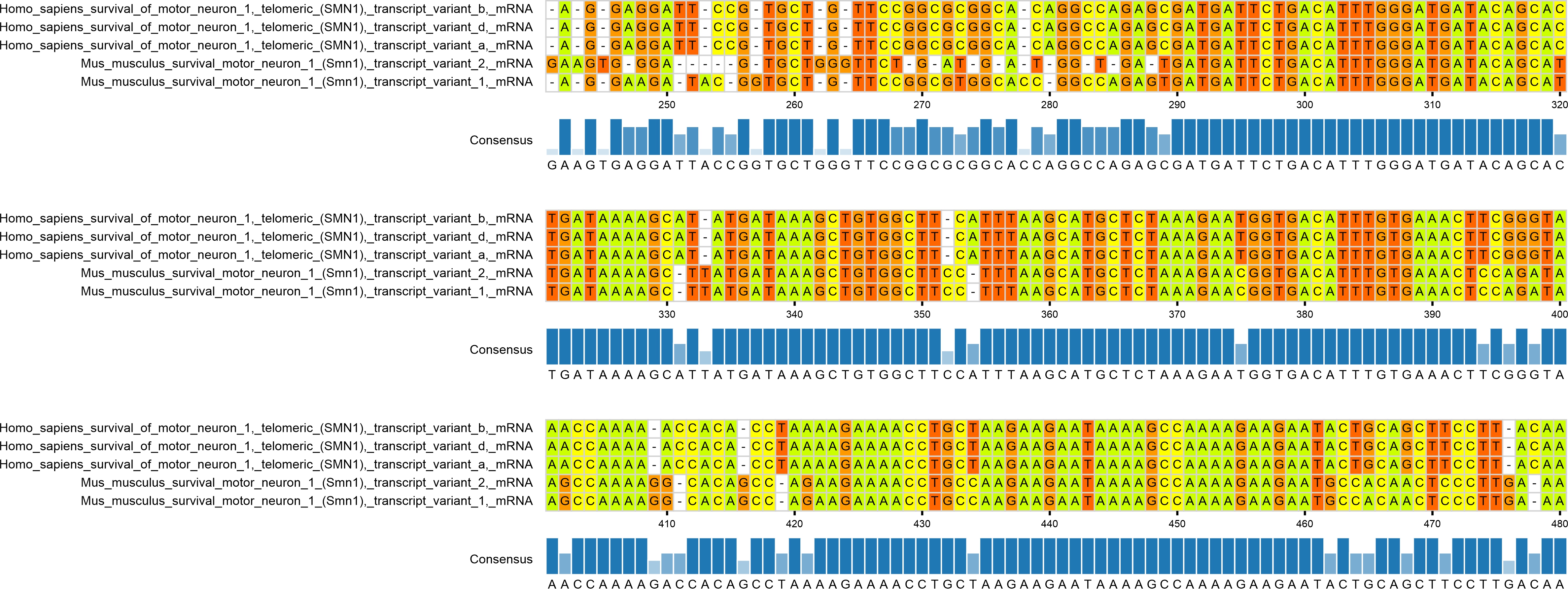
1.10. Decoding alignment file
decoded_alignment_file = st.decode_alignments(alignment_file)
This function decodes the alignment file from MuscleMultipleSequenceAlignment() and converts it to FASTA-ALIGNMENT
Args:
alignment_file (Bio.Align.MultipleSeqAlignment class) - the output file from MuscleMultipleSequenceAlignment()
Returns:
txt: FASTA-ALIGNMENT format of input file
Example output:
print(decoded_alignment_file)
Homo_sapiens_survival_of_motor_neuron_1,_telomeric_(SMN1),_transcript_variant_b,_mRNA | -CCACAAATGTGGGAGGGCGATAACCACTCGTAG-AAAGCGTGAGAAGTTACTACAAGC-G-G-TCCTCCC--G-GCCACCG-TACT-G-TT-C--C-----GCTCC-C-AGAAGCC--C--CGGGCGGCGG-AAGTCGTCACTCTTAAGAAGGGACGGGGCCCCACGCTG-CGCACCCGCGGGT-TTGCTATGGCGATGAGCAGCGG-C-GGCAGTGG-TGGC-G-G-CGTCCCGGAGC-A-G-GAGGATT-CCG-TGCT-G-TTCCGGCGCGGCA-CAGGCCAGAGCGATGATTCTGACATTTGGGATGATACAGCACTGATAAAAGCAT-ATGATAAAGCTGTGGCTT-CATTTAAGCATGCTCTAAAGAATGGTGACATTTGTGAAACTTCGGGTAAACCAAAA-ACCACA-CCTAAAAGAAAACCTGCTAAGAAGAATAAAAGCCAAAAGAAGAATACTGCAGCTTCCTT-ACAACAGTGGAAAGTTGGG-GACAAATGTTCTGCCATTTGGTCAGAAGACGGTTGCATTTACCCAGCTACCATTGC-TTCAATTGATTTTAAGAGAGAAACCTGTGTTGTGGTTTACACTGGATATGGAAATAGAGAGGAGCAAAATC-TGTCCGATCTACTTTCCCCAATCTGTGAAGTAGCTAATAATATAGAACA-AA-ATGCTCAAGAGAATGAAAATGAAAGCCAAGTTT-CAACAGATGAA-AGTGAGA-ACT-C-C-AGGTCTC-CT-GGAAATAAATCAGATA-ACATCAAGCCCAAATCTGCTCCATGGAA-CTC-TTTTCTCCCTCCACCACCCCCC-ATGCCAGGGCCAAGACTGGGACCAGGAAAG--A--T--A-A-TT------C-C--C--C-C-A-C-----C-A-C--C--T---C-C-C--A---T---A---T-G-T--C--C-A-----------G------A-T---T-----C-------T--C---T---T----GA-T--------GATG---CTGATGCTTTGGG-AAGTATGTTAATTTCAT-GGTACATGAGTGGCTATCATACTGGCTATTATATGGGTTTCAGACAAAATCAAAAAGAAGGAAGGTGCTCACATTCCTTAAATTAAGG-AGAAATGCTGGCATAGAGCAGCACTAAATGAC-ACCACTAAAGAAACGAT-CAGAC-A-GATCT--GGA-ATGTGAAGCGTTATAGAAGATAACTGGCCTCATTTCTTCAAAATATCAAGTGTTGGGA-AAGAAAAAAGGA-AGTGGAATGGGTAACTCTTCTTGATTAAAAGTTATGTAATAAC-CAAAT-G-C-AATGTGAAATATTTTACTGGACTCTATTTTGAAAAACCATCTGTAAAAGAC-TGGGG-TGGGGGTGGGAGGCCAGCACGGTGGTGAGGCAGTTGAGAAAATTTGAATGTGGATTAG-ATTTTG-AATGATATTG-GATAATTATTGGTAATTTTATGAGCTGTGAG-AAGGGTGTTGTAGTTTATAAAAGACTGTCTTAATTTGCATACTTAAGCATTTAGGAATGAAGTGTTAGAGTGTCTTA-AAATGTTTCAAATGGTTTAA--CAAAATGTATGTGAGGCGTATGTGGCA-AAATGTTACAGAATCTAACTGGTGGACATGGCTGTTCATT-GTACTGTTT----TTTTCTATCTTCTATATG--TTTAAAAGTATATAATAAAAATATTTAATTTTTTTTTAAAAAAAAAAAAAAAAA----AA
Homo_sapiens_survival_of_motor_neuron_1,_telomeric_(SMN1),_transcript_variant_d,_mRNA | ----------------G-------C----------A---------------C--C---C-G-----C------G-G-----G-T--T----T----------GCT-----A----------T--G--G-C-G--A-T-G----------------A----G--C-A-G----CG-----GCG-G-----C-A----G-TG-------G-T--G-----G----C-G-G-CGTCCCGGAGC-A-G-GAGGATT-CCG-TGCT-G-TTCCGGCGCGGCA-CAGGCCAGAGCGATGATTCTGACATTTGGGATGATACAGCACTGATAAAAGCAT-ATGATAAAGCTGTGGCTT-CATTTAAGCATGCTCTAAAGAATGGTGACATTTGTGAAACTTCGGGTAAACCAAAA-ACCACA-CCTAAAAGAAAACCTGCTAAGAAGAATAAAAGCCAAAAGAAGAATACTGCAGCTTCCTT-ACAACAGTGGAAAGTTGGG-GACAAATGTTCTGCCATTTGGTCAGAAGACGGTTGCATTTACCCAGCTACCATTGC-TTCAATTGATTTTAAGAGAGAAACCTGTGTTGTGGTTTACACTGGATATGGAAATAGAGAGGAGCAAAATC-TGTCCGATCTACTTTCCCCAATCTGTGAAGTAGCTAATAATATAGAACA-AA-ATGCTCAAGAGAATGAAAATGAAAGCCAAGTTT-CAACAGATGAA-AGTGAGA-ACT-C-C-AGGTCTC-CT-GGAAATAAATCAGATA-ACATCAAGCCCAAATCTGCTCCATGGAA-CTC-TTTTCTCCCTCCACCACCCCCC-ATGCCAGGGCCAAGACTGGGACCAGGAAAGCCAGGTCTAAAATTCAATGGCCCACCACCGCCA-C-CGC-C-ACCACCAC-CACCCCACTTACTAT-C-ATGCTGGCTG-CCTCCATTTC-CTTCTGGACCACCAATAATTCCC-CCA-CCACCTCCCA--TATGT-CCAGA-T-TCTCTT-GATGATGCTGATGCTTTGGG-AAGTATGTTAATTTCAT-GGTACATGAGTGGCTATCATACTGGCTATTATATGGGTTTCAGACAAAATCAAAAAGAAGGAAGGTGCTCACATTCCTTAAATTAAGG-AGAAATGCTGGCATAGAGCAGCACTAAATGAC-ACCACTAAAGAAACGAT-CAGAC-A-GATCT--GGA-ATGTGAAGCGTTATAGAAGATAACTGGCCTCATTTCTTCAAAATATCAAGTGTTGGGA-AAGAAAAAAGGA-AGTGGAATGGGTAACTCTTCTTGATTAAAAGTTATGTAATAAC-CAAAT-G-C-AATGTGAAATATTTTACTGGACTCTATTTTGAAAAACCATCTGTAAAAGAC-TGGGG-TGGGGGTGGGAGGCCAGCACGGTGGTGAGGCAGTTGAGAAAATTTGAATGTGGATTAG-ATTTTG-AATGATATTG-GATAATTATTGGTAATTTTATGAGCTGTGAG-AAGGGTGTTGTAGTTTATAAAAGACTGTCTTAATTTGCATACTTAAGCATTTAGGAATGAAGTGTTAGAGTGTCTTA-AAATGTTTCAAATGGTTTAA--CAAAATGTATGTGAGGCGTATGTGGCA-AAATGTTACAGAATCTAACTGGTGGACATGGCTGTTCATT-GTACTGTTT----TTTTCTATCTTCTATATG--TTTAAAAGTATATAATAAAAATATTTAATTTTTTTTT------------A-A-AT-T-A-
Homo_sapiens_survival_of_motor_neuron_1,_telomeric_(SMN1),_transcript_variant_a,_mRNA | -CCACAAATGTGGGAGGGCGATAACCACTCGTAG-AAAGCGTGAGAAGTTACTACAAGC-G-G-TCCTCCC--G-GCCACCG-TACT-G-TT-C--C-----GCTCC-C-AGAAGCC--C--CGGGCGGCGG-AAGTCGTCACTCTTAAGAAGGGACGGGGCCCCACGCTG-CGCACCCGCGGGT-TTGCTATGGCGATGAGCAGCGG-C-GGCAGTGG-TGGC-G-G-CGTCCCGGAGC-A-G-GAGGATT-CCG-TGCT-G-TTCCGGCGCGGCA-CAGGCCAGAGCGATGATTCTGACATTTGGGATGATACAGCACTGATAAAAGCAT-ATGATAAAGCTGTGGCTT-CATTTAAGCATGCTCTAAAGAATGGTGACATTTGTGAAACTTCGGGTAAACCAAAA-ACCACA-CCTAAAAGAAAACCTGCTAAGAAGAATAAAAGCCAAAAGAAGAATACTGCAGCTTCCTT-ACAACAGTGGAAAGTTGGG-GACAAATGTTCTGCCATTTGGTCAGAAGACGGTTGCATTTACCCAGCTACCATTGC-TTCAATTGATTTTAAGAGAGAAACCTGTGTTGTGGTTTACACTGGATATGGAAATAGAGAGGAGCAAAATC-TGTCCGATCTACTTTCCCCAATCTGTGAAGTAGCTAATAATATAGAACA-AA-ATGCTCAAGAGAATGAAAATGAAAGCCAAGTTT-CAACAGATGAA-AGTGAGA-ACT-C-C-AGGTCTC-CT-GGAAATAAATCAGATA-ACATCAAGCCCAAATCTGCTCCATGGAA-CTC-TTTTCTCCCTCCACCACCCCCC-ATGCCAGGGCCAAGACTGGGACCAGGAAAGCCAGGTCTAAAATTCAATGGCCCACCACCGCCA-C-CGC-C-ACCACCAC-CACCCCACTTACTAT-C-ATGCTGGCTG-CCTCCATTTC-CTTCTGGACCACCAATAATTCCC-CCA-CCACCTCCCA--TATGT-CCAGA-T-TCTCTT-GATGATGCTGATGCTTTGGG-AAGTATGTTAATTTCAT-GGTACATGAGTGGCTATCATACTGGCTATTATAT--G-----GA-A-A-T----------------G---------C------T--GGCAT--A-G--AGC--A--GCA-C--TAAATGAC-ACCACTAAAGAAACGAT-CAGAC-A-GATCT--GGA-ATGTGAAGCGTTATAGAAGATAACTGGCCTCATTTCTTCAAAATATCAAGTGTTGGGA-AAGAAAAAAGGA-AGTGGAATGGGTAACTCTTCTTGATTAAAAGTTATGTAATAAC-CAAAT-G-C-AATGTGAAATATTTTACTGGACTCTATTTTGAAAAACCATCTGTAAAAGAC-TGGGG-TGGGGGTGGGAGGCCAGCACGGTGGTGAGGCAGTTGAGAAAATTTGAATGTGGATTAG-ATTTTG-AATGATATTG-GATAATTATTGGTAATTTTATGAGCTGTGAG-AAGGGTGTTGTAGTTTATAAAAGACTGTCTTAATTTGCATACTTAAGCATTTAGGAATGAAGTGTTAGAGTGTCTTA-AAATGTTTCAAATGGTTTAA--CAAAATGTATGTGAGGCGTATGTGGCA-AAATGTTACAGAATCTAACTGGTGGACATGGCTGTTCATT-GTACTGTTT----TTTTCTATCTTCTATATG--TTTAAAAGTATATAATAAAAATATTTAATTTTTTTTT------------A-A-A------
Mus_musculus_survival_motor_neuron_1_(Smn1),_transcript_variant_2,_mRNA | G--AC---T-T-----G----TAACT-CTT-TAGCA--G--T--G---TT-CT-C---CTGTATTTCT-TTAAGTG--A--GTTATTAAATTCCTTCTTTATG-TCCTCTA----CCATCATC-A-TG-T-GAAA-T-G-C-TT-TT-A--A---AT-----CC-A-G--GTC-TA-C--C---TTTT-C-A--G-G-TG-----T-GTTAGG-A-T-GCC--CTG-GAC-T---GG-GCGAAGTG-GGA-----G-TGCTGGGTTCT-G-AT-G-A-T-GG-T-GA-TGATGATTCTGACATTTGGGATGATACAGCATTGATAAAAGC-TTATGATAAAGCTGTGGCTTCC-TTTAAGCATGCTCTAAAGAACGGTGACATTTGTGAAACTCCAGATAAGCCAAAAGG-CACAGCC-AGAAGAAAACCTGCCAAGAAGAATAAAAGCCAAAAGAAGAATGCCACAACTCCCTTGA-AACAGTGGAAAGTT-GGTGACAAGTGTTCTGCTGTTTGGTCAGAAGACGGCTGCATTTACCCAGCTACTATTACGTCC-ATTGACTTTAAGAGAGAAACCTGTGTCGTGGTTTATACTGGATATGGAAACAGAGAGGAGCAAAA-CTTATCTGACCTACTTTCCCCGACCTGTGAAGTAGCTAATAGTACAGAACAGAACA--CTC-AG-G-A-G-A-ATGAAAGTCAAGTTTCC-ACAGACG-ACAGTGA-ACACTCCTCCAGA--TCGCTCAGAAGT-AA--AG-CACACAGCAAGTCCAAAGCTGCTCCGTGG-ACCTCAT-TTCTTCCTCCACCA-CCCCCAATGCCAGGGTCAGGATTAGGACCAGGAAAGCCAGGTCTAAAATTCAACGGCCCGCCGCCGCCGCCTC-CACTACC-CC-CTC-CCCC-CTT-CC-TGCCGTGCTGGATGCCC-CCGTT-CCCTTCAGGACCACCAATAA-TCCCGCCACCC-CCTCCCATCT-C-TCCC-GACTGT--C-TGGATGACACTGATGCCCTGGGCA-GTATGCTAATCTC-TTGGTACATGAGTGGCTACCACACTGGCTACTATATGGGTTTCAGACAAAAT-A-A-A--A--AA---G--------------A--A-GG-A-A-A-G-T-GC-T----CA-C----A-T-ACAA--ATT-AAG-AA-G-TTCAG-CTCTG-TCTCAGGAGATG-G--G-G---T----G-T--C-GG--T-G-T-C--C----C-T---G-GTC-G-ACAAG-----A--ACA--G-A-C-G-T--CTC--CTCG-TC---A-TCA-GT-G-GACTC---TTGGCTAA-GTG-G-TG---T-C--G--TC-A-T-C--A-G-C-ATCT-C------CCC---GCT---G-TGGGA-GTC--CA---T--C----CA-TC-------C-T-AA-GT---C-AGCA----GCA--G--A--GCG-T-G-C-CTGG---------G-GC-GTGAGCA--G-T-T-G--G---A-G-G-GAC---C-------G-A--C-C-AG----T-GG-A-G---TG-T-GCGTGTC--GGAAG-G---C--A-G-TCT-ACCC---A-GT-CGTGA--C-T--G-AGCACAAATG-TGCA-A-T-T----G-T---CAT---T-TTC-TTAGCA-TG-TCAAGATTTT-TA---T-TA-ATGCCTTTAGAA-T-TA-AAT-AAAA-G-TC--C-T-TTTTT----------G-A-A-ATCTTG-
Mus_musculus_survival_motor_neuron_1_(Smn1),_transcript_variant_1,_mRNA | ----------------G----T--C-A-T--T-G-A--G--T--GA-G---C--C---C-G-G---C-----A--G---C-G---------T-C--C-----G-T-------------------G--G-T----A---G-C--------------A--G-G-CC-ATG--G-CG-A-T-G-G-G-----C-A----G-TG-------G-C-GG-A---G----C-GGG-C-TC-C-GAGC-A-G-GAAGA-TAC-GGTGCT-G-TTCCGGCGTGGCACC-GGCCAGAGTGATGATTCTGACATTTGGGATGATACAGCATTGATAAAAGC-TTATGATAAAGCTGTGGCTTCC-TTTAAGCATGCTCTAAAGAACGGTGACATTTGTGAAACTCCAGATAAGCCAAAAGG-CACAGCC-AGAAGAAAACCTGCCAAGAAGAATAAAAGCCAAAAGAAGAATGCCACAACTCCCTTGA-AACAGTGGAAAGTT-GGTGACAAGTGTTCTGCTGTTTGGTCAGAAGACGGCTGCATTTACCCAGCTACTATTACGTCC-ATTGACTTTAAGAGAGAAACCTGTGTCGTGGTTTATACTGGATATGGAAACAGAGAGGAGCAAAA-CTTATCTGACCTACTTTCCCCGACCTGTGAAGTAGCTAATAGTACAGAACAGAACA--CTC-AG-G-A-G-A-ATGAAAGTCAAGTTTCC-ACAGACG-ACAGTGA-ACACTCCTCCAGA--TCGCTCAGAAGT-AA--AG-CACACAGCAAGTCCAAAGCTGCTCCGTGG-ACCTCAT-TTCTTCCTCCACCA-CCCCCAATGCCAGGGTCAGGATTAGGACCAGGAAAGCCAGGTCTAAAATTCAACGGCCCGCCGCCGCCGCCTC-CACTACC-CC-CTC-CCCC-CTT-CC-TGCCGTGCTGGATGCCC-CCGTT-CCCTTCAGGACCACCAATAA-TCCCGCCACCC-CCTCCCATCT-C-TCCC-GACTGT--C-TGGATGACACTGATGCCCTGGGCA-GTATGCTAATCTC-TTGGTACATGAGTGGCTACCACACTGGCTACTATATGGGTTTCAGACAAAAT-A-A-A--A--AA---G--------------A--A-GG-A-A-A-G-T-GC-T----CA-C----A-T-ACAA--ATT-AAG-AA-G-TTCAG-CTCTG-TCTCAGGAGATG-G--G-G---T----G-T--C-GG--T-G-T-C--C----C-T---G-GTC-G-ACAAG-----A--ACA--G-A-C-G-T--CTC--CTCG-TC---A-TCA-GT-G-GACTC---TTGGCTAA-GTG-G-TG---T-C--G--TC-A-T-C--A-G-C-ATCT-C------CCC---GCT---G-TGGGA-GTC--CA---T--C----CA-TC-------C-T-AA-GT---C-AGCA----GCA--G--A--GCG-T-G-C-CTGG---------G-GC-GTGAGCA--G-T-T-G--G---A-G-G-GAC---C-------G-A--C-C-AG----T-GG-A-G---TG-T-GCGTGTC--GGAAG-G---C--A-G-TCT-ACCC---A-GT-CGTGA--C-T--G-AGCACAAATG-TGCA-A-T-T----G-T---CAT---T-TTC-TTAGCA-TG-TCAAGATTTT-TA---T-TA-ATGCCTTTAGAA-T-TA-AAT-AAAA-G-TC--C-T-TTTTT----------G-A-A-ATCTTG-
1.11. Writing to ALIGN format *.ALIGN
st.write_alignments(decoded_alignment_file, path = None, name = 'alignments_file')
This function saves into FASTA-ALIGNMENT *.align
Args:
decoded_alignment_file (str/FASTA) - sequences provided in FASTA format from decode_alignments()
path (str | None) - the path to save. If None save it to the current working directory. Default: None
name (str) - the name of the saving file. Default: 'alignments_file'
1.12. Extracting consensus parts of alignments
consensuse = st.ExtractConsensuse(alignment_file, refseq_sequences = None)
This function extracts consensus fragments of sequences alignments from alignment_file obtained in the MuscleMultipleSequence Alignment() function
Args:
alignment_file (Bio.Align.MultipleSeqAlignment class) - the output file from MuscleMultipleSequenceAlignment()
refseq_sequences (dict | None) - dictionary obtained from get_sequences_gene() or get_sequences_accesion(), which add additional information to results. If None results will be reduced to only consensus sequences.
Returns:
dict: The dictionary containing consensus fragments
1.13. Passing sequence from an external source
sequence = st.load_sequence()
This function makes it easy to load the genetic sequences to variable
Returns:
str: Input sequence in a variable
1.14. Clearing junk characters from the sequence
sequence = st.clear_sequence(sequence)
This function clear sequence from special characters, spaces, numbers that may be in the sequence when copied from external sources.
Args:
sequence (str) - nucleotide sequence provided in UPAC code
upac_code (list) - list of nucleotides in which the sequence is encoded. Default: ['A','C','T','G','N','M','R','W','S','Y','K','V','H','D','B','U', 'Ψ']
Returns:
str: The genetic sequence after clearing
1.15. Checking that the sequence is coding protein (CDS)
dec_coding = st.check_coding(sequence)
This function checks that the input sequence belongs to the (CDS) coding sequence. Checkpoints include the start with ATG and include 3-nucleotide repeats.
Args:
sequence (str) - nucleotide sequence provided in UPAC code
Returns:
bool: True/False
1.16. Checking that all bases in sequence are contained in the UPAC code
dec_upac = st.check_upac(sequence)
This function checks that the input sequence elements are included in UPAC code.
Args:
sequence (str) - nucleotide sequence provided in UPAC code
upac_code (list) - list of nucleotides in which the sequence is encoded. Default: ['A','C','T','G','N','M','R','W','S','Y','K','V','H','D','B','U', 'Ψ']
Returns:
bool: True/False
1.17. Reversing the DNA / RNA sequence: 5' --> 3' and 3' --> 5'
reversed_sequence = st.reverse(sequence)
This function reverses the input genetic sequence from 5' -> 3' to 3' -> 5' and opposite.
Args:
sequence (str) - nucleotide sequence provided in UPAC code
Returns:
str: The genetic sequence after reversing
1.18. Complementing the DNA / RNA second strand sequence
complementary_sequence = st.complement(sequence)
This function makes a complementary sequence to the input genetic sequence on the nucleotide pairs from the extended UPAC code.
Args:
sequence (str) - nucleotide sequence provided in UPAC code
Returns:
str: The complementary sequence
1.19. Changing DNA to RNA sequence
rna_sequence = st.dna_to_rna(sequence, enrichment= False)
This function changes the sequence from DNA format to RNA.
Args:
sequence (str) - nucleotide sequence provided in UPAC code
enrichment (bool) - If True, the nucleotides in the RNA sequence instead of uracil (U) will be replaced with pseudouridine (Ψ) (True/False). Default: False
Returns:
str: The RNA sequence
1.20. Changing RNA to DNA sequence
dna_sequence = st.rna_to_dna(rna_sequence)
This function changes the sequence from RNA format to DNA.
Args:
sequence (str) - nucleotide sequence provided in UPAC code
Returns:
str: The DNA sequence
1.21. Changing DNA or RNA sequence to amino acid / protein sequence
protein_sequence = st.seuqence_to_protein(dna_sequence, metadata)
This function changes the sequence from (CDS) RNA or DNA format to protein sequence.
Args:
sequence (str) - nucleotide sequence provided in UPAC code
metadata (dict) - set of metadata loaded vie load_metadata()
Returns:
str: The protein sequence
1.22. Prediction of RNA / DNA sequence secondary structure
figure, dot = predict_structure(sequence,
anty_sequence = '',
height=None,
width=None,
dis_alpha = 0.35,
seq_force = 27,
pair_force = 8,
show_plot = True)
figure.savefig('predicted_structure.svg')
Predict and visualize the secondary structure of RNA or DNA sequences.
The function uses the ViennaRNA package to fold a nucleotide sequence (or
a sequence with its complementary/antisense strand) and generates a
secondary structure in dot-bracket notation. The structure is visualized
as a network graph where nodes represent nucleotides and edges represent
base pairs and backbone connections.
Args:
sequence (str): Nucleotide sequence in IUPAC notation (ATGC/U).
anty_sequence (str, optional): Antisense/complementary sequence.
If provided, duplex folding is performed instead of single-strand folding.
Default is '' (single-strand folding).
height (int | float | None, optional): Height of the figure in inches.
If None, it is determined automatically based on sequence length.
width (int | float | None, optional): Width of the figure in inches.
If None, it is determined automatically based on sequence length.
dis_alpha (float, optional): Adjustment factor (0–1) controlling the
attraction of paired nucleotides in the visualization. Default is 0.35.
- Recommended for short sequences (~25 nt): 0.35 (default).
- Recommended for longer sequences (e.g. mRNA): ~0.1.
seq_force (int, optional): Edge weight for base-pair connections
in the layout algorithm. Default is 27.
pair_force (int, optional): Edge weight for backbone connections
in the layout algorithm. Default is 8.
- Recommended for short sequences (~25 nt): 8 (default).
- Recommended for longer sequences (e.g. mRNA): ~3.
show_plot (bool, optional): If True, the plot is displayed.
If False, the figure is returned without showing. Default is True.
Returns:
tuple:
- fig (matplotlib.figure.Figure): Generated Matplotlib figure object.
- dot_bracket_structure (str): Predicted secondary structure in
dot-bracket notation (e.g. "..((..))..").
Example of structure prediction graph:
- shRNA

- siRNA

- mRNA

1.23. Prediction of RNAi on the provided sequence
RNAi_data = st.FindRNAi(sequence, metadata, length = 23, n = 200, max_repeat_len = 3, max_off = 1, species = 'human', output = None, database_name = "refseq_select_rna", evalue = 1e-3, outfmt = 5, word_size = 7, max_hsps = 20, reward = 1, penalty = -3, gapopen = 5, gapextend = 2, dust = "no", extension = 'xml')
Example of data frame output:
| RNAi_name | RNAi_seq | target | e-value | bit_score | alignment_length | target_seq | target_gene_name | species | specificity | complemenatry_regions | complemenatry_pct | RNAi_sense | repeated_motif | repeated_motif_pct | score | GC% |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| RNAi_66 | TGATTTTGTCTGAAACCCATATA | ['Homo sapiens survival of motor neuron 1'] | [2.07974e-05] | [46.087] | [23] | ['TGATTTTGTCTGAAACCCATATA'] | ['SMN1'] | ['Homo sapiens'] | 1 | ['ATAT', 'TATA'] | 0.34782608695652173 | TATATGGGTTTCAGACAAAATCA | ['TTTT'] | 0.17 | 0.0 | 17.04 |
| RNAi_11 | TTGATTTTGTCTGAAACCCATAT | ['Homo sapiens survival of motor neuron 1'] | [2.07974e-05] | [46.087] | [23] | ['TTGATTTTGTCTGAAACCCATAT'] | ['SMN1'] | ['Homo sapiens'] | 1 | ['ATAT'] | 0.17391304347826086 | ATATGGGTTTCAGACAAAATCAA | ['TTTTTT'] | 0.26 | -3.0 | 17.04 |
Columns explanation:
-
RNAi_name - name added by algorithms in the course of searching and predicting RNAi's
-
RNAi_seq - the sequence of RNAi (reverse-complementary to a sequence that should be silenced)
-
target - name of the targeted sequence
-
e-value (expect value) - the probability of finding the observed alignment or a better one by random chance
-
bit_score - statistical measure that provides an indication of the significance of a sequence alignment
-
alignment_length - size of the alignment complementarity to the blasted sequence
-
target_seq - part of the blasted sequence complementary to the RNAi_seq (in the same rotation as RNAi)
-
target_gene_name - list of genes to which the RNAi is complementary
-
species - list of species to which the gene sequence is complementary
-
specificity - number of genes to which the RNAi is complementary
-
complemenatry_regions - complementary part of RNAi inside the RNAi sequence
-
complemenatry_pct - percent of complementary nucleotides inside RNAi
-
RNAi_sense - sense strand of RNAi (reverse-complement to the RNAi_seq)
-
repeated_motif - sequences with repeated sequences of the same nucleotide inside RNAi
-
repeated_motif_pct - percent of nucleotides from sequences with repeated sequences of the same nucleotide inside RNAi
-
score - algorithmically given points to particular RNAi sequences based on knowledge from previous works and papers:
-DOI:10.1093/nar/gkn902
-DOI:10.1093/nar/gkh247
-DOI:10.1038/nbt936
-DOI:10.1016/j.bbrc.2004.02.157 -
GC% - percent content of G and C nucleotides in RNAi sequence (best target between 35-55%)
1.24. Correcting of RNAi_data for complementarity to the loop sequence
RNAi_data = st.loop_complementary_adjustment(RNAi_data, loop_seq, min_length=3)
This function takes output DataFrame from Find RNAi() or remove_specific_to_sequence() reducing the RNAi score on their complementarity to the provided loop sequence.
It allows the choice of RNAi with better biological functionality.
Args:
RNAi_data (DataFrame) - data frame obtained in the FindRNAi() or remove_specific_to_sequence() function
loop_seq (str) - sequence of the loop in the UPAC code for siRNA / shRNA creation
min_length (int) - min value of loop complementary nucleotides. Default: 3
Returns:
DataFrame: Data frame containing predicted RNA sequence (sense / antisense), target, scores, and statistics corrected by reducing complementary to loop sequence.
Example of data frame output:
| RNAi_name | RNAi_seq | target | e-value | bit_score | alignment_length | target_seq | target_gene_name | species | specificity | complemenatry_regions | complemenatry_pct | RNAi_sense | repeated_motif | repeated_motif_pct | score | GC% | sense_loop_complementary | sense_loop_complementary_pct | antisense_loop_complementary | antisense_loop_complementary_pct |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| RNAi_66 | TGATTTTGTCTGAAACCCATATA | ['Homo sapiens survival of motor neuron 1'] | [2.07974e-05] | [46.087] | [23] | ['TGATTTTGTCTGAAACCCATATA'] | ['SMN1'] | ['Homo sapiens'] | 1 | ['ATAT', 'TATA'] | 0.34782608695652173 | TATATGGGTTTCAGACAAAATCA | ['TTTT'] | 0.17 | 0.0 | 17.04 | [] | 0.0 | [] | 0.0 |
| RNAi_11 | TTGATTTTGTCTGAAACCCATAT | ['Homo sapiens survival of motor neuron 1'] | [2.07974e-05] | [46.087] | [23] | ['TTGATTTTGTCTGAAACCCATAT'] | ['SMN1'] | ['Homo sapiens'] | 1 | ['ATAT'] | 0.17391304347826086 | ATATGGGTTTCAGACAAAATCAA | ['TTTTTT'] | 0.26 | -3.0 | 17.04 | [] | 0.0 | [] | 0.0 |
Columns explanation:
-
RNAi_name - name added by algorithms in the course of searching and predicting RNAi's
-
RNAi_seq - the sequence of RNAi (reverse-complementary to a sequence that should be silenced)
-
target - name of the targeted sequence
-
e-value (expect value) - the probability of finding the observed alignment or a better one by random chance
-
bit_score - statistical measure that provides an indication of the significance of a sequence alignment
-
alignment_length - size of the alignment complementarity to the blasted sequence
-
target_seq - part of the blasted sequence complementary to the RNAi_seq (in the same rotation as RNAi)
-
target_gene_name - list of genes to which the RNAi is complementary
-
species - list of species to which the gene sequence is complementary
-
specificity - number of genes to which the RNAi is complementary
-
complemenatry_regions - complementary part of RNAi inside the RNAi sequence
-
complemenatry_pct - percent of complementary nucleotides inside RNAi
-
RNAi_sense - sense strand of RNAi (reverse-complement to the RNAi_seq)
-
repeated_motif - sequences with repeated sequences of the same nucleotide inside RNAi
-
repeated_motif_pct - percent of nucleotides from sequences with repeated sequences of the same nucleotide inside RNAi
-
score - algorithmically given points to particular RNAi sequences based on knowledge from previous works and papers:
-DOI:10.1093/nar/gkn902
-DOI:10.1093/nar/gkh247
-DOI:10.1038/nbt936
-DOI:10.1016/j.bbrc.2004.02.157 -
GC% - percent content of G and C nucleotides in RNAi sequence (best target between 35-55%)
-
sense_loop_complementary - part of sense strand RNAi complementary to the loop
-
sense_loop_complementary_pct - percent of nucleotides from the sense strand of RNAi complementary to the loop
-
antisense_loop_complementary - part of antisense strand RNAi complementary to the loop
-
antisense_loop_complementary_pct - percent of nucleotides from the antisense strand of RNAi complementary to the loop
1.25. Correcting of RNAi_data for complementarity to the additional external sequence
RNAi_data = st.remove_specific_to_sequence(RNAi_data, sequence, min_length=4)
This function takes output DataFrame from Find RNAi() or loop_complementary_adjustment() reducing the RNAi score on their complementarity to the provided external genetic sequence. eg sequence after codon optimization which is not included in NCBI ref_seq db.
It allows the choice of RNAi with better biological functionality.
Args:
RNAi_data (DataFrame) - data frame obtained in the FindRNAi() or loop_complementary_adjustment() function
sequences (list | str) - nucleotide sequence provided in UPAC code
min_length (int) - min value of sequence complementary nucleotides. Default: 4
Returns:
DataFrame: Data frame containing predicted RNA sequence (sense / antisense), target, scores, and statistics corrected by reducing complementary to another sequence.
Example of data frame output:
| RNAi_name | RNAi_seq | target | e-value | bit_score | alignment_length | target_seq | target_gene_name | species | specificity | complemenatry_regions | complemenatry_pct | RNAi_sense | repeated_motif | repeated_motif_pct | score | GC% | sense_loop_complementary | sense_loop_complementary_pct | antisense_loop_complementary | antisense_loop_complementary_pct |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| RNAi_183 | TTTGATTTTGTCTGAAACCCATA | ['Homo sapiens survival of motor neuron 1'] | [2.07974e-05] | [46.087] | [23] | ['TTTGATTTTGTCTGAAACCCATA'] | ['SMN1'] | ['Homo sapiens'] | 1 | [] | 0.0 | TATGGGTTTCAGACAAAATCAAA | ['TTTTTTT'] | 0.3 | 0 | 17.04 | [] | 0.0 | [] | 0.0 |
| RNAi_164 | TTTTGATTTTGTCTGAAACCCAT | ['Homo sapiens survival of motor neuron 1'] | [2.07974e-05] | [46.087] | [23] | ['TTTTGATTTTGTCTGAAACCCAT'] | ['SMN1'] | ['Homo sapiens'] | 1 | [] | 0.0 | ATGGGTTTCAGACAAAATCAAAA | ['TTTTTTTT'] | 0.35 | 1.0 | 17.04 | [] | 0.0 | [] | 0.0 |
- Data reduced by RNAi records with too high complementarity of the provided sequence
- Columns explanation as in the paragraphs 1.23 and / or 1.24
1.26. Codon optimization
optimized_data = st.codon_otymization(sequence, metadata, species = 'human')
This function optimize procided genetic sequence.
Args:
sequence (str) - nucleotide sequence provided in UPAC (ATGC)
metadata (dict) - set of metadata loaded vie load_metadata()
species (str) - species for which the codons are optimized in the sequence (human / mouse / rat). Default: 'human'
GC_pct (int) - desired GC content percentage of the optimized sequence. Default is 58.
correct_rep (int) - codon repetition threshold for optimization (eg. CCCCCCC, AAAAAAA). Default is 7.
Returns:
DataFrame: Data frame containing the optimized sequence / input sequence and their statistics
1.27. Checking susceptibility to restriction enzymes
all_restriction_places, reduced_restriction_places_with_indexes = st.check_restriction(sequence, metadata)
This function finds restriction places inside the sequence.
Args:
sequence (str) - nucleotide sequence provided in UPAC (ATGC)
metadata (dict) - set of metadata loaded vie load_metadata()
Returns:
DataFrame: Data frame containing the restriction places and position of its occurrence in the provided genetic sequence
DataFrame: Data frame containing the parsed information about restriction places and their indexes to mapping to the first DataFrame
1.28. Checking and removing susceptibility to restriction enzymes
repaired_sequence_data = st.sequence_restriction_removal(sequence, metadata, restriction_places = [], species = 'human')
This function finds and removes restriction places inside the sequence.
Args:
sequence (str) - nucleotide sequence provided in UPAC (ATGC)
metadata (dict) - set of metadata loaded vie load_metadata()
restriction_places (list) - list of potential restriction places defined by the user to remove from the sequence. Default: []
*if the user did not define (empty list []) the potential restriction places to remove, the algorithm checks all possible restriction places, present it to the user (print), and asks him to choose which should be removed by writing IDs in consol.
species (str) - species for which the codons are exchanged to remove restriction places (human / mouse / rat). Default: 'human'
Returns:
DataFrame: Data frame containing the sequence before restriction removal and sequence after restriction removal and their statistics
1.29. Sequence comparison (Interval Fold Change Estimation)
comparison = compare_sequences(sequence_1,
sequence_name_1,
sequence_2,
sequence_name_2,
sep = 1)
Compares two character sequences and identifies differences at specified intervals.
This function compares two input sequences (e.g., DNA, RNA, amino acids) character by character
or in chunks of length `sep`. It returns a formatted text report showing the number of changes, the
percentage of positions that differ, and a list of specific changes with their positions.
Args:
sequence_1 (str): The first sequence to compare.
sequence_name_1 (str): A label or name for the first sequence (for display in the report).
sequence_2 (str): The second sequence to compare.
sequence_name_2 (str): A label or name for the second sequence (for display in the report).
sep (int, optional): Comparison step size; the number of characters to compare at a time. Default is 1.
Returns:
str: A formatted textual report that includes:
- Names of the compared sequences,
- Number and percentage of positions with changes,
- A table listing the positions and the specific changes in the format: `original -> new`.
2. vector_build - part of the library containing building plasmid vectors with optimization elements from seq_tools
2.1. Import part of library
from jbst import vector_build as vb
2.2. Creating vector plasmid
project = vb.vector_create_on_dict(metadata, input_dict, show_plot=True)
This function change provided by user metadata into three types of vector plasmids:
-expression (artificial gene expression)
-RNAi (silencing)
-invitro transcription (used in mRNA vaccine, mRNA, RNAi, and peptides production)
in three types of delivery systems:
-AAVs
-lentiviruses
-regular plasmid (for liposomes or other delivery systems)
Args:
metadata (dict) - matadata loaded in the load_metadata() function
input_dict (dict) - dictionary of metadata provided by the user
Examples:
-expression vector
-RNAi vector
-in-vitro transcription:
-RNAi
-mRNA
Avaiable on https://github.com/jkubis96/JBioSeqTools
If you have any problem, don't hesitate to contact us!
Args
show_plot (bool) - if True the plot will be displayed, if False only the graph will be returned to the project. Default: True
Returns:
dict: Dictionary including all vector data (graphs, sequences, fasta) created based on user definition
2.2.1 Creating expression of the plasmid vector
Empty input dictionary schema:
input_dict = {
# REQUIRED!
# name of current project (defined by user)
'project_name':''
# REQUIRED!
# avaiable of vector types (ssAAV / scAAV / lentiviral / regular)
'vector_type':'',
# REQUIRED!
# in this case 'vector_function':'expression'
'vector_function':'expression',
# REQUIRED!
# avaiable options (human / mouse / rat / both (mouse + human) / both2 (rat + human) / multi (mouse + rat + human))
# 'both / both2 / multi' - creating vector function adjusted for all species taking into consideration most adjustments for Homo sapiens
'species':'human',
# list of coding sequences (CDS) provided to make expression from the vector
# the CSD sequences the user can obtain from ...
# amount of sequences is not restricted as the user must remember that the length of whole vector is limited
# excide the relevant vector size can decrease vector working
# if the user wants to not include any sequences only fluorescent_tag, provide ['']
# sequences orientation 5' ---> 3' - sense
'sequences':[''],
# list of names of coding sequences
# amount of names should be equal with amount of sequences
# if provided no sequences, provide ['']
'sequences_names':[''],
# REQUIRED!
# sequence of provided promoter
# name and sequence the user can take from metadata['promoters'] (load_metadata())
# for coding sequences the user should choose the promoter of coding genes (metadata['promoters']['type'] == 'coding')
# sequence orientation 5' ---> 3' - sense
'promoter_sequence':'',
# REQUIRED!
# name of provided promoter sequence
'promoter_name':'',
# OPTIONAL!
# sequence of provided enhancer
# name and sequence the user can take from metadata['regulators'] (load_metadata())
# sequence orientation 5' ---> 3' - sense
'regulator_sequence':'',
# OPTIONAL!
# name of provided enhancer sequence
'regulator_name':'',
# REQUIRED!
# sequence of provided polyA signal
# name and sequence the user can take from metadata['polya_seq'] (load_metadata())
'polya_sequence':'',
# REQUIRED!
# name of provided polyA signal sequence
'polya_name':'',
# REQUIRED if more than one sequence of transcripts!
# sequences of provided linkers
# number of linkers_sequences should be equal number of sequences (transcripts) - 1. One linker for each pair of sequences.
# name and sequence the user can take from metadata['linkers'] (load_metadata())
# if the number of transcript sequences is equal 1 then provide empty list []
# if the user wants to not provide any linkers between the transcript sequences, provide an empty string '' for each pair of transcripts where the user wants to avoid linker; empty strings '' provide inside the list ['']
# sequence orientation 5' ---> 3' - sense
'linkers_sequences':[''],
# REQUIRED if more than one sequence!
# names of provided linkers
# if the number of transcript sequences is equal 1 then provide empty list []
# if the user wants to not provide any linkers between the transcript sequences, provide an empty string '' for each pair of transcripts where the user wants to avoid linker; empty strings '' provide inside the list ['']
'linkers_names':[''],
# OPTIONAL!
# sequence of provided fluorescent tag
# name and sequence the user can take from metadata['fluorescent_tag'] (load_metadata())
# if the user does not need fluorescent tag, provide ''
# sequence orientation 5' ---> 3' - sense
'fluorescence_sequence':'',
# OPTIONAL!
# name of provided fluorescent tag
# if the user does not need fluorescent tag, provide ''
'fluorescence_name':'',
# WARNING! If the user wants to use an additional promoter for the fluorescent tag expression, provide data for fluorescence_promoter_sequence & fluorescence_polya_sequence!
# OPTIONAL!
# sequence of provided fluorescence promoter
# name and sequence the user can take from metadata['promoters'] (load_metadata())
# if the user does not need additional promoter for fluorescent tag, provide ''
# sequence orientation 5' ---> 3' - sense
'fluorescence_promoter_sequence':'',
# OPTIONAL!
# name of provided fluorescence promoter
# if the user does not need additional promoter for fluorescent tag, provide ''
'fluorescence_promoter_name':'',
# OPTIONAL!
# sequence of provided fluorescence polyA signal
# name and sequence the user can take from metadata['polya_seq'] (load_metadata())
# if the user does not need additional promoter for fluorescent tag, provide ''
# sequence orientation 5' ---> 3' - sense
'fluorescence_polya_sequence':'',
# OPTIONAL!
# name of provided fluorescence polyA signal
# if the user does not need additional promoter for fluorescent tag, provide ''
'fluorescence_polya_name':'',
# WARNING! If provided sequences for transcripts (> 0) and do not need additional promoter for fluorescent tag, provide fluorescence_linker_sequence or provide empty string ''.
# OPTIONAL!
# sequence of provided fluorescence tag linker
# name and sequence the user can take from metadata['linkers'] (load_metadata())
# if the user has provided additional promoter, so the fluorescence_linker_sequence is not needed, provide ''
# sequence orientation 5' ---> 3' - sense
'fluorescence_linker_sequence':'',
# OPTIONAL!
# name of provided fluorescence tag linker
# if the user has provided additional promoter, so the fluorescence_linker_sequence is not needed, provide ''
'fluorescence_linker_name':'',
# REQUIRED!
# sequence of provided selection marker
# name and sequence the user can take from metadata['selection_markers'] (load_metadata())
# sequence orientation 5' ---> 3' - sense
'selection_marker_sequence':'',
# REQUIRED!
# name of provided selection marker
'selection_marker_name':'',
# OPTIONAL!
# restriction enzymes protection of transcript sequences
# enzymes the user can take from metadata['restriction'] (load_metadata())
# if do not need any restriction places protection, provide an empty list []
'restriction_list':[],
# REQUIRED!
# available options (True / False)
# decision; if the user wants the transcription sequences optimized based on the provided species
'optimize':True,
# REQUIRED; if optimize == True!
# user-defined percent of GC% content in predicted/optimized sequence
'transcript_GC':58,
# REQUIRED; if optimize == True!
# user-defined maximum number of consecutive repeats of a single nucleotide (A, C, T(U), G) in the predicted/optimized sequence, eg. AAAAAA
'poly_len':7
}
Example dictionary:
input_dict = {
'project_name':'test_expression',
'vector_type':'ssAAV',
'vector_function':'expression',
'species':'human',
'sequences':['ATGGCGATGAGCAGCGGCGGCAGTGGTGGCGGCGTCCCGGAGCAGGAGGATTCCGTGCTGTTCCGGCGCGGCACAGGCCAGAGCGATGATTCTGACATTTGGGATGATACAGCACTGATAAAAGCATATGATAAAGCTGTGGCTTCATTTAAGCATGCTCTAAAGAATGGTGACATTTGTGAAACTTCGGGTAAACCAAAAACCACACCTAAAAGAAAACCTGCTAAGAAGAATAAAAGCCAAAAGAAGAATACTGCAGCTTCCTTACAACAGTGGAAAGTTGGGGACAAATGTTCTGCCATTTGGTCAGAAGACGGTTGCATTTACCCAGCTACCATTGCTTCAATTGATTTTAAGAGAGAAACCTGTGTTGTGGTTTACACTGGATATGGAAATAGAGAGGAGCAAAATCTGTCCGATCTACTTTCCCCAATCTGTGAAGTAGCTAATAATATAGAACAAAATGCTCAAGAGAATGAAAATGAAAGCCAAGTTTCAACAGATGAAAGTGAGAACTCCAGGTCTCCTGGAAATAAATCAGATAACATCAAGCCCAAATCTGCTCCATGGAACTCTTTTCTCCCTCCACCACCCCCCATGCCAGGGCCAAGACTGGGACCAGGAAAGCCAGGTCTAAAATTCAATGGCCCACCACCGCCACCGCCACCACCACCACCCCACTTACTATCATGCTGGCTGCCTCCATTTCCTTCTGGACCACCAATAATTCCCCCACCACCTCCCATATGTCCAGATTCTCTTGATGATGCTGATGCTTTGGGAAGTATGTTAATTTCATGGTACATGAGTGGCTATCATACTGGCTATTATATGTTTCCTGAGGCCTCCCTAAAAGCCGAGCAGATGCCAGCACCATGCTTCCTGTAA',
'ATGGCGATGAGCAGCGGCGGCAGTGGTGGCGGCGTCCCGGAGCAGGAGGATTCCGTGCTGTTCCGGCGCGGCACAGGCCAGAGCGATGATTCTGACATTTGGGATGATACAGCACTGATAAAAGCATATGATAAAGCTGTGGCTTCATTTAAGCATGCTCTAAAGAATGGTGACATTTGTGAAACTTCGGGTAAACCAAAAACCACACCTAAAAGAAAACCTGCTAAGAAGAATAAAAGCCAAAAGAAGAATACTGCAGCTTCCTTACAACAGTGGAAAGTTGGGGACAAATGTTCTGCCATTTGGTCAGAAGACGGTTGCATTTACCCAGCTACCATTGCTTCAATTGATTTTAAGAGAGAAACCTGTGTTGTGGTTTACACTGGATATGGAAATAGAGAGGAGCAAAATCTGTCCGATCTACTTTCCCCAATCTGTGAAGTAGCTAATAATATAGAACAAAATGCTCAAGAGAATGAAAATGAAAGCCAAGTTTCAACAGATGAAAGTGAGAACTCCAGGTCTCCTGGAAATAAATCAGATAACATCAAGCCCAAATCTGCTCCATGGAACTCTTTTCTCCCTCCACCACCCCCCATGCCAGGGCCAAGACTGGGACCAGGAAAGCCAGGTCTAAAATTCAATGGCCCACCACCGCCACCGCCACCACCACCACCCCACTTACTATCATGCTGGCTGCCTCCATTTCCTTCTGGACCACCAATAATTCCCCCACCACCTCCCATATGTCCAGATTCTCTTGATGATGCTGATGCTTTGGGAAGTATGTTAATTTCATGGTACATGAGTGGCTATCATACTGGCTATTATATGTTTCCTGAGGCCTCCCTAAAAGCCGAGCAGATGCCAGCACCATGCTTCCTGTAA'],
'sequences_names':['SMN1','SMN2'],
'promoter_sequence':'GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGAT',
'promoter_name':'TBG',
'regulator_sequence':'CGATAATCAACCTCTGGATTACAAAATTTGTGAAAGATTGACTGGTATTCTTAACTATGTTGCTCCTTTTACGCTATGTGGATACGCTGCTTTAATGCCTTTGTATCATGCTATTGCTTCCCGTATGGCTTTCATTTTCTCCTCCTTGTATAAATCCTGGTTGCTGTCTCTTTATGAGGAGTTGTGGCCCGTTGTCAGGCAACGTGGCGTGGTGTGCACTGTGTTTGCTGACGCAACCCCCACTGGTTGGGGCATTGCCACCACCTGTCAGCTCCTTTCCGGGACTTTCGCTTTCCCCCTCCCTATTGCCACGGCGGAACTCATCGCCGCCTGCCTTGCCCGCTGCTGGACAGGGGCTCGGCTGTTGGGCACTGACAATTCCGTGGTGTTGTCGGGGAAGCTGACGTCCTTTCCATGGCTGCTCGCCTGTGTTGCCACCTGGATTCTGCGCGGGACGTCCTTCTGCTACGTCCCTTCGGCCCTCAATCCAGCGGACCTTCCTTCCCGCGGCCTGCTGCCGGCTCTGCGGCCTCTTCCGCGTCTTCGCCTTCGCCCTCAGACGAGTCGGATCTCCCTTTGGGCCGCCTCCCCGCATCGG',
'regulator_name':'WPRE',
'polya_sequence':'CAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTA',
'polya_name':'SV40_late',
'linkers_sequences':['GGAAGCGGAGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCCCGGCCCC'],
'linkers_names':['T2A'],
'fluorescence_sequence':'ATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGGGCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCATCTGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGACCTACGGCGTGCAGTGCTTCAGCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTCTTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGGACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCATGGCCGACAAGCAGAAGAACGGCATCAAGGTGAACTTCAAGATCCGCCACAACATCGAGGACGGCAGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCACTACCTGAGCACCCAGTCCGCCCTGAGCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCGTGACCGCCGCCGGGATCACTCTCGGCATGGACGAGCTGTACAAGTAA',
'fluorescence_name':'EGFP',
'fluorescence_promoter_sequence':'CTCGACATTGATTATTGACTAGTTATTAATAGTAATCAATTACGGGGTCATTAGTTCATAGCCCATATATGGAGTTCCGCGTTACATAACTTACGGTAAATGGCCCGCCTGGCTGACCGCCCAACGACCCCCGCCCATTGACGTCAATAATGACGTATGTTCCCATAGTAACGCCAATAGGGACTTTCCATTGACGTCAATGGGTGGAGTATTTACGGTAAACTGCCCACTTGGCAGTACATCAAGTGTATCATATGCCAAGTACGCCCCCTATTGACGTCAATGACGGTAAATGGCCCGCCTGGCATTATGCCCAGTACATGACCTTATGGGACTTTCCTACTTGGCAGTACATCTACGTATTAGTCATCGCTATTACCATGGTCGAGGTGAGCCCCACGTTCTGCTTCACTCTCCCCATCTCCCCCCCCTCCCCACCCCCAATTTTGTATTTATTTATTTTTTAATTATTTTGTGCAGCGATGGGGGCGGGGGGGGGGGGGGGGCGCGCGCCAGGCGGGGCGGGGCGGGGCGAGGGGCGGGGCGGGGCGAGGCGGAGAGGTGCGGCGGCAGCCAATCAGAGCGGCGCGCTCCGAAAGTTTCCTTTTATGGCGAGGCGGCGGCGGCGGCGGCCCTATAAAAAGCGAAGCGCGCGGCGGGCGGGAGTCGCTGCGCGCTGCCTTCGCCCCGTGCCCCGCTCCGCCGCCGCCTCGCGCCGCCCGCCCCGGCTCTGACTGACCGCGTTACTCCCACAGGTGAGCGGGCGGGACGGCCCTTCTCCTCCGGGCTGTAATTAGCGCTTGGTTTAATGACGGCTTGTTTCTTTTCTGTGGCTGCGTGAAAGCCTTGAGGGGCTCCGGGAGGGCCCTTTGTGCGGGGGGAGCGGCTCGGGGGGTGCGTGCGTGTGTGTGTGCGTGGGGAGCGCCGCGTGCGGCTCCGCGCTGCCCGGCGGCTGTGAGCGCTGCGGGCGCGGCGCGGGGCTTTGTGCGCTCCGCAGTGTGCGCGAGGGGAGCGCGGCCGGGGGCGGTGCCCCGCGGTGCGGGGGGGGCTGCGAGGGGAACAAAGGCTGCGTGCGGGGTGTGTGCGTGGGGGGGTGAGCAGGGGGTGTGGGCGCGTCGGTCGGGCTGCAACCCCCCCTGCACCCCCCTCCCCGAGTTGCTGAGCACGGCCCGGCTTCGGGTGCGGGGCTCCGTACGGGGCGTGGCGCGGGGCTCGCCGTGCCGGGCGGGGGGTGGCGGCAGGTGGGGGTGCCGGGCGGGGCGGGGCCGCCTCGGGCCGGGGAGGGCTCGGGGGAGGGGCGCGGCGGCCCCCGGAGCGCCGGCGGCTGTCGAGGCGCGGCGAGCCGCAGCCATTGCCTTTTATGGTAATCGTGCGAGAGGGCGCAGGGACTTCCTTTGTCCCAAATCTGTGCGGAGCCGAAATCTGGGAGGCGCCGCCGCACCCCCTCTAGCGGGCGCGGGGCGAAGCGGTGCGGCGCCGGCAGGAAGGAAATGGGCGGGGAGGGCCTTCGTGCGTCGCCGCGCCGCCGTCCCCTTCTCCCTCTCCAGCCTCGGGGCTGTCCGCGGGGGGACGGCTGCCTTCGGGGGGGACGGGGCAGGGCGGGGTTCGGCTTCTGGCGTGTGACCGGCGGCTCTAGAGCCTCTGCTAACCATGTTCATGCCTTCTTCTTTTTCCTACAGCTCCTGGGCAACGTGCTGGTTATTGTGCTGTCTCATCATTTTGGCAAAGAATTG',
'fluorescence_promoter_name':'CAG',
'fluorescence_polya_sequence':'CTGTGCCTTCTAGTTGCCAGCCATCTGTTGTTTGCCCCTCCCCCGTGCCTTCCTTGACCCTGGAAGGTGCCACTCCCACTGTCCTTTCCTAATAAAATGAGGAAATTGCATCGCATTGTCTGAGTAGGTGTCATTCTATTCTGGGGGGTGGGGTGGGGCAGGACAGCAAGGGGGAGGATTGGGAAGAGAATAGCAGGCATGCTGGGGA',
'fluorescence_polya_name':'bGH',
'fluorescence_linker_sequence':'GGAAGCGGAGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCCCGGCCCC',
'fluorescence_linker_name':'T2A',
'selection_marker_sequence':'ATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATTTTGCCTTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGTTACATCGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTTTTAAAGTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTCTCAGAATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGTGCTGCCATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTTTTTTGCACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGAGCGTGACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCCCGGCAACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGTTTATTGCTGATAAATCTGGAGCCGGTGAGCGTGGAAGCCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTCCCGTATCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCCTCACTGATTAAGCATTGGTAA',
'selection_marker_name':'Ampicillin',
'restriction_list':['RsaI', 'MnlI', 'AciI', 'AluI', 'BmrI'],
'optimize':True,
'transcript_GC':58,
'poly_len':7
}
Output:
# Name of the project
project['project']
# Graph of the designed vector
vector_plot = project['vector']['graph']
vector_plot.savefig('expression_vector.svg')
Example return:

# Complete FASTA file of the designed vecotr
project['vector']['full_fasta']
Example return:
>test_expression_ssAAV_expression_8780nc
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTCAGTGAGCGAGCGAGCGCGCAGAGAGGGAGTGGCCAACTCCATCACTAGGGGTTCCTTCTAGACAACTTTGTATAGAAAAGTTGGGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGATCAAGTTTGTACAAAAAAGCAGGCTGCCACCATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTGGGAAGCGGAGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCCCGGCCCCATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTGTGACAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTACTCGACATTGATTATTGACTAGTTATTAATAGTAATCAATTACGGGGTCATTAGTTCATAGCCCATATATGGAGTTCCGCGTTACATAACTTACGGTAAATGGCCCGCCTGGCTGACCGCCCAACGACCCCCGCCCATTGACGTCAATAATGACGTATGTTCCCATAGTAACGCCAATAGGGACTTTCCATTGACGTCAATGGGTGGAGTATTTACGGTAAACTGCCCACTTGGCAGTACATCAAGTGTATCATATGCCAAGTACGCCCCCTATTGACGTCAATGACGGTAAATGGCCCGCCTGGCATTATGCCCAGTACATGACCTTATGGGACTTTCCTACTTGGCAGTACATCTACGTATTAGTCATCGCTATTACCATGGTCGAGGTGAGCCCCACGTTCTGCTTCACTCTCCCCATCTCCCCCCCCTCCCCACCCCCAATTTTGTATTTATTTATTTTTTAATTATTTTGTGCAGCGATGGGGGCGGGGGGGGGGGGGGGGCGCGCGCCAGGCGGGGCGGGGCGGGGCGAGGGGCGGGGCGGGGCGAGGCGGAGAGGTGCGGCGGCAGCCAATCAGAGCGGCGCGCTCCGAAAGTTTCCTTTTATGGCGAGGCGGCGGCGGCGGCGGCCCTATAAAAAGCGAAGCGCGCGGCGGGCGGGAGTCGCTGCGCGCTGCCTTCGCCCCGTGCCCCGCTCCGCCGCCGCCTCGCGCCGCCCGCCCCGGCTCTGACTGACCGCGTTACTCCCACAGGTGAGCGGGCGGGACGGCCCTTCTCCTCCGGGCTGTAATTAGCGCTTGGTTTAATGACGGCTTGTTTCTTTTCTGTGGCTGCGTGAAAGCCTTGAGGGGCTCCGGGAGGGCCCTTTGTGCGGGGGGAGCGGCTCGGGGGGTGCGTGCGTGTGTGTGTGCGTGGGGAGCGCCGCGTGCGGCTCCGCGCTGCCCGGCGGCTGTGAGCGCTGCGGGCGCGGCGCGGGGCTTTGTGCGCTCCGCAGTGTGCGCGAGGGGAGCGCGGCCGGGGGCGGTGCCCCGCGGTGCGGGGGGGGCTGCGAGGGGAACAAAGGCTGCGTGCGGGGTGTGTGCGTGGGGGGGTGAGCAGGGGGTGTGGGCGCGTCGGTCGGGCTGCAACCCCCCCTGCACCCCCCTCCCCGAGTTGCTGAGCACGGCCCGGCTTCGGGTGCGGGGCTCCGTACGGGGCGTGGCGCGGGGCTCGCCGTGCCGGGCGGGGGGTGGCGGCAGGTGGGGGTGCCGGGCGGGGCGGGGCCGCCTCGGGCCGGGGAGGGCTCGGGGGAGGGGCGCGGCGGCCCCCGGAGCGCCGGCGGCTGTCGAGGCGCGGCGAGCCGCAGCCATTGCCTTTTATGGTAATCGTGCGAGAGGGCGCAGGGACTTCCTTTGTCCCAAATCTGTGCGGAGCCGAAATCTGGGAGGCGCCGCCGCACCCCCTCTAGCGGGCGCGGGGCGAAGCGGTGCGGCGCCGGCAGGAAGGAAATGGGCGGGGAGGGCCTTCGTGCGTCGCCGCGCCGCCGTCCCCTTCTCCCTCTCCAGCCTCGGGGCTGTCCGCGGGGGGACGGCTGCCTTCGGGGGGGACGGGGCAGGGCGGGGTTCGGCTTCTGGCGTGTGACCGGCGGCTCTAGAGCCTCTGCTAACCATGTTCATGCCTTCTTCTTTTTCCTACAGCTCCTGGGCAACGTGCTGGTTATTGTGCTGTCTCATCATTTTGGCAAAGAATTGATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGGGCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCATCTGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGACCTACGGCGTGCAGTGCTTCAGCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTCTTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGGACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCATGGCCGACAAGCAGAAGAACGGCATCAAGGTGAACTTCAAGATCCGCCACAACATCGAGGACGGCAGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCACTACCTGAGCACCCAGTCCGCCCTGAGCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCGTGACCGCCGCCGGGATCACTCTCGGCATGGACGAGCTGTACAAGTAAACCCAGCTTTCTTGTACAAAGTGGGAATTCCGATAATCAACCTCTGGATTACAAAATTTGTGAAAGATTGACTGGTATTCTTAACTATGTTGCTCCTTTTACGCTATGTGGATACGCTGCTTTAATGCCTTTGTATCATGCTATTGCTTCCCGTATGGCTTTCATTTTCTCCTCCTTGTATAAATCCTGGTTGCTGTCTCTTTATGAGGAGTTGTGGCCCGTTGTCAGGCAACGTGGCGTGGTGTGCACTGTGTTTGCTGACGCAACCCCCACTGGTTGGGGCATTGCCACCACCTGTCAGCTCCTTTCCGGGACTTTCGCTTTCCCCCTCCCTATTGCCACGGCGGAACTCATCGCCGCCTGCCTTGCCCGCTGCTGGACAGGGGCTCGGCTGTTGGGCACTGACAATTCCGTGGTGTTGTCGGGGAAGCTGACGTCCTTTCCATGGCTGCTCGCCTGTGTTGCCACCTGGATTCTGCGCGGGACGTCCTTCTGCTACGTCCCTTCGGCCCTCAATCCAGCGGACCTTCCTTCCCGCGGCCTGCTGCCGGCTCTGCGGCCTCTTCCGCGTCTTCGCCTTCGCCCTCAGACGAGTCGGATCTCCCTTTGGGCCGCCTCCCCGCATCGGGAATTCCTAGAGCTCGCTGATCAGCCTCGACTGTGCCTTCTAGTTGCCAGCCATCTGTTGTTTGCCCCTCCCCCGTGCCTTCCTTGACCCTGGAAGGTGCCACTCCCACTGTCCTTTCCTAATAAAATGAGGAAATTGCATCGCATTGTCTGAGTAGGTGTCATTCTATTCTGGGGGGTGGGGTGGGGCAGGACAGCAAGGGGGAGGATTGGGAAGAGAATAGCAGGCATGCTGGGGAGGGCCGCCTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTCAGTGAGCGAGCGAGCGCGCAGAGAGGGAGTGGCCAACTCCATCACTAGGGGTTCCTCTGCCTGCAGGGGCGCCTGATGCGGTATTTTCTCCTTACGCATCTGTGCGGTATTTCACACCGCATACGTCAAAGCAACCATAGTACGCGCCCTGTAGCGGCGCATTAAGCGCGGCGGGGGTGGTGGTTACGCGCAGCGTGACCGCTACACTTGCCAGCGCCTTAGCGCCCGCTCCTTTCGCTTTCTTCCCTTCCTTTCTCGCCACGTTCGCCGGCTTTCCCCGTCAAGCTCTAAATCGGGGGCTCCCTTTAGGGTTCCGATTTAGTGCTTTACGGCACCTCGACCCCAAAAAACTTGATTTGGGTGATGGTTCACGTAGTGGGCCATCGCCCTGATAGACGGTTTTTCGCCCTTTGACGTTGGAGTCCACGTTCTTTAATAGTGGACTCTTGTTCCAAACTGGAACAACACTCAACTCTATCTCGGGCTATTCTTTTGATTTATAAGGGATTTTGCCGATTTCGGTCTATTGGTTAAAAAATGAGCTGATTTAACAAAAATTTAACGCGAATTTTAACAAAATATTAACGTTTACAATTTTATGGTGCACTCTCAGTACAATCTGCTCTGATGCCGCATAGTTAAGCCAGCCCCGACACCCGCCAACACCCGCTGACGCGCCCTGACGGGCTTGTCTGCTCCCGGCATCCGCTTACAGACAAGCTGTGACCGTCTCCGGGAGCTGCATGTGTCAGAGGTTTTCACCGTCATCACCGAAACGCGCGAGACGAAAGGGCCTCGTGATACGCCTATTTTTATAGGTTAATGTCATGATAATAATGGTTTCTTAGACGTCAGGTGGCACTTTTCGGGGAAATGTGCGCGGAACCCCTATTTGTTTATTTTTCTAAATACATTCAAATATGTATCCGCTCATGAGACAATAACCCTGATAAATGCTTCAATAATATTGAAAAAGGAAGAGTATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATTTTGCCTTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGTTACATCGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTTTTAAAGTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTCTCAGAATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGTGCTGCCATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTTTTTTGCACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGAGCGTGACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCCCGGCAACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGTTTATTGCTGATAAATCTGGAGCCGGTGAGCGTGGAAGCCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTCCCGTATCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCCTCACTGATTAAGCATTGGTAACTGTCAGACCAAGTTTACTCATATATACTTTAGATTGATTTAAAACTTCATTTTTAATTTAAAAGGATCTAGGTGAAGATCCTTTTTGATAATCTCATGACCAAAATCCCTTAACGTGAGTTTTCGTTCCACTGAGCGTCAGACCCCGTAGAAAAGATCAAAGGATCTTCTTGAGATCCTTTTTTTCTGCGCGTAATCTGCTGCTTGCAAACAAAAAAACCACCGCTACCAGCGGTGGTTTGTTTGCCGGATCAAGAGCTACCAACTCTTTTTCCGAAGGTAACTGGCTTCAGCAGAGCGCAGATACCAAATACTGTTCTTCTAGTGTAGCCGTAGTTAGGCCACCACTTCAAGAACTCTGTAGCACCGCCTACATACCTCGCTCTGCTAATCCTGTTACCAGTGGCTGCTGCCAGTGGCGATAAGTCGTGTCTTACCGGGTTGGACTCAAGACGATAGTTACCGGATAAGGCGCAGCGGTCGGGCTGAACGGGGGGTTCGTGCACACAGCCCAGCTTGGAGCGAACGACCTACACCGAACTGAGATACCTACAGCGTGAGCTATGAGAAAGCGCCACGCTTCCCGAAGGGAGAAAGGCGGACAGGTATCCGGTAAGCGGCAGGGTCGGAACAGGAGAGCGCACGAGGGAGCTTCCAGGGGGAAACGCCTGGTATCTTTATAGTCCTGTCGGGTTTCGCCACCTCTGACTTGAGCGTCGATTTTTGTGATGCTCGTCAGGGGGGCGGAGCCTATGGAAAAACGCCAGCAACGCGGCCTTTTTACGGTTCCTGGCCTTTTGCTGGCCTTTTGCTCACATGTCCTGCAGGCAG
# The FASTA file is divided into particular elements of the designed vector
project['vector']['fasta']
Example return:
# test_expression_ssAAV_expression_8780nc
>5`ITR
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTCAGTGAGCGAGCGAGCGCGCAGAGAGGGAGTGGCCAACTCCATCACTAGGGGTTCCT
>backbone_element
TCTAGACAACTTTGTATAGAAAAGTTG
>Promoter : TBG
GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGAT
>backbone_element
CAAGTTTGTACAAAAAAGCAGGCT
>Kozak_sequence
GCCACC
>SEQ1 : SMN1
ATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTG
>Linker_1 : T2A
GGAAGCGGAGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCCCGGCCCC
>SEQ2 : SMN2
ATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTGTGA
>PolyA_signal : SV40_late
CAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTA
>2nd_promoter : CAG
CTCGACATTGATTATTGACTAGTTATTAATAGTAATCAATTACGGGGTCATTAGTTCATAGCCCATATATGGAGTTCCGCGTTACATAACTTACGGTAAATGGCCCGCCTGGCTGACCGCCCAACGACCCCCGCCCATTGACGTCAATAATGACGTATGTTCCCATAGTAACGCCAATAGGGACTTTCCATTGACGTCAATGGGTGGAGTATTTACGGTAAACTGCCCACTTGGCAGTACATCAAGTGTATCATATGCCAAGTACGCCCCCTATTGACGTCAATGACGGTAAATGGCCCGCCTGGCATTATGCCCAGTACATGACCTTATGGGACTTTCCTACTTGGCAGTACATCTACGTATTAGTCATCGCTATTACCATGGTCGAGGTGAGCCCCACGTTCTGCTTCACTCTCCCCATCTCCCCCCCCTCCCCACCCCCAATTTTGTATTTATTTATTTTTTAATTATTTTGTGCAGCGATGGGGGCGGGGGGGGGGGGGGGGCGCGCGCCAGGCGGGGCGGGGCGGGGCGAGGGGCGGGGCGGGGCGAGGCGGAGAGGTGCGGCGGCAGCCAATCAGAGCGGCGCGCTCCGAAAGTTTCCTTTTATGGCGAGGCGGCGGCGGCGGCGGCCCTATAAAAAGCGAAGCGCGCGGCGGGCGGGAGTCGCTGCGCGCTGCCTTCGCCCCGTGCCCCGCTCCGCCGCCGCCTCGCGCCGCCCGCCCCGGCTCTGACTGACCGCGTTACTCCCACAGGTGAGCGGGCGGGACGGCCCTTCTCCTCCGGGCTGTAATTAGCGCTTGGTTTAATGACGGCTTGTTTCTTTTCTGTGGCTGCGTGAAAGCCTTGAGGGGCTCCGGGAGGGCCCTTTGTGCGGGGGGAGCGGCTCGGGGGGTGCGTGCGTGTGTGTGTGCGTGGGGAGCGCCGCGTGCGGCTCCGCGCTGCCCGGCGGCTGTGAGCGCTGCGGGCGCGGCGCGGGGCTTTGTGCGCTCCGCAGTGTGCGCGAGGGGAGCGCGGCCGGGGGCGGTGCCCCGCGGTGCGGGGGGGGCTGCGAGGGGAACAAAGGCTGCGTGCGGGGTGTGTGCGTGGGGGGGTGAGCAGGGGGTGTGGGCGCGTCGGTCGGGCTGCAACCCCCCCTGCACCCCCCTCCCCGAGTTGCTGAGCACGGCCCGGCTTCGGGTGCGGGGCTCCGTACGGGGCGTGGCGCGGGGCTCGCCGTGCCGGGCGGGGGGTGGCGGCAGGTGGGGGTGCCGGGCGGGGCGGGGCCGCCTCGGGCCGGGGAGGGCTCGGGGGAGGGGCGCGGCGGCCCCCGGAGCGCCGGCGGCTGTCGAGGCGCGGCGAGCCGCAGCCATTGCCTTTTATGGTAATCGTGCGAGAGGGCGCAGGGACTTCCTTTGTCCCAAATCTGTGCGGAGCCGAAATCTGGGAGGCGCCGCCGCACCCCCTCTAGCGGGCGCGGGGCGAAGCGGTGCGGCGCCGGCAGGAAGGAAATGGGCGGGGAGGGCCTTCGTGCGTCGCCGCGCCGCCGTCCCCTTCTCCCTCTCCAGCCTCGGGGCTGTCCGCGGGGGGACGGCTGCCTTCGGGGGGGACGGGGCAGGGCGGGGTTCGGCTTCTGGCGTGTGACCGGCGGCTCTAGAGCCTCTGCTAACCATGTTCATGCCTTCTTCTTTTTCCTACAGCTCCTGGGCAACGTGCTGGTTATTGTGCTGTCTCATCATTTTGGCAAAGAATTG
>Fluorescent_tag : EGFP
ATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGGGCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCATCTGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGACCTACGGCGTGCAGTGCTTCAGCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTCTTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGGACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCATGGCCGACAAGCAGAAGAACGGCATCAAGGTGAACTTCAAGATCCGCCACAACATCGAGGACGGCAGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCACTACCTGAGCACCCAGTCCGCCCTGAGCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCGTGACCGCCGCCGGGATCACTCTCGGCATGGACGAGCTGTACAAGTAA
>backbone_element
ACCCAGCTTTCTTGTACAAAGTGGGAATTC
>Enhancer : WPRE
CGATAATCAACCTCTGGATTACAAAATTTGTGAAAGATTGACTGGTATTCTTAACTATGTTGCTCCTTTTACGCTATGTGGATACGCTGCTTTAATGCCTTTGTATCATGCTATTGCTTCCCGTATGGCTTTCATTTTCTCCTCCTTGTATAAATCCTGGTTGCTGTCTCTTTATGAGGAGTTGTGGCCCGTTGTCAGGCAACGTGGCGTGGTGTGCACTGTGTTTGCTGACGCAACCCCCACTGGTTGGGGCATTGCCACCACCTGTCAGCTCCTTTCCGGGACTTTCGCTTTCCCCCTCCCTATTGCCACGGCGGAACTCATCGCCGCCTGCCTTGCCCGCTGCTGGACAGGGGCTCGGCTGTTGGGCACTGACAATTCCGTGGTGTTGTCGGGGAAGCTGACGTCCTTTCCATGGCTGCTCGCCTGTGTTGCCACCTGGATTCTGCGCGGGACGTCCTTCTGCTACGTCCCTTCGGCCCTCAATCCAGCGGACCTTCCTTCCCGCGGCCTGCTGCCGGCTCTGCGGCCTCTTCCGCGTCTTCGCCTTCGCCCTCAGACGAGTCGGATCTCCCTTTGGGCCGCCTCCCCGCATCGG
>backbone_element
GAATTCCTAGAGCTCGCTGATCAGCCTCGA
>2nd_polyA_signal : bGH
CTGTGCCTTCTAGTTGCCAGCCATCTGTTGTTTGCCCCTCCCCCGTGCCTTCCTTGACCCTGGAAGGTGCCACTCCCACTGTCCTTTCCTAATAAAATGAGGAAATTGCATCGCATTGTCTGAGTAGGTGTCATTCTATTCTGGGGGGTGGGGTGGGGCAGGACAGCAAGGGGGAGGATTGGGAAGAGAATAGCAGGCATGCTGGGGA
>backbone_element
GGGCCGC
>3`ITR
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTCAGTGAGCGAGCGAGCGCGCAGAGAGGGAGTGGCCAACTCCATCACTAGGGGTTCCT
>backbone_element
CTGCCTGCAGGGGCGCCTGATGCGGTATTTTCTCCTTACGCATCTGTGCGGTATTTCACACCGCATACGTCAAAGCAACCATAGTACGCGCCCTGTAGCGGCGCATTAAGCGCGGCGGGGGTGGTGGTTACGCGCAGCGTGACCGCTACACTTGCCAGCGCCTTAGCGCCCGCTCCTTTCGCTTTCTTCCCTTCCTTTCTCGCCACGTTCGCCGGCTTTCCCCGTCAAGCTCTAAATCGGGGGCTCCCTTTAGGGTTCCGATTTAGTGCTTTACGGCACCTCGACCCCAAAAAACTTGATTTGGGTGATGGTTCACGTAGTGGGCCATCGCCCTGATAGACGGTTTTTCGCCCTTTGACGTTGGAGTCCACGTTCTTTAATAGTGGACTCTTGTTCCAAACTGGAACAACACTCAACTCTATCTCGGGCTATTCTTTTGATTTATAAGGGATTTTGCCGATTTCGGTCTATTGGTTAAAAAATGAGCTGATTTAACAAAAATTTAACGCGAATTTTAACAAAATATTAACGTTTACAATTTTATGGTGCACTCTCAGTACAATCTGCTCTGATGCCGCATAGTTAAGCCAGCCCCGACACCCGCCAACACCCGCTGACGCGCCCTGACGGGCTTGTCTGCTCCCGGCATCCGCTTACAGACAAGCTGTGACCGTCTCCGGGAGCTGCATGTGTCAGAGGTTTTCACCGTCATCACCGAAACGCGCGAGACGAAAGGGCCTCGTGATACGCCTATTTTTATAGGTTAATGTCATGATAATAATGGTTTCTTAGACGTCAGGTGGCACTTTTCGGGGAAATGTGCGCGGAACCCCTATTTGTTTATTTTTCTAAATACATTCAAATATGTATCCGCTCATGAGACAATAACCCTGATAAATGCTTCAATAATATTGAAAAAGGAAGAGT
>Resistance : Ampicillin
ATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATTTTGCCTTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGTTACATCGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTTTTAAAGTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTCTCAGAATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGTGCTGCCATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTTTTTTGCACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGAGCGTGACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCCCGGCAACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGTTTATTGCTGATAAATCTGGAGCCGGTGAGCGTGGAAGCCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTCCCGTATCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCCTCACTGATTAAGCATTGGTAA
>backbone_element
CTGTCAGACCAAGTTTACTCATATATACTTTAGATTGATTTAAAACTTCATTTTTAATTTAAAAGGATCTAGGTGAAGATCCTTTTTGATAATCTCATGACCAAAATCCCTTAACGTGAGTTTTCGTTCCACTGAGCGTCAGACCCCGTAGAAAAGATCAAAGGATCTTC
>pUC_ori
TTGAGATCCTTTTTTTCTGCGCGTAATCTGCTGCTTGCAAACAAAAAAACCACCGCTACCAGCGGTGGTTTGTTTGCCGGATCAAGAGCTACCAACTCTTTTTCCGAAGGTAACTGGCTTCAGCAGAGCGCAGATACCAAATACTGTTCTTCTAGTGTAGCCGTAGTTAGGCCACCACTTCAAGAACTCTGTAGCACCGCCTACATACCTCGCTCTGCTAATCCTGTTACCAGTGGCTGCTGCCAGTGGCGATAAGTCGTGTCTTACCGGGTTGGACTCAAGACGATAGTTACCGGATAAGGCGCAGCGGTCGGGCTGAACGGGGGGTTCGTGCACACAGCCCAGCTTGGAGCGAACGACCTACACCGAACTGAGATACCTACAGCGTGAGCTATGAGAAAGCGCCACGCTTCCCGAAGGGAGAAAGGCGGACAGGTATCCGGTAAGCGGCAGGGTCGGAACAGGAGAGCGCACGAGGGAGCTTCCAGGGGGAAACGCCTGGTATCTTTATAGTCCTGTCGGGTTTCGCCACCTCTGACTTGAGCGTCGATTTTTGTGATGCTCGTCAGGGGGGCGGAGCCTATGGAAA
>backbone_element
AACGCCAGCAACGCGGCCTTTTTACGGTTCCTGGCCTTTTGCTGGCCTTTTGCTCACATGTCCTGCAGGCAG
Sequences output data:
## genes names
project['transcripts']['sequences']['name']
## proteins sequences
project['transcripts']['sequences']['sequence_aa']
## average codon frequency in the input sequence
project['transcripts']['sequences']['vector_sequence_frequence']
## GC% content in the input sequence
project['transcripts']['sequences']['vector_sequence_GC']
############################################################################
## average codon frequency in the output sequence
project['transcripts']['sequences']['optimized_vector_sequence_frequence']
## GC% content in the output sequence
project['transcripts']['sequences']['optimized_vector_sequence_GC']
# Sequences
# Input sequence
project['transcripts']['sequences']['sequence']
# Predicted structure - input
project['transcripts']['sequences']['sequence_figure']
# Optimized sequence
project['transcripts']['sequences']['vector_sequence']
# Predicted structure - optimized
project['transcripts']['sequences']['optimized_sequence_figure']
Example return:
- Otimized sequence structure:

- Input sequence structure:

- Alternative options:
# Sequences results and metadata
# optimized sequence - main
project['transcripts']['sequences']
# optimized sequence - alternatives
project['transcripts']['alternative']
# additional variant 1
project['transcripts']['alternative']['var0]
# additional variant 2
project['transcripts']['alternative']['var1]
Example return:
>Optimized sequence - main
ATGCTGACCAGCGGATTGGTTGTGAGCAACATGTTCAGCTACCACCTGGCCGCCCTGGGATTGATGCCTTCTTTTCAGATGGAGGGCAGGGGCAGAGTGAACCAGTTGGGAGGAGTGTTTATCAACGGCCGGCCTCTGCCTAATCACATCCGGCTGAAAATCGTGGAGTTGGCCGCTCAGGGAGTTAGGCCTTGTGTGATTAGCAGGCAGCTGAGGGTGTCTCACGGATGTGTGAGCAAGATCCTGCAGCGGTACCAGGAGACCGGCTCTATCAGGCCTGGAGTGATTGGCGGCTCTAAACCCAGGGTGGCTACACCTGAAGTGGAAAAGAAGATCGAGCAGTACAAGAAGGACAACCCCGGCATCTTCAGCTGGGAGATCCGGGACCGGCTGCTGAAGGAGGGCATCTGCGACAGAAGCACCGTGCCTTCTGTGTCTAGCATCAGCAGGGTGTTGAGGAGCAGATTTCAGAAGTGCGACAGCGACGACAACGACAACGACAACGACAACGAGGACGACGACGGCGACGACGGCAGCAACAGCAGCGTGGCCGATAGGTCTGTGAACTTCAGCGTGAGCGGCCTGCTGTCTGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGAGCGAGCCCGGACTGTCTGTGAAACGGAAGCAGAGAAGGAGCAGAACAACCTTCACAGCTGAACAGCTGGAGGAACTGGAGAGGGCCTTTGAGAGGACACACTACCCTGATATCTACACCCGGGAAGAGCTGGCTCAGAGAACAAAGTTGACCGAGGCCAGAGTTCAGGTGTGGTTCAGCAACCGGAGGGCCAGATGGAGAAAGCAGATGGGCTCTAACCAGTTGACCGCTCTGAATAGCATCCTGCAGGTGCCCCAGGGAATGGGAACACCTAGCTACATGCTGCACGAGCCCGGATACCCTTTGTCTCACAACGCCGATAACCTGTGGCACAGGTCTAGCATGGCTCAGTCTCTGCAGAGCTTCGGACAGACAATCAAGCCCGAGAACAGCTACGCCGGATTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGACTGCCTACACATAGCAGCAGCGACCCTTTGTCTTCTACCTGGTCTAGCCCTGTGTCTACCTCTGTGCCTGCTCTGGGATATACCCCTTCTAGCGGACACTATCACCACTACAGCGACGTGACCAAGTCTACCCTGCACAGCTACAACGCCCACATCCCCTCTGTGACCAACATGGAGCGGTGCAGCGTGGACGACTCTCTGGTGGCTCTGAGAATGAAAAGCCGGGAGCACAGCGCCGCCCTGTCTTTGATGCAGGTGGCCGATAACAAGATGGCCACCTCTTTCTGA
>Optimized sequence - alternative variant 1
ATGCTGACCAGCGGCCTGGTTGTGAGCAACATGTTCTCTTACCACCTGGCCGCTCTGGGATTAATGCCTTCTTTTCAGATGGAGGGCAGGGGCAGAGTGAATCAGTTAGGAGGAGTGTTTATTAACGGCAGGCCTTTACCTAATCACATTCGGCTGAAAATCGTGGAGTTAGCTGCTCAGGGAGTTAGACCTTGTGTGATCAGCAGGCAGTTGAGAGTGTCTCACGGATGTGTGTCTAAAATCCTGCAGAGATACCAGGAGACAGGCAGCATCAGGCCTGGAGTGATTGGAGGCTCTAAACCCAGAGTGGCTACACCTGAAGTGGAGAAAAAGATCGAGCAATACAAGAAGGACAACCCCGGCATCTTCTCTTGGGAGATCCGGGACAGGTTGCTGAAGGAGGGAATCTGTGATAGGTCTACAGTGCCTTCTGTGTCTAGCATCAGCAGGGTGTTGAGAAGCAGATTCCAGAAATGCGACAGCGACGATAACGACAACGACAACGACAACGAGGACGACGACGGCGATGATGGCTCTAATAGCAGCGTGGCCGATAGATCTGTGAACTTCAGCGTGAGCGGATTGCTGTCTGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGTCTGAGCCTGGACTGTCTGTTAAACGGAAGCAGAGGAGGAGCAGAACAACATTCACAGCCGAGCAGTTGGAGGAACTGGAGAGGGCCTTTGAGAGAACCCACTATCCCGATATCTACACCAGGGAAGAACTGGCTCAGAGAACAAAACTGACCGAGGCCAGAGTTCAGGTGTGGTTTAGCAACCGGAGGGCCAGATGGAGAAAACAGATGGGCAGCAACCAGTTAACAGCCCTGAACAGCATCCTGCAGGTGCCTCAAGGAATGGGAACACCTTCTTATATGCTGCACGAGCCTGGATATCCTTTATCTCATAACGCCGATAACCTGTGGCACAGGTCTTCTATGGCTCAATCTCTGCAGTCTTTCGGACAGACAATTAAGCCCGAGAACTCTTACGCCGGATTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGACTGCCTACACATTCTTCTAGCGACCCTTTATCTTCTACATGGTCTAGCCCTGTGTCTACATCTGTGCCTGCTCTGGGATATACACCTTCTTCTGGCCACTATCACCACTACAGCGACGTGACCAAGAGCACCCTGCATTCTTACAACGCCCACATTCCCAGCGTGACCAACATGGAGCGGTGCTCTGTGGATGACAGCTTAGTGGCTCTGAGAATGAAAAGCAGGGAACACTCTGCTGCTCTGTCTTTAATGCAGGTGGCCGATAATAAGATGGCCACCTCTTTCTGA
>Optimized sequence - alternative variant 2
ATGCTGACCAGCGGATTGGTGGTGAGCAACATGTTCTCTTACCACCTGGCTGCTCTGGGCCTGATGCCCTCTTTCCAGATGGAGGGCCGGGGCAGGGTGAATCAGTTGGGCGGAGTGTTTATCAACGGCCGGCCCCTGCCTAATCACATCCGGCTGAAGATCGTGGAGTTGGCTGCTCAGGGAGTGAGACCCTGCGTGATCAGCCGGCAGTTGAGGGTGTCTCACGGATGTGTGAGCAAGATCCTGCAGAGATACCAGGAGACCGGCAGCATCCGGCCCGGAGTGATTGGCGGCTCTAAGCCCAGGGTGGCTACACCTGAAGTGGAGAAGAAGATCGAGCAGTATAAGAAGGACAACCCCGGCATCTTCTCTTGGGAGATCCGGGACCGGCTGCTGAAGGAGGGCATCTGCGACCGGAGCACCGTGCCCAGCGTGAGCAGCATCAGCCGGGTGCTGCGGAGCCGGTTCCAGAAGTGCGACAGCGACGACAACGACAACGACAACGACAACGAGGACGACGACGGCGACGACGGCAGCAACAGCAGCGTGGCCGACCGGAGCGTGAACTTCAGCGTGAGCGGCCTGCTGAGCGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGAGCGAGCCCGGCCTGAGCGTGAAGCGGAAGCAGCGGCGGAGCCGGACCACCTTCACCGCTGAGCAGTTGGAGGAGTTGGAGCGGGCCTTCGAGCGGACCCACTACCCCGACATCTACACCCGGGAGGAGTTGGCCCAGCGGACCAAGTTGACCGAGGCCCGGGTGCAGGTGTGGTTCAGCAACCGGCGGGCCAGATGGAGGAAACAGATGGGCAGCAACCAGTTGACCGCTCTGAACAGCATCCTGCAGGTGCCCCAGGGCATGGGCACCCCTTCTTACATGCTGCACGAGCCCGGATACCCTTTGAGCCACAACGCCGACAACCTGTGGCACCGGAGCAGCATGGCCCAGAGCCTGCAGTCTTTCGGCCAGACCATCAAGCCCGAGAACTCTTACGCCGGCCTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGCCTGCCCACCCACAGCAGCAGCGACCCCCTGAGCAGCACCTGGAGCAGCCCTGTGTCTACCAGCGTGCCTGCTCTGGGATATACCCCCAGCAGCGGACACTATCACCACTACAGCGACGTGACCAAGAGCACCCTGCACTCTTACAACGCCCACATCCCCAGCGTGACCAACATGGAGCGGTGCAGCGTGGACGACAGCCTGGTGGCCCTGCGGATGAAGAGCCGGGAGCACAGCGCTGCTCTGAGCCTGATGCAGGTGGCCGACAACAAGATGGCCACCTCTTTCTGA
2.2.2 Creating RNAi / RNAi + expression of the plasmid vector
Empty input dictionary schema:
input_dict = {
# REQUIRED!
# name of current project (defined by user)
'project_name':'',
# REQUIRED!
# avaiable of vector types (ssAAV / scAAV / lentiviral / regular)
'vector_type':'ssAAV',
# REQUIRED!
# in this case 'vector_function':'rnai'
'vector_function':'rnai',
# REQUIRED!
# avaiable options (human / mouse / rat / both (mouse + human) / both2 (rat + human) / multi (mouse + rat + human))
# 'both / both2 / multi' - creating vector function adjusted for all species taking into consideration most adjustments for Homo sapiens
'species':'human',
# REQUIRED!
# sequence of provided non-coding promoter
# for coding sequences the user should choose the promoter of non-coding genes (metadata['promoters']['type'] == 'non-coding')
# sequence orientation 5' ---> 3' - sense
'promoter_ncrna_sequence':'',
# REQUIRED!
# name of provided promoter sequence
'promoter_ncrna_name':'',
# OPTIONAL!
# sequence of custom RNAi, which can be provided by user
# if provided, then the algorithm of RNAi estimation is off
# if empt '' the algorithm share the best possible RNAi based on 'rnai_gene_name'
# sequence orientation 5' ---> 3' - sense
'rnai_sequence':'',
# REQUIRED; if rnai_sequence is empty!
# length of RNAi (defined by user)
# recomended range: 20–25
'rnai_length':20,
# OPTIONAL; if rnai_sequence is empty!
# overhang sequence of RNAi sequence 3' end (defined by user)
# recomended 'UU'
'overhang_3_prime':'UU',
# REQUIRED!
# name of the target gene for the RNAi searching algorithm (gene name for Homo sapien / Mus musculus / Rattus norvegicus)
# algorithm is working when the rnai_sequence is empty ''
# if the user defines 'rnai_sequence' this 'rnai_gene_name' is just a name for a user-supplied sequence
'rnai_gene_name':'',
# REQUIRED!
# sequence of the loop to create the structure of the hairpin of shRNA or siRNA depending on the loop sequence
# algorithm is working when the rnai_sequence is empty ''
# if the user defines 'rnai_sequence' this 'rnai_gene_name' is just a name for a user-supplied sequence
# sequence orientation 5' ---> 3' - sense
'loop_sequence':'',
# WARNING! If the user wants to add additional CDS sequences to parallel transcript expression with silencing by RNAi in one vector; provide sequences, linkers_sequences, promoter_sequence, etc.
# list of coding sequences (CDS) provided to make expression from the vector
# amount of sequences is not restricted as the user must remember that the length of whole vector is limited
# excide the relevant vector size can decrease vector working
# if the user wants to not include any sequences only fluorescent_tag, provide ['']
# sequences orientation 5' ---> 3' - sense
'sequences':[''],
# list of names of coding sequences
# amount of names should be equal with amount of sequences
# if provided no sequences, provide ['']
'sequences_names':[''],
# REQUIRED if more than one sequence of transcripts!
# sequences of provided linkers
# number of linkers_sequences should be equal number of sequences (transcripts) - 1. One linker for each pair of sequences.
# if the number of transcript sequences is equal 1 then provide empty list []
# if the user wants to not provide any linkers between the transcript sequences, provide an empty string '' for each pair of transcripts where the user wants to avoid linker; empty strings '' provide inside the list ['']
'linkers_sequences':[''],
# REQUIRED if transcript sequence occures, if not empty string ''!
# names of provided linkers
# if the number of transcript sequences is equal 1 then provide empty list []
# if the user wants to not provide any linkers between the transcript sequences, provide an empty string '' for each pair of transcripts where the user wants to avoid linker; empty strings '' provide inside the list ['']
'linkers_names':[''],
# REQUIRED if transcript sequence occurs or fluorescent tag will be included, if not empty string ''!
# sequence of provided promoter
# sequence orientation 5' ---> 3' - sense
'promoter_sequence':'',
# REQUIRED if transcript sequence occurs or fluorescent tag will be included, if not empty string ''!
# name of provided promoter sequence
'promoter_name':'',
# OPTIONAL if transcript sequence occures or fluorescent tag will be included, if not empty string ''!
# sequence of provided enhancer
# sequence orientation 5' ---> 3' - sense
'regulator_sequence':'',
# OPTIONAL if transcript sequence occures or fluorescent tag will be included, if not empty string ''!
# name of provided enhancer sequence
'regulator_name':'',
# REQUIRED if transcript sequence occurs or fluorescent tag will be included, if not empty string ''!
# sequence of provided polyA signal
# sequence orientation 5' ---> 3' - sense
'polya_sequence':'',
# REQUIRED if transcript sequence occurs or fluorescent tag will be included, if not empty string ''!
# name of provided polyA singla sequence
'polya_name':'',
# OPTIONAL!
# sequence of provided fluorescent tag
# if the user does not need fluorescent tag, provide ''
# sequence orientation 5' ---> 3' - sense
'fluorescence_sequence':'',
# OPTIONAL!
# name of provided fluorescent tag
# if the user does not need fluorescent tag, provide ''
'fluorescence_name':'',
# WARNING! If provided sequences for transcripts (> 0), provide fluorescence_linker_sequence or provide empty string ''.
# OPTIONAL if transcript sequence occures, if not empty string ''!
# sequence of provided fluorescence tag linker
# sequence orientation 5' ---> 3' - sense
'fluorescence_linker_sequence':'',
# OPTIONAL if transcript sequence occures, if not empty string ''!
# name of provided fluorescence tag linker
'fluorescence_linker_name':'',
# REQUIRED!
# sequence of provided selection marker
# sequence orientation 5' ---> 3' - sense
'selection_marker_sequence':'',
# REQUIRED!
# name of provided selection marker
'selection_marker_name':'',
# OPTIONAL!
# restriction enzymes protection of transcript sequences
# if the user does not need any restriction places protection, provide empty list []
'restriction_list':[],
# REQUIRED!
# available options (True / False)
# decision; if the user wants the transcription sequences optimized based on the provided species
# if the user has omitted the additional transcript sequences, provide False
'optimize':True,
# REQUIRED; if optimize == True!
# user-defined percent of GC% content in predicted/optimized sequence
'transcript_GC':58,
# REQUIRED; if optimize == True!
# user-defined maximum number of consecutive repeats of a single nucleotide (A, C, T(U), G) in the predicted/optimized sequence, eg. AAAAAA
'poly_len':7
}
Example dictionary:
input_dict = {
'project_name':'test_RNAi',
'vector_type':'ssAAV',
'vector_function':'rnai',
'species':'human',
'promoter_ncrna_name':'U6',
'promoter_ncrna_sequence':'GAGGGCCTATTTCCCATGATTCCTTCATATTTGCATATACGATACAAGGCTGTTAGAGAGATAATTGGAATTAATTTGACTGTAAACACAAAGATATTAGTACAAAATACGTGACGTAGAAAGTAATAATTTCTTGGGTAGTTTGCAGTTTTAAAATTATGTTTTAAAATGGACTATCATATGCTTACCGTAACTTGAAAGTATTTCGATTTCTTGGCTTTATATATCTTGTGGAAAGGACGAAACACC',
'rnai_sequence':'',
'rnai_length':20,
'overhang_3_prime':'UU',
'rnai_gene_name':'PAX3',
'loop_sequence':'TAGTGAAGCCACAGATGTAC',
'sequences':['ATGGCGATGAGCAGCGGCGGCAGTGGTGGCGGCGTCCCGGAGCAGGAGGATTCCGTGCTGTTCCGGCGCGGCACAGGCCAGAGCGATGATTCTGACATTTGGGATGATACAGCACTGATAAAAGCATATGATAAAGCTGTGGCTTCATTTAAGCATGCTCTAAAGAATGGTGACATTTGTGAAACTTCGGGTAAACCAAAAACCACACCTAAAAGAAAACCTGCTAAGAAGAATAAAAGCCAAAAGAAGAATACTGCAGCTTCCTTACAACAGTGGAAAGTTGGGGACAAATGTTCTGCCATTTGGTCAGAAGACGGTTGCATTTACCCAGCTACCATTGCTTCAATTGATTTTAAGAGAGAAACCTGTGTTGTGGTTTACACTGGATATGGAAATAGAGAGGAGCAAAATCTGTCCGATCTACTTTCCCCAATCTGTGAAGTAGCTAATAATATAGAACAAAATGCTCAAGAGAATGAAAATGAAAGCCAAGTTTCAACAGATGAAAGTGAGAACTCCAGGTCTCCTGGAAATAAATCAGATAACATCAAGCCCAAATCTGCTCCATGGAACTCTTTTCTCCCTCCACCACCCCCCATGCCAGGGCCAAGACTGGGACCAGGAAAGCCAGGTCTAAAATTCAATGGCCCACCACCGCCACCGCCACCACCACCACCCCACTTACTATCATGCTGGCTGCCTCCATTTCCTTCTGGACCACCAATAATTCCCCCACCACCTCCCATATGTCCAGATTCTCTTGATGATGCTGATGCTTTGGGAAGTATGTTAATTTCATGGTACATGAGTGGCTATCATACTGGCTATTATATGTTTCCTGAGGCCTCCCTAAAAGCCGAGCAGATGCCAGCACCATGCTTCCTGTAA'],
'sequences_names':['SMN1'],
'linkers_sequences':[''],
'linkers_names':[''],
'promoter_sequence':'GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGAT',
'promoter_name':'TBG',
'regulator_sequence':'CGATAATCAACCTCTGGATTACAAAATTTGTGAAAGATTGACTGGTATTCTTAACTATGTTGCTCCTTTTACGCTATGTGGATACGCTGCTTTAATGCCTTTGTATCATGCTATTGCTTCCCGTATGGCTTTCATTTTCTCCTCCTTGTATAAATCCTGGTTGCTGTCTCTTTATGAGGAGTTGTGGCCCGTTGTCAGGCAACGTGGCGTGGTGTGCACTGTGTTTGCTGACGCAACCCCCACTGGTTGGGGCATTGCCACCACCTGTCAGCTCCTTTCCGGGACTTTCGCTTTCCCCCTCCCTATTGCCACGGCGGAACTCATCGCCGCCTGCCTTGCCCGCTGCTGGACAGGGGCTCGGCTGTTGGGCACTGACAATTCCGTGGTGTTGTCGGGGAAGCTGACGTCCTTTCCATGGCTGCTCGCCTGTGTTGCCACCTGGATTCTGCGCGGGACGTCCTTCTGCTACGTCCCTTCGGCCCTCAATCCAGCGGACCTTCCTTCCCGCGGCCTGCTGCCGGCTCTGCGGCCTCTTCCGCGTCTTCGCCTTCGCCCTCAGACGAGTCGGATCTCCCTTTGGGCCGCCTCCCCGCATCGG',
'regulator_name':'WPRE',
'polya_sequence':'CAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTA',
'polya_name':'SV40_late',
'fluorescence_sequence':'ATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGGGCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCATCTGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGACCTACGGCGTGCAGTGCTTCAGCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTCTTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGGACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCATGGCCGACAAGCAGAAGAACGGCATCAAGGTGAACTTCAAGATCCGCCACAACATCGAGGACGGCAGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCACTACCTGAGCACCCAGTCCGCCCTGAGCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCGTGACCGCCGCCGGGATCACTCTCGGCATGGACGAGCTGTACAAGTAA',
'fluorescence_name':'EGFP',
'fluorescence_linker_sequence':'GGAAGCGGAGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCCCGGCCCC',
'fluorescence_linker_name':'T2A',
'selection_marker_sequence':'ATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATTTTGCCTTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGTTACATCGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTTTTAAAGTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTCTCAGAATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGTGCTGCCATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTTTTTTGCACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGAGCGTGACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCCCGGCAACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGTTTATTGCTGATAAATCTGGAGCCGGTGAGCGTGGAAGCCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTCCCGTATCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCCTCACTGATTAAGCATTGGTAA',
'selection_marker_name':'Ampicillin',
'restriction_list':['RsaI', 'MnlI', 'AciI', 'AluI', 'BmrI'],
'optimize':True,
'transcript_GC':58,
'poly_len':7
}
Output:
# Name of project
project['project']
# Graph of the designed vector
vector_plot = project['vector']['graph']
vector_plot.savefig('RNAi_vector.svg')
Example return:

# Complete FASTA file of the designed vecotr
project['vector']['full_fasta']
Example return:
>test_RNAi_ssAAV_rnai_6790nc
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTCAGTGAGCGAGCGAGCGCGCAGAGAGGGAGTGGCCAACTCCATCACTAGGGGTTCCTATCGATGAGGGCCTATTTCCCATGATTCCTTCATATTTGCATATACGATACAAGGCTGTTAGAGAGATAATTGGAATTAATTTGACTGTAAACACAAAGATATTAGTACAAAATACGTGACGTAGAAAGTAATAATTTCTTGGGTAGTTTGCAGTTTTAAAATTATGTTTTAAAATGGACTATCATATGCTTACCGTAACTTGAAAGTATTTCGATTTCTTGGCTTTATATATCTTGTGGAAAGGACGAAACACCGGGCCTTTCCGTTTCGCCTTCACCTTAGTGAAGCCACAGATGTACAGGTGAAGGCGAAACGGAAAGGCTTTTTTTGAATTCCAACTTTGTATAGAAAAGTTGGGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGATCAAGTTTGTACAAAAAAGCAGGCTGCCACCATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTGGGAAGCGGAGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCCCGGCCCCATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGGGCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCATCTGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGACCTACGGCGTGCAGTGCTTCAGCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTCTTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGGACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCATGGCCGACAAGCAGAAGAACGGCATCAAGGTGAACTTCAAGATCCGCCACAACATCGAGGACGGCAGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCACTACCTGAGCACCCAGTCCGCCCTGAGCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCGTGACCGCCGCCGGGATCACTCTCGGCATGGACGAGCTGTACAAGTAACGATAATCAACCTCTGGATTACAAAATTTGTGAAAGATTGACTGGTATTCTTAACTATGTTGCTCCTTTTACGCTATGTGGATACGCTGCTTTAATGCCTTTGTATCATGCTATTGCTTCCCGTATGGCTTTCATTTTCTCCTCCTTGTATAAATCCTGGTTGCTGTCTCTTTATGAGGAGTTGTGGCCCGTTGTCAGGCAACGTGGCGTGGTGTGCACTGTGTTTGCTGACGCAACCCCCACTGGTTGGGGCATTGCCACCACCTGTCAGCTCCTTTCCGGGACTTTCGCTTTCCCCCTCCCTATTGCCACGGCGGAACTCATCGCCGCCTGCCTTGCCCGCTGCTGGACAGGGGCTCGGCTGTTGGGCACTGACAATTCCGTGGTGTTGTCGGGGAAGCTGACGTCCTTTCCATGGCTGCTCGCCTGTGTTGCCACCTGGATTCTGCGCGGGACGTCCTTCTGCTACGTCCCTTCGGCCCTCAATCCAGCGGACCTTCCTTCCCGCGGCCTGCTGCCGGCTCTGCGGCCTCTTCCGCGTCTTCGCCTTCGCCCTCAGACGAGTCGGATCTCCCTTTGGGCCGCCTCCCCGCATCGGACCCAGCTTTCTTGTACAAAGTGGTGATGGCCGGCCGCTTCGAGCAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTAATCGATAGATCTAGGAACCCCTAGTGATGGAGTTGGCCACTCCCTCTCTGCGCGCTCGCTCGCTCACTGAGGCCGGGCGACCAAAGGTCGCCCGACGCCCGGGCTTTGCCCGGGCGGCCTCAGTGAGCGAGCGAGCGCGCAGCTGCCTGCAGGCAGCTTGGCACTGGCCGTCGTTTTACAACGTCGTGACTGGGAAAACCCTGGCGTTACCCAACTTAATCGCCTTGCAGCACATCCCCCTTTCGCCAGCTGGCGTAATAGCGAAGAGGCCCGCACCGATCGCCCTTCCCAACAGTTGCGCAGCCTGAATGGCGAATGGCGCCTGATGCGGTATTTTCTCCTTACGCATCTGTGCGGTATTTCACACCGCATACGTCAAAGCAACCATAGTACGCGCCCTGTAGCGGCGCATTAAGCGCGGCGGGTGTGGTGGTTACGCGCAGCGTGACCGCTACACTTGCCAGCGCCCTAGCGCCCGCTCCTTTCGCTTTCTTCCCTTCCTTTCTCGCCACGTTCGCCGGCTTTCCCCGTCAAGCTCTAAATCGGGGGCTCCCTTTAGGGTTCCGATTTAGTGCTTTACGGCACCTCGACCCCAAAAAACTTGATTTGGGTGATGGTTCACGTAGTGGGCCATCGCCCTGATAGACGGTTTTTCGCCCTTTGACGTTGGAGTCCACGTTCTTTAATAGTGGACTCTTGTTCCAAACTGGAACAACACTCAACCCTATCTCGGGCTATTCTTTTGATTTATAAGGGATTTTGCCGATTTCGGCCTATTGGTTAAAAAATGAGCTGATTTAACAAAAATTTAACGCGAATTTTAACAAAATATTAACGTTTACAATTTTATGGTGCACTCTCAGTACAATCTGCTCTGATGCCGCATAGTTAAGCCAGCCCCGACACCCGCCAACACCCGCTGACGCGCCCTGACGGGCTTGTCTGCTCCCGGCATCCGCTTACAGACAAGCTGTGACCGTCTCCGGGAGCTGCATGTGTCAGAGGTTTTCACCGTCATCACCGAAACGCGCGAGACGAAAGGGCCTCGTGATACGCCTATTTTTATAGGTTAATGTCATGATAATAATGGTTTCTTAGACGTCAGGTGGCACTTTTCGGGGAAATGTGCGCGGAACCCCTATTTGTTTATTTTTCTAAATACATTCAAATATGTATCCGCTCATGAGACAATAACCCTGATAAATGCTTCAATAATATTGAAAAAGGAAGAGTATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATTTTGCCTTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGTTACATCGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTTTTAAAGTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTCTCAGAATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGTGCTGCCATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTTTTTTGCACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGAGCGTGACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCCCGGCAACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGTTTATTGCTGATAAATCTGGAGCCGGTGAGCGTGGAAGCCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTCCCGTATCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCCTCACTGATTAAGCATTGGTAACTGTCAGACCAAGTTTACTCATATATACTTTAGATTGATTTAAAACTTCATTTTTAATTTAAAAGGATCTAGGTGAAGATCCTTTTTGATAATCTCATGACCAAAATCCCTTAACGTGAGTTTTCGTTCCACTGAGCGTCAGACCCCGTAGAAAAGATCAAAGGATCTTCTTGAGATCCTTTTTTTCTGCGCGTAATCTGCTGCTTGCAAACAAAAAAACCACCGCTACCAGCGGTGGTTTGTTTGCCGGATCAAGAGCTACCAACTCTTTTTCCGAAGGTAACTGGCTTCAGCAGAGCGCAGATACCAAATACTGTTCTTCTAGTGTAGCCGTAGTTAGGCCACCACTTCAAGAACTCTGTAGCACCGCCTACATACCTCGCTCTGCTAATCCTGTTACCAGTGGCTGCTGCCAGTGGCGATAAGTCGTGTCTTACCGGGTTGGACTCAAGACGATAGTTACCGGATAAGGCGCAGCGGTCGGGCTGAACGGGGGGTTCGTGCACACAGCCCAGCTTGGAGCGAACGACCTACACCGAACTGAGATACCTACAGCGTGAGCTATGAGAAAGCGCCACGCTTCCCGAAGGGAGAAAGGCGGACAGGTATCCGGTAAGCGGCAGGGTCGGAACAGGAGAGCGCACGAGGGAGCTTCCAGGGGGAAACGCCTGGTATCTTTATAGTCCTGTCGGGTTTCGCCACCTCTGACTTGAGCGTCGATTTTTGTGATGCTCGTCAGGGGGGCGGAGCCTATGGAAAAACGCCAGCAACGCGGCCTTTTTACGGTTCCTGGCCTTTTGCTGGCCTTTTGCTCACATGTTCTTTCCTGCGTTATCCCCTGATTCTGTGGATAACCGTATTACCGCCTTTGAGTGAGCTGATACCGCTCGCCGCAGCCGAACGACCGAGCGCAGCGAGTCAGTGAGCGAGGAAGCGGAAGAGCGCCCAATACGCAAACCGCCTCTCCCCGCGCGTTGGCCGATTCATTAATGCAGCTGGCACGACAGGTTTCCCGACTGGAAAGCGGGCAGTGAGCGCAACGCAATTAATGTGAGTTAGCTCACTCATTAGGCACCCCAGGCTTTACACTTTATGCTTCCGGCTCGTATGTTGTGTGGAATTGTGAGCGGATAACAATTTCACACAGGAAACAGCTATGACCATGATTACGAATTGCCTGCAGGCAG
# The FASTA file is divided into particular elements of the designed vector
project['vector']['fasta']
Example return:
# test_RNAi_ssAAV_rnai_6790nc
>5`ITR
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTCAGTGAGCGAGCGAGCGCGCAGAGAGGGAGTGGCCAACTCCATCACTAGGGGTTCCT
>backbone_element
ATCGAT
>Promoter_ncRNA : U6
GAGGGCCTATTTCCCATGATTCCTTCATATTTGCATATACGATACAAGGCTGTTAGAGAGATAATTGGAATTAATTTGACTGTAAACACAAAGATATTAGTACAAAATACGTGACGTAGAAAGTAATAATTTCTTGGGTAGTTTGCAGTTTTAAAATTATGTTTTAAAATGGACTATCATATGCTTACCGTAACTTGAAAGTATTTCGATTTCTTGGCTTTATATATCTTGTGGAAAGGACGAAACACC
>backbone_element
GG
>RNAi : PAX3_RNAi_35_hs
GCCTTTCCGTTTCGCCTTCACCTTAGTGAAGCCACAGATGTACAGGTGAAGGCGAAACGGAAAGGCTT
>Terminator
TTTTT
>backbone_element
GAATTCCAACTTTGTATAGAAAAGTTG
>Promoter : TBG
GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGAT
>backbone_element
CAAGTTTGTACAAAAAAGCAGGCTGCCACC
>SEQ1 : SMN1
ATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTG
>Fluorescent_tag_linker : T2A
GGAAGCGGAGAGGGCAGGGGAAGTCTTCTAACATGCGGGGACGTGGAGGAAAATCCCGGCCCC
>Fluorescent_tag : EGFP
ATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGGGCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCATCTGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGACCTACGGCGTGCAGTGCTTCAGCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTCTTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGGACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCATGGCCGACAAGCAGAAGAACGGCATCAAGGTGAACTTCAAGATCCGCCACAACATCGAGGACGGCAGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCACTACCTGAGCACCCAGTCCGCCCTGAGCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCGTGACCGCCGCCGGGATCACTCTCGGCATGGACGAGCTGTACAAGTAA
>Enhancer : WPRE
CGATAATCAACCTCTGGATTACAAAATTTGTGAAAGATTGACTGGTATTCTTAACTATGTTGCTCCTTTTACGCTATGTGGATACGCTGCTTTAATGCCTTTGTATCATGCTATTGCTTCCCGTATGGCTTTCATTTTCTCCTCCTTGTATAAATCCTGGTTGCTGTCTCTTTATGAGGAGTTGTGGCCCGTTGTCAGGCAACGTGGCGTGGTGTGCACTGTGTTTGCTGACGCAACCCCCACTGGTTGGGGCATTGCCACCACCTGTCAGCTCCTTTCCGGGACTTTCGCTTTCCCCCTCCCTATTGCCACGGCGGAACTCATCGCCGCCTGCCTTGCCCGCTGCTGGACAGGGGCTCGGCTGTTGGGCACTGACAATTCCGTGGTGTTGTCGGGGAAGCTGACGTCCTTTCCATGGCTGCTCGCCTGTGTTGCCACCTGGATTCTGCGCGGGACGTCCTTCTGCTACGTCCCTTCGGCCCTCAATCCAGCGGACCTTCCTTCCCGCGGCCTGCTGCCGGCTCTGCGGCCTCTTCCGCGTCTTCGCCTTCGCCCTCAGACGAGTCGGATCTCCCTTTGGGCCGCCTCCCCGCATCGG
>backbone_element
ACCCAGCTTTCTTGTACAAAGTGGTGATGGCCGGCCGCTTCGAG
>PolyA_signal : SV40_late
CAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTA
>backbone_element
ATCGATAGATCT
>3`ITR
AGGAACCCCTAGTGATGGAGTTGGCCACTCCCTCTCTGCGCGCTCGCTCGCTCACTGAGGCCGGGCGACCAAAGGTCGCCCGACGCCCGGGCTTTGCCCGGGCGGCCTCAGTGAGCGAGCGAGCGCGCAG
>backbone_element
CTGCCTGCAGGCAGCTTGGCACTGGCCGTCGTTTTACAACGTCGTGACTGGGAAAACCCTGGCGTTACCCAACTTAATCGCCTTGCAGCACATCCCCCTTTCGCCAGCTGGCGTAATAGCGAAGAGGCCCGCACCGATCGCCCTTCCCAACAGTTGCGCAGCCTGAATGGCGAATGGCGCCTGATGCGGTATTTTCTCCTTACGCATCTGTGCGGTATTTCACACCGCATACGTCAAAGCAACCATAGTACGCGCCCTGTAGCGGCGCATTAAGCGCGGCGGGTGTGGTGGTTACGCGCAGCGTGACCGCTACACTTGCCAGCGCCCTAGCGCCCGCTCCTTTCGCTTTCTTCCCTTCCTTTCTCGCCACGTTCGCCGGCTTTCCCCGTCAAGCTCTAAATCGGGGGCTCCCTTTAGGGTTCCGATTTAGTGCTTTACGGCACCTCGACCCCAAAAAACTTGATTTGGGTGATGGTTCACGTAGTGGGCCATCGCCCTGATAGACGGTTTTTCGCCCTTTGACGTTGGAGTCCACGTTCTTTAATAGTGGACTCTTGTTCCAAACTGGAACAACACTCAACCCTATCTCGGGCTATTCTTTTGATTTATAAGGGATTTTGCCGATTTCGGCCTATTGGTTAAAAAATGAGCTGATTTAACAAAAATTTAACGCGAATTTTAACAAAATATTAACGTTTACAATTTTATGGTGCACTCTCAGTACAATCTGCTCTGATGCCGCATAGTTAAGCCAGCCCCGACACCCGCCAACACCCGCTGACGCGCCCTGACGGGCTTGTCTGCTCCCGGCATCCGCTTACAGACAAGCTGTGACCGTCTCCGGGAGCTGCATGTGTCAGAGGTTTTCACCGTCATCACCGAAACGCGCGAGACGAAAGGGCCTCGTGATACGCCTATTTTTATAGGTTAATGTCATGATAATAATGGTTTCTTAGACGTCAGGTGGCACTTTTCGGGGAAATGTGCGCGGAACCCCTATTTGTTTATTTTTCTAAATACATTCAAATATGTATCCGCTCATGAGACAATAACCCTGATAAATGCTTCAATAATATTGAAAAAGGAAGAGT
>Resistance : Ampicillin
ATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATTTTGCCTTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGTTACATCGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTTTTAAAGTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTCTCAGAATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGTGCTGCCATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTTTTTTGCACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGAGCGTGACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCCCGGCAACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGTTTATTGCTGATAAATCTGGAGCCGGTGAGCGTGGAAGCCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTCCCGTATCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCCTCACTGATTAAGCATTGGTAA
>backbone_element
CTGTCAGACCAAGTTTACTCATATATACTTTAGATTGATTTAAAACTTCATTTTTAATTTAAAAGGATCTAGGTGAAGATCCTTTTTGATAATCTCATGACCAAAATCCCTTAACGTGAGTTTTCGTTCCACTGAGCGTCAGACCCCGTAGAAAAGATCAAAGGATCTTC
>pUC_ori
TTGAGATCCTTTTTTTCTGCGCGTAATCTGCTGCTTGCAAACAAAAAAACCACCGCTACCAGCGGTGGTTTGTTTGCCGGATCAAGAGCTACCAACTCTTTTTCCGAAGGTAACTGGCTTCAGCAGAGCGCAGATACCAAATACTGTTCTTCTAGTGTAGCCGTAGTTAGGCCACCACTTCAAGAACTCTGTAGCACCGCCTACATACCTCGCTCTGCTAATCCTGTTACCAGTGGCTGCTGCCAGTGGCGATAAGTCGTGTCTTACCGGGTTGGACTCAAGACGATAGTTACCGGATAAGGCGCAGCGGTCGGGCTGAACGGGGGGTTCGTGCACACAGCCCAGCTTGGAGCGAACGACCTACACCGAACTGAGATACCTACAGCGTGAGCTATGAGAAAGCGCCACGCTTCCCGAAGGGAGAAAGGCGGACAGGTATCCGGTAAGCGGCAGGGTCGGAACAGGAGAGCGCACGAGGGAGCTTCCAGGGGGAAACGCCTGGTATCTTTATAGTCCTGTCGGGTTTCGCCACCTCTGACTTGAGCGTCGATTTTTGTGATGCTCGTCAGGGGGGCGGAGCCTATGGAAA
>backbone_element
AACGCCAGCAACGCGGCCTTTTTACGGTTCCTGGCCTTTTGCTGGCCTTTTGCTCACATGTTCTTTCCTGCGTTATCCCCTGATTCTGTGGATAACCGTATTACCGCCTTTGAGTGAGCTGATACCGCTCGCCGCAGCCGAACGACCGAGCGCAGCGAGTCAGTGAGCGAGGAAGCGGAAGAGCGCCCAATACGCAAACCGCCTCTCCCCGCGCGTTGGCCGATTCATTAATGCAGCTGGCACGACAGGTTTCCCGACTGGAAAGCGGGCAGTGAGCGCAACGCAATTAATGTGAGTTAGCTCACTCATTAGGCACCCCAGGCTTTACACTTTATGCTTCCGGCTCGTATGTTGTGTGGAATTGTGAGCGGATAACAATTTCACACAGGAAACAGCTATGACCATGATTACGAATTGCCTGCAGGCAG
# Top 1 designed RNAi in shRNA form information
# RNAi name
project['rnai']['name']
# RNAi sequence
project['rnai']['sequence']
# RNAi prediction structure
rnai_prediction = project['rnai']['figure']
rnai_prediction.savefig('rnai_predicted_structure.svg')
Example return:

# other selected RNAi with statistics
pd.DataFrame(project['rnai']['full_data'])
Examples with structure description presented in:
- 1.23. Prediction of RNAi on the provided sequence
- 1.24. Correcting of RNAi_data for complementarity to the loop sequence
- 1.25. Correcting of RNAi_data for complementarity to the additional external sequence
Sequences output data:
## *if occur user-defined sequences for additional expression in the plasmid vector
## genes names
project['transcripts']['sequences']['name']
## proteins sequences
project['transcripts']['sequences']['sequence_aa']
## average codon frequency in the input sequence
project['transcripts']['sequences']['vector_sequence_frequence']
## GC% content in the input sequence
project['transcripts']['sequences']['vector_sequence_GC']
############################################################################
## average codon frequency in the output sequence
project['transcripts']['sequences']['optimized_vector_sequence_frequence']
## GC% content in the output sequence
project['transcripts']['sequences']['optimized_vector_sequence_GC']
# Sequences
# Input sequence
project['transcripts']['sequences']['sequence']
# Predicted structure - input
project['transcripts']['sequences']['sequence_figure']
# Optimized sequence
project['transcripts']['sequences']['vector_sequence']
# Predicted structure - optimized
project['transcripts']['sequences']['optimized_sequence_figure']
Example return:
- Otimized sequence structure:

- Input sequence structure:

- Alternative options:
# Sequences results and metadata
# optimized sequence - main
project['transcripts']['sequences']
# optimized sequence - alternatives
project['transcripts']['alternative']
# additional variant 1
project['transcripts']['alternative']['var0]
# additional variant 2
project['transcripts']['alternative']['var1]
Example return:
>Optimized sequence - main
ATGCTGACCAGCGGATTGGTTGTGAGCAACATGTTCAGCTACCACCTGGCCGCCCTGGGATTGATGCCTTCTTTTCAGATGGAGGGCAGGGGCAGAGTGAACCAGTTGGGAGGAGTGTTTATCAACGGCCGGCCTCTGCCTAATCACATCCGGCTGAAAATCGTGGAGTTGGCCGCTCAGGGAGTTAGGCCTTGTGTGATTAGCAGGCAGCTGAGGGTGTCTCACGGATGTGTGAGCAAGATCCTGCAGCGGTACCAGGAGACCGGCTCTATCAGGCCTGGAGTGATTGGCGGCTCTAAACCCAGGGTGGCTACACCTGAAGTGGAAAAGAAGATCGAGCAGTACAAGAAGGACAACCCCGGCATCTTCAGCTGGGAGATCCGGGACCGGCTGCTGAAGGAGGGCATCTGCGACAGAAGCACCGTGCCTTCTGTGTCTAGCATCAGCAGGGTGTTGAGGAGCAGATTTCAGAAGTGCGACAGCGACGACAACGACAACGACAACGACAACGAGGACGACGACGGCGACGACGGCAGCAACAGCAGCGTGGCCGATAGGTCTGTGAACTTCAGCGTGAGCGGCCTGCTGTCTGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGAGCGAGCCCGGACTGTCTGTGAAACGGAAGCAGAGAAGGAGCAGAACAACCTTCACAGCTGAACAGCTGGAGGAACTGGAGAGGGCCTTTGAGAGGACACACTACCCTGATATCTACACCCGGGAAGAGCTGGCTCAGAGAACAAAGTTGACCGAGGCCAGAGTTCAGGTGTGGTTCAGCAACCGGAGGGCCAGATGGAGAAAGCAGATGGGCTCTAACCAGTTGACCGCTCTGAATAGCATCCTGCAGGTGCCCCAGGGAATGGGAACACCTAGCTACATGCTGCACGAGCCCGGATACCCTTTGTCTCACAACGCCGATAACCTGTGGCACAGGTCTAGCATGGCTCAGTCTCTGCAGAGCTTCGGACAGACAATCAAGCCCGAGAACAGCTACGCCGGATTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGACTGCCTACACATAGCAGCAGCGACCCTTTGTCTTCTACCTGGTCTAGCCCTGTGTCTACCTCTGTGCCTGCTCTGGGATATACCCCTTCTAGCGGACACTATCACCACTACAGCGACGTGACCAAGTCTACCCTGCACAGCTACAACGCCCACATCCCCTCTGTGACCAACATGGAGCGGTGCAGCGTGGACGACTCTCTGGTGGCTCTGAGAATGAAAAGCCGGGAGCACAGCGCCGCCCTGTCTTTGATGCAGGTGGCCGATAACAAGATGGCCACCTCTTTCTGA
>Optimized sequence - alternative variant 1
ATGCTGACCAGCGGCCTGGTTGTGAGCAACATGTTCTCTTACCACCTGGCCGCTCTGGGATTAATGCCTTCTTTTCAGATGGAGGGCAGGGGCAGAGTGAATCAGTTAGGAGGAGTGTTTATTAACGGCAGGCCTTTACCTAATCACATTCGGCTGAAAATCGTGGAGTTAGCTGCTCAGGGAGTTAGACCTTGTGTGATCAGCAGGCAGTTGAGAGTGTCTCACGGATGTGTGTCTAAAATCCTGCAGAGATACCAGGAGACAGGCAGCATCAGGCCTGGAGTGATTGGAGGCTCTAAACCCAGAGTGGCTACACCTGAAGTGGAGAAAAAGATCGAGCAATACAAGAAGGACAACCCCGGCATCTTCTCTTGGGAGATCCGGGACAGGTTGCTGAAGGAGGGAATCTGTGATAGGTCTACAGTGCCTTCTGTGTCTAGCATCAGCAGGGTGTTGAGAAGCAGATTCCAGAAATGCGACAGCGACGATAACGACAACGACAACGACAACGAGGACGACGACGGCGATGATGGCTCTAATAGCAGCGTGGCCGATAGATCTGTGAACTTCAGCGTGAGCGGATTGCTGTCTGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGTCTGAGCCTGGACTGTCTGTTAAACGGAAGCAGAGGAGGAGCAGAACAACATTCACAGCCGAGCAGTTGGAGGAACTGGAGAGGGCCTTTGAGAGAACCCACTATCCCGATATCTACACCAGGGAAGAACTGGCTCAGAGAACAAAACTGACCGAGGCCAGAGTTCAGGTGTGGTTTAGCAACCGGAGGGCCAGATGGAGAAAACAGATGGGCAGCAACCAGTTAACAGCCCTGAACAGCATCCTGCAGGTGCCTCAAGGAATGGGAACACCTTCTTATATGCTGCACGAGCCTGGATATCCTTTATCTCATAACGCCGATAACCTGTGGCACAGGTCTTCTATGGCTCAATCTCTGCAGTCTTTCGGACAGACAATTAAGCCCGAGAACTCTTACGCCGGATTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGACTGCCTACACATTCTTCTAGCGACCCTTTATCTTCTACATGGTCTAGCCCTGTGTCTACATCTGTGCCTGCTCTGGGATATACACCTTCTTCTGGCCACTATCACCACTACAGCGACGTGACCAAGAGCACCCTGCATTCTTACAACGCCCACATTCCCAGCGTGACCAACATGGAGCGGTGCTCTGTGGATGACAGCTTAGTGGCTCTGAGAATGAAAAGCAGGGAACACTCTGCTGCTCTGTCTTTAATGCAGGTGGCCGATAATAAGATGGCCACCTCTTTCTGA
>Optimized sequence - alternative variant 2
ATGCTGACCAGCGGATTGGTGGTGAGCAACATGTTCTCTTACCACCTGGCTGCTCTGGGCCTGATGCCCTCTTTCCAGATGGAGGGCCGGGGCAGGGTGAATCAGTTGGGCGGAGTGTTTATCAACGGCCGGCCCCTGCCTAATCACATCCGGCTGAAGATCGTGGAGTTGGCTGCTCAGGGAGTGAGACCCTGCGTGATCAGCCGGCAGTTGAGGGTGTCTCACGGATGTGTGAGCAAGATCCTGCAGAGATACCAGGAGACCGGCAGCATCCGGCCCGGAGTGATTGGCGGCTCTAAGCCCAGGGTGGCTACACCTGAAGTGGAGAAGAAGATCGAGCAGTATAAGAAGGACAACCCCGGCATCTTCTCTTGGGAGATCCGGGACCGGCTGCTGAAGGAGGGCATCTGCGACCGGAGCACCGTGCCCAGCGTGAGCAGCATCAGCCGGGTGCTGCGGAGCCGGTTCCAGAAGTGCGACAGCGACGACAACGACAACGACAACGACAACGAGGACGACGACGGCGACGACGGCAGCAACAGCAGCGTGGCCGACCGGAGCGTGAACTTCAGCGTGAGCGGCCTGCTGAGCGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGAGCGAGCCCGGCCTGAGCGTGAAGCGGAAGCAGCGGCGGAGCCGGACCACCTTCACCGCTGAGCAGTTGGAGGAGTTGGAGCGGGCCTTCGAGCGGACCCACTACCCCGACATCTACACCCGGGAGGAGTTGGCCCAGCGGACCAAGTTGACCGAGGCCCGGGTGCAGGTGTGGTTCAGCAACCGGCGGGCCAGATGGAGGAAACAGATGGGCAGCAACCAGTTGACCGCTCTGAACAGCATCCTGCAGGTGCCCCAGGGCATGGGCACCCCTTCTTACATGCTGCACGAGCCCGGATACCCTTTGAGCCACAACGCCGACAACCTGTGGCACCGGAGCAGCATGGCCCAGAGCCTGCAGTCTTTCGGCCAGACCATCAAGCCCGAGAACTCTTACGCCGGCCTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGCCTGCCCACCCACAGCAGCAGCGACCCCCTGAGCAGCACCTGGAGCAGCCCTGTGTCTACCAGCGTGCCTGCTCTGGGATATACCCCCAGCAGCGGACACTATCACCACTACAGCGACGTGACCAAGAGCACCCTGCACTCTTACAACGCCCACATCCCCAGCGTGACCAACATGGAGCGGTGCAGCGTGGACGACAGCCTGGTGGCCCTGCGGATGAAGAGCCGGGAGCACAGCGCTGCTCTGAGCCTGATGCAGGTGGCCGACAACAAGATGGCCACCTCTTTCTGA
2.2.3 Creating plasmid vector of in-vitro transcription of mRNA
Empty input dictionary schema:
input_dict = {
# REQUIRED!
# name of current project (defined by user)
'project_name':'',
# REQUIRED!
# avaiable of vector types (transcription)
'vector_type':'transcription',
# REQUIRED!
# in this case 'vector_function':'mrna'
'vector_function':'mrna',
# REQUIRED!
# avaiable options (human / mouse / rat / both (mouse + human) / both2 (rat + human) / multi (mouse + rat + human))
# 'both / both2 / multi' - creating vector function adjusted for all species taking into consideration most adjustments for Homo sapiens
'species':'human',
# REQUIRED!
# list of coding sequences (CDS) provided to make expression from the vector
# amount of sequences is not restricted as the user must remember that the length of whole vector is limited
# excide the relevant vector size can decrease vector working
# sequences orientation 5' ---> 3' - sense
'sequences':[''],
# REQUIRED!
# list of names of coding sequences
# amount of names should be equal with amount of sequences
# if provided no sequences, provide ['']
'sequences_names':[''],
# REQUIRED if more than one sequence of transcripts!
# sequences of provided linkers
# number of linkers_sequences should be equal number of sequences (transcripts) - 1. One linker for each pair of sequences.
# if the number of transcript sequences is equal 1 then provide empty list []
# if the user wants to not provide any linkers between the transcript sequences, provide an empty string '' for each pair of transcripts where the user wants to avoid linker; empty strings '' provide inside the list ['']
# sequence orientation 5' ---> 3' - sense
'linkers_sequences':[''],
# REQUIRED if transcript sequence occures, if not empty string ''!
# names of provided linkers
# if the number of transcript sequences is equal 1 then provide empty list []
# if the user wants to not provide any linkers between the transcript sequences, provide an empty string '' for each pair of transcripts where the user wants to avoid linker; empty strings '' provide inside the list ['']
'linkers_names':[''],
# REQUIRED!
# sequence of provided 5`UTR
# sequence orientation 5' ---> 3' - sense
'utr5_sequence':'',
# REQUIRED!
# name of provided 5`UTR
'utr5_name':'',
# REQUIRED!
# sequence of provided 3`UTR
# sequence orientation 5' ---> 3' - sense
'utr3_sequence':'',
# REQUIRED!
# name of provided 3`UTR
'utr3_name':'',
# REQUIRED!
# number (integer) of A repeat in the polyA tail
'polya_tail_x':00,
# REQUIRED!
# sequence of provided selection marker
# sequence orientation 5' ---> 3' - sense
'selection_marker_sequence':'',
# REQUIRED!
# name of provided selection marker
'selection_marker_name':'',
# OPTIONAL!
# restriction enzymes protection of transcript sequences
# if the user does not need any restriction places protection, provide empty list []
'restriction_list':[],
# REQUIRED!
# available options (True / False)
# decision; if the user wants the transcription sequences optimized based on the provided species
'optimize':True
# REQUIRED; if optimize == True!
# user-defined percent of GC% content in predicted/optimized sequence
'transcript_GC':58,
# REQUIRED; if optimize == True!
# user-defined maximum number of consecutive repeats of a single nucleotide (A, C, T(U), G) in the predicted/optimized sequence, eg. AAAAAA
'poly_len':7
}
Example dictionary:
input_dict = {
'project_name':'test_invitro_transcription_mRNA',
'vector_type':'transcription',
'vector_function':'mrna',
'species':'human',
'sequences':['ATGGCGATGAGCAGCGGCGGCAGTGGTGGCGGCGTCCCGGAGCAGGAGGATTCCGTGCTGTTCCGGCGCGGCACAGGCCAGAGCGATGATTCTGACATTTGGGATGATACAGCACTGATAAAAGCATATGATAAAGCTGTGGCTTCATTTAAGCATGCTCTAAAGAATGGTGACATTTGTGAAACTTCGGGTAAACCAAAAACCACACCTAAAAGAAAACCTGCTAAGAAGAATAAAAGCCAAAAGAAGAATACTGCAGCTTCCTTACAACAGTGGAAAGTTGGGGACAAATGTTCTGCCATTTGGTCAGAAGACGGTTGCATTTACCCAGCTACCATTGCTTCAATTGATTTTAAGAGAGAAACCTGTGTTGTGGTTTACACTGGATATGGAAATAGAGAGGAGCAAAATCTGTCCGATCTACTTTCCCCAATCTGTGAAGTAGCTAATAATATAGAACAAAATGCTCAAGAGAATGAAAATGAAAGCCAAGTTTCAACAGATGAAAGTGAGAACTCCAGGTCTCCTGGAAATAAATCAGATAACATCAAGCCCAAATCTGCTCCATGGAACTCTTTTCTCCCTCCACCACCCCCCATGCCAGGGCCAAGACTGGGACCAGGAAAGCCAGGTCTAAAATTCAATGGCCCACCACCGCCACCGCCACCACCACCACCCCACTTACTATCATGCTGGCTGCCTCCATTTCCTTCTGGACCACCAATAATTCCCCCACCACCTCCCATATGTCCAGATTCTCTTGATGATGCTGATGCTTTGGGAAGTATGTTAATTTCATGGTACATGAGTGGCTATCATACTGGCTATTATATGTTTCCTGAGGCCTCCCTAAAAGCCGAGCAGATGCCAGCACCATGCTTCCTGTAA'],
'sequences_names':['SMN1'],
'linkers_sequences':[''],
'linkers_names':[''],
'utr5_sequence':'GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGAT',
'utr5_name':'SMN1',
'utr3_sequence':'CAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTA',
'utr3_name':'KIT',
'polya_tail_x':50,
'selection_marker_sequence':'ATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATTTTGCCTTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGTTACATCGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTTTTAAAGTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTCTCAGAATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGTGCTGCCATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTTTTTTGCACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGAGCGTGACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCCCGGCAACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGTTTATTGCTGATAAATCTGGAGCCGGTGAGCGTGGAAGCCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTCCCGTATCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCCTCACTGATTAAGCATTGGTAA',
'selection_marker_name':'Ampicillin',
'restriction_list':['RsaI', 'MnlI', 'AciI', 'AluI', 'BmrI'],
'optimize':True,
'transcript_GC':58,
'poly_len':7
}
Output:
# Name of project
project['project']
# Graph of the designed vector
vector_plot = project['vector']['graph']
vector_plot.savefig('expression_vector.svg')
Example return:

# Complete FASTA file of the designed vecotr
project['vector']['full_fasta']
Example return:
>test_invitro_transcription_mRNA_Regular_plasmid_mrna_3676nc
TAATACGACTCACTATAGGGGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGATGCCACCATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTGTGACAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAATGAAGAGCCGTACGGGCGCGCCTAGGCGCGATTCCGCTTCCTCGCTCACTGACTCGCTGCGCTCGGTCGTTCGGCTGCGGCGAGCGGTATCAGCTCACTCAAAGGCGGTAATACGGTTATCCACAGAATCAGGGGATAACGCAGGAAAGAACATGTGAGCAAAAGGCCAGCAAAAGGCCAGGAACCGTAAAAAGGCCGCGTTGCTGGCGTTTTTCCATAGGCTCCGCCCCCCTGACGAGCATCACAAAAATCGACGCTCAAGTCAGAGGTGGCGAAACCCGACAGGACTATAAAGATACCAGGCGTTTCCCCCTGGAAGCTCCCTCGTGCGCTCTCCTGTTCCGACCCTGCCGCTTACCGGATACCTGTCCGCCTTTCTCCCTTCGGGAAGCGTGGCGCTTTCTCATAGCTCACGCTGTAGGTATCTCAGTTCGGTGTAGGTCGTTCGCTCCAAGCTGGGCTGTGTGCACGAACCCCCCGTTCAGCCCGACCGCTGCGCCTTATCCGGTAACTATCGTCTTGAGTCCAACCCGGTAAGACACGACTTATCGCCACTGGCAGCAGCCACTGGTAACAGGATTAGCAGAGCGAGGTATGTAGGCGGTGCTACAGAGTTCTTGAAGTGGTGGCCTAACTACGGCTACACTAGAAGAACAGTATTTGGTATCTGCGCTCTGCTGAAGCCAGTTACCTTCGGAAAAAGAGTTGGTAGCTCTTGATCCGGCAAACAAACCACCGCTGGTAGCGGTGGTTTTTTTGTTTGCAAGCAGCAGATTACGCGCAGAAAAAAAGGATCTCAAGAAGATCCTTTGATCTTTTCTACGGGGTCTGACGCTCAGTGGAACGAAAACTCACGTTAAGGGATTTTGGTCATGAGATTATCAAAAAGGATCTTCACCTAGATCCTTTTAAATTAAAAATGAAGTTTTAAATCAATCTAAAGTATATATGAGTAAACTTGGTCTGACAGAATGGTTACGAATTAGTCACTCCGTGGATAGAGTCGCTAGACAGATAAAGCAAGTAGGTATCAACGGACTGAGGGGCAGCACATCTATTGATGCTATGCCCTCCCGAATGGTAGACCGGGGTCACGACGTTACTATGGCGCCGAAGGTGCGAGTGGCCGAGGTCTAAATAGTCGTTATTTGGTCGGTCGGCCTTCCCGGCTCGCGTCTTCACCAGGACGTTGAAATAGGCGGAGGTAGGTCAGATAATTAACAACGGCCCTTCGATCTCATTCATCAAGCGGTCAATTATCAAACGCGTTGCAACAACGGTAACGATGTCCGTAGCACCACAGTGCGAGCAGCAAACCATACCGAAGTAAGTCGAGGCCAAGGGTTGCTAGTTCCGCTCAATGTACTAGGGGGTACAACACGTTTTTTCGCCAATCGAGGAAGCCAGGAGGCTAGCAACAGTCTTCATTCAACCGGCGTCACAATAGTGAGTACCAATACCGTCGTGACGTATTAAGAGAATGACAGTACGGTAGGCATTCTACGAAAAGACACTGACCACTCATGAGTTGGTTCAGTAAGACTCTTATCACATACGCCGCTGGCTCAACGAGAACGGGCCGCAGTTATGCCCTATTATGGCGCGGTGTATCGTCTTGAAATTTTCACGAGTAGTAACCTTTTGCAAGAAGCCCCGCTTTTGAGAGTTCCTAGAATGGCGACAACTCTAGGTCAAGCTACATTGGGTGAGCACGTGGGTTGACTAGAAGTCGTAGAAAATGAAAGTGGTCGCAAAGACCCACTCGTTTTTGTCCTTCCGTTTTACGGCGTTTTTTCCCTTATTCCCGCTGTGCCTTTACAACTTATGAGTAACTCTTCCTTTTTCAATATTATTGAAGCATTTATCAGGGTTATTGTCTCATGAGCGGATACATATTTGAATGTATTTAGAAAAATAAACAAATAGGGGTTCCGCGCACATTTCCCCGAAAAGTGCCACCTGACGTCTAAGAAACCATTATTATCATGACATTAACCTATAAAAATAGGCGTATCACGAGGCCCTTTCGTC
# The FASTA file is divided into particular elements of the designed vector
project['vector']['fasta']
Example return:
#test_invitro_transcription_mRNA_Regular_plasmid_mrna_3676nc
>T7
TAATACGACTCACTATAGG
>5`UTR : SMN1
GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGAT
>Kozak_sequence
GCCACC
>SEQ1 : SMN1
ATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTGTGA
>3`UTR : KIT
CAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTA
>PolyA_tail : x 50
AAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA
>backbone_element
T
>SapI
GAAGAGC
>BsiWI
CGTACG
>AscI
GGCGCGCC
>backbone_element
TAGGCGCGATTCCGCTTCCTCGCTCACTGACTCGCTGCGCTCGGTCGTTCGGCTGCGGCGAGCGGTATCAGCTCACTCAAAGGCGGTAATACGGTTATCCACAGAATCAGGGGATAACGCAGGAAAGAACATGTGAGCAAAAGGCCAGCAAAAGGCCAGGAACCGTAAAAAGGCCGCGTTGCTGGCGTT
>pUC_ori
TTTCCATAGGCTCCGCCCCCCTGACGAGCATCACAAAAATCGACGCTCAAGTCAGAGGTGGCGAAACCCGACAGGACTATAAAGATACCAGGCGTTTCCCCCTGGAAGCTCCCTCGTGCGCTCTCCTGTTCCGACCCTGCCGCTTACCGGATACCTGTCCGCCTTTCTCCCTTCGGGAAGCGTGGCGCTTTCTCATAGCTCACGCTGTAGGTATCTCAGTTCGGTGTAGGTCGTTCGCTCCAAGCTGGGCTGTGTGCACGAACCCCCCGTTCAGCCCGACCGCTGCGCCTTATCCGGTAACTATCGTCTTGAGTCCAACCCGGTAAGACACGACTTATCGCCACTGGCAGCAGCCACTGGTAACAGGATTAGCAGAGCGAGGTATGTAGGCGGTGCTACAGAGTTCTTGAAGTGGTGGCCTAACTACGGCTACACTAGAAGAACAGTATTTGGTATCTGCGCTCTGCTGAAGCCAGTTACCTTCGGAAAAAGAGTTGGTAGCTCTTGATCCGGCAAACAAACCACCGCTGGTAGCGGTGGTTTTTTTGTTTGCAAGCAGCAGATTACGCGCAGAAAAAAAGGATCTCAA
>backbone_element
GAAGATCCTTTGATCTTTTCTACGGGGTCTGACGCTCAGTGGAACGAAAACTCACGTTAAGGGATTTTGGTCATGAGATTATCAAAAAGGATCTTCACCTAGATCCTTTTAAATTAAAAATGAAGTTTTAAATCAATCTAAAGTATATATGAGTAAACTTGGTCTGACAG
>Resistance : Ampicillin
AATGGTTACGAATTAGTCACTCCGTGGATAGAGTCGCTAGACAGATAAAGCAAGTAGGTATCAACGGACTGAGGGGCAGCACATCTATTGATGCTATGCCCTCCCGAATGGTAGACCGGGGTCACGACGTTACTATGGCGCCGAAGGTGCGAGTGGCCGAGGTCTAAATAGTCGTTATTTGGTCGGTCGGCCTTCCCGGCTCGCGTCTTCACCAGGACGTTGAAATAGGCGGAGGTAGGTCAGATAATTAACAACGGCCCTTCGATCTCATTCATCAAGCGGTCAATTATCAAACGCGTTGCAACAACGGTAACGATGTCCGTAGCACCACAGTGCGAGCAGCAAACCATACCGAAGTAAGTCGAGGCCAAGGGTTGCTAGTTCCGCTCAATGTACTAGGGGGTACAACACGTTTTTTCGCCAATCGAGGAAGCCAGGAGGCTAGCAACAGTCTTCATTCAACCGGCGTCACAATAGTGAGTACCAATACCGTCGTGACGTATTAAGAGAATGACAGTACGGTAGGCATTCTACGAAAAGACACTGACCACTCATGAGTTGGTTCAGTAAGACTCTTATCACATACGCCGCTGGCTCAACGAGAACGGGCCGCAGTTATGCCCTATTATGGCGCGGTGTATCGTCTTGAAATTTTCACGAGTAGTAACCTTTTGCAAGAAGCCCCGCTTTTGAGAGTTCCTAGAATGGCGACAACTCTAGGTCAAGCTACATTGGGTGAGCACGTGGGTTGACTAGAAGTCGTAGAAAATGAAAGTGGTCGCAAAGACCCACTCGTTTTTGTCCTTCCGTTTTACGGCGTTTTTTCCCTTATTCCCGCTGTGCCTTTACAACTTATGAGTA
>backbone_element
ACTCTTCCTTTTTCAATATTATTGAAGCATTTATCAGGGTTATTGTCTCATGAGCGGATACATATTTGAATGTATTTAGAAAAATAAACAAATAGGGGTTCCGCGCACATTTCCCCGAAAAGTGCCACCTGACGTCTAAGAAACCATTATTATCATGACATTAACCTATAAAAATAGGCGTATCACGAGGCCCTTTCGTC
## genes names
project['transcripts']['sequences']['name']
## proteins sequences
project['transcripts']['sequences']['sequence_aa']
## average codon frequency in the input sequence
project['transcripts']['sequences']['vector_sequence_frequence']
## GC% content in the input sequence
project['transcripts']['sequences']['vector_sequence_GC']
############################################################################
## average codon frequency in the output sequence
project['transcripts']['sequences']['optimized_vector_sequence_frequence']
## GC% content in the output sequence
project['transcripts']['sequences']['optimized_vector_sequence_GC']
Sequences output data:
# Sequences
# Input sequence
project['transcripts']['sequences']['sequence']
# Predicted structure - input
project['transcripts']['sequences']['sequence_figure']
# Optimized sequence
project['transcripts']['sequences']['vector_sequence']
# Predicted structure - optimized
project['transcripts']['sequences']['optimized_sequence_figure']
Example return:
- Otimized sequence structure:

- Input sequence structure:

- Alternative options:
# Sequences results and metadata
# optimized sequence - main
project['transcripts']['sequences']
# optimized sequence - alternatives
project['transcripts']['alternative']
# additional variant 1
project['transcripts']['alternative']['var0]
# additional variant 2
project['transcripts']['alternative']['var1]
Example return:
>Optimized sequence - main
ATGCTGACCAGCGGATTGGTTGTGAGCAACATGTTCAGCTACCACCTGGCCGCCCTGGGATTGATGCCTTCTTTTCAGATGGAGGGCAGGGGCAGAGTGAACCAGTTGGGAGGAGTGTTTATCAACGGCCGGCCTCTGCCTAATCACATCCGGCTGAAAATCGTGGAGTTGGCCGCTCAGGGAGTTAGGCCTTGTGTGATTAGCAGGCAGCTGAGGGTGTCTCACGGATGTGTGAGCAAGATCCTGCAGCGGTACCAGGAGACCGGCTCTATCAGGCCTGGAGTGATTGGCGGCTCTAAACCCAGGGTGGCTACACCTGAAGTGGAAAAGAAGATCGAGCAGTACAAGAAGGACAACCCCGGCATCTTCAGCTGGGAGATCCGGGACCGGCTGCTGAAGGAGGGCATCTGCGACAGAAGCACCGTGCCTTCTGTGTCTAGCATCAGCAGGGTGTTGAGGAGCAGATTTCAGAAGTGCGACAGCGACGACAACGACAACGACAACGACAACGAGGACGACGACGGCGACGACGGCAGCAACAGCAGCGTGGCCGATAGGTCTGTGAACTTCAGCGTGAGCGGCCTGCTGTCTGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGAGCGAGCCCGGACTGTCTGTGAAACGGAAGCAGAGAAGGAGCAGAACAACCTTCACAGCTGAACAGCTGGAGGAACTGGAGAGGGCCTTTGAGAGGACACACTACCCTGATATCTACACCCGGGAAGAGCTGGCTCAGAGAACAAAGTTGACCGAGGCCAGAGTTCAGGTGTGGTTCAGCAACCGGAGGGCCAGATGGAGAAAGCAGATGGGCTCTAACCAGTTGACCGCTCTGAATAGCATCCTGCAGGTGCCCCAGGGAATGGGAACACCTAGCTACATGCTGCACGAGCCCGGATACCCTTTGTCTCACAACGCCGATAACCTGTGGCACAGGTCTAGCATGGCTCAGTCTCTGCAGAGCTTCGGACAGACAATCAAGCCCGAGAACAGCTACGCCGGATTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGACTGCCTACACATAGCAGCAGCGACCCTTTGTCTTCTACCTGGTCTAGCCCTGTGTCTACCTCTGTGCCTGCTCTGGGATATACCCCTTCTAGCGGACACTATCACCACTACAGCGACGTGACCAAGTCTACCCTGCACAGCTACAACGCCCACATCCCCTCTGTGACCAACATGGAGCGGTGCAGCGTGGACGACTCTCTGGTGGCTCTGAGAATGAAAAGCCGGGAGCACAGCGCCGCCCTGTCTTTGATGCAGGTGGCCGATAACAAGATGGCCACCTCTTTCTGA
>Optimized sequence - alternative variant 1
ATGCTGACCAGCGGCCTGGTTGTGAGCAACATGTTCTCTTACCACCTGGCCGCTCTGGGATTAATGCCTTCTTTTCAGATGGAGGGCAGGGGCAGAGTGAATCAGTTAGGAGGAGTGTTTATTAACGGCAGGCCTTTACCTAATCACATTCGGCTGAAAATCGTGGAGTTAGCTGCTCAGGGAGTTAGACCTTGTGTGATCAGCAGGCAGTTGAGAGTGTCTCACGGATGTGTGTCTAAAATCCTGCAGAGATACCAGGAGACAGGCAGCATCAGGCCTGGAGTGATTGGAGGCTCTAAACCCAGAGTGGCTACACCTGAAGTGGAGAAAAAGATCGAGCAATACAAGAAGGACAACCCCGGCATCTTCTCTTGGGAGATCCGGGACAGGTTGCTGAAGGAGGGAATCTGTGATAGGTCTACAGTGCCTTCTGTGTCTAGCATCAGCAGGGTGTTGAGAAGCAGATTCCAGAAATGCGACAGCGACGATAACGACAACGACAACGACAACGAGGACGACGACGGCGATGATGGCTCTAATAGCAGCGTGGCCGATAGATCTGTGAACTTCAGCGTGAGCGGATTGCTGTCTGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGTCTGAGCCTGGACTGTCTGTTAAACGGAAGCAGAGGAGGAGCAGAACAACATTCACAGCCGAGCAGTTGGAGGAACTGGAGAGGGCCTTTGAGAGAACCCACTATCCCGATATCTACACCAGGGAAGAACTGGCTCAGAGAACAAAACTGACCGAGGCCAGAGTTCAGGTGTGGTTTAGCAACCGGAGGGCCAGATGGAGAAAACAGATGGGCAGCAACCAGTTAACAGCCCTGAACAGCATCCTGCAGGTGCCTCAAGGAATGGGAACACCTTCTTATATGCTGCACGAGCCTGGATATCCTTTATCTCATAACGCCGATAACCTGTGGCACAGGTCTTCTATGGCTCAATCTCTGCAGTCTTTCGGACAGACAATTAAGCCCGAGAACTCTTACGCCGGATTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGACTGCCTACACATTCTTCTAGCGACCCTTTATCTTCTACATGGTCTAGCCCTGTGTCTACATCTGTGCCTGCTCTGGGATATACACCTTCTTCTGGCCACTATCACCACTACAGCGACGTGACCAAGAGCACCCTGCATTCTTACAACGCCCACATTCCCAGCGTGACCAACATGGAGCGGTGCTCTGTGGATGACAGCTTAGTGGCTCTGAGAATGAAAAGCAGGGAACACTCTGCTGCTCTGTCTTTAATGCAGGTGGCCGATAATAAGATGGCCACCTCTTTCTGA
>Optimized sequence - alternative variant 2
ATGCTGACCAGCGGATTGGTGGTGAGCAACATGTTCTCTTACCACCTGGCTGCTCTGGGCCTGATGCCCTCTTTCCAGATGGAGGGCCGGGGCAGGGTGAATCAGTTGGGCGGAGTGTTTATCAACGGCCGGCCCCTGCCTAATCACATCCGGCTGAAGATCGTGGAGTTGGCTGCTCAGGGAGTGAGACCCTGCGTGATCAGCCGGCAGTTGAGGGTGTCTCACGGATGTGTGAGCAAGATCCTGCAGAGATACCAGGAGACCGGCAGCATCCGGCCCGGAGTGATTGGCGGCTCTAAGCCCAGGGTGGCTACACCTGAAGTGGAGAAGAAGATCGAGCAGTATAAGAAGGACAACCCCGGCATCTTCTCTTGGGAGATCCGGGACCGGCTGCTGAAGGAGGGCATCTGCGACCGGAGCACCGTGCCCAGCGTGAGCAGCATCAGCCGGGTGCTGCGGAGCCGGTTCCAGAAGTGCGACAGCGACGACAACGACAACGACAACGACAACGAGGACGACGACGGCGACGACGGCAGCAACAGCAGCGTGGCCGACCGGAGCGTGAACTTCAGCGTGAGCGGCCTGCTGAGCGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGAGCGAGCCCGGCCTGAGCGTGAAGCGGAAGCAGCGGCGGAGCCGGACCACCTTCACCGCTGAGCAGTTGGAGGAGTTGGAGCGGGCCTTCGAGCGGACCCACTACCCCGACATCTACACCCGGGAGGAGTTGGCCCAGCGGACCAAGTTGACCGAGGCCCGGGTGCAGGTGTGGTTCAGCAACCGGCGGGCCAGATGGAGGAAACAGATGGGCAGCAACCAGTTGACCGCTCTGAACAGCATCCTGCAGGTGCCCCAGGGCATGGGCACCCCTTCTTACATGCTGCACGAGCCCGGATACCCTTTGAGCCACAACGCCGACAACCTGTGGCACCGGAGCAGCATGGCCCAGAGCCTGCAGTCTTTCGGCCAGACCATCAAGCCCGAGAACTCTTACGCCGGCCTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGCCTGCCCACCCACAGCAGCAGCGACCCCCTGAGCAGCACCTGGAGCAGCCCTGTGTCTACCAGCGTGCCTGCTCTGGGATATACCCCCAGCAGCGGACACTATCACCACTACAGCGACGTGACCAAGAGCACCCTGCACTCTTACAACGCCCACATCCCCAGCGTGACCAACATGGAGCGGTGCAGCGTGGACGACAGCCTGGTGGCCCTGCGGATGAAGAGCCGGGAGCACAGCGCTGCTCTGAGCCTGATGCAGGTGGCCGACAACAAGATGGCCACCTCTTTCTGA
2.2.4 Creating plasmid vector of in-vitro transcription of RNAi
Empty input dictionary schema:
input_dict = {
# REQUIRED!
# name of current project (defined by user)
'project_name':'',
# REQUIRED!
# avaiable of vector types (transcription)
'vector_type':'transcription',
# REQUIRED!
# in this case 'vector_function':'rnai'
'vector_function':'rnai',
# REQUIRED!
# avaiable options (human / mouse / rat / both (mouse + human) / both2 (rat + human) / multi (mouse + rat + human))
# 'both / both2 / multi' - creating vector function adjusted for all species taking into consideration most adjustments for Homo sapiens
'species':'human',
# OPTIONAL!
# sequence of custom RNAi, which can be provided by user
# if provided, then the algorithm of RNAi estimation is off
# if empt '' the algorithm share the best possible RNAi based on 'rnai_gene_name'
# sequence orientation 5' ---> 3' - sense
'rnai_sequence':'',
# REQUIRED; if rnai_sequence is empty!
# length of RNAi (defined by user)
# recomended range: 20–25
'rnai_length':20,
# OPTIONAL; if rnai_sequence is empty!
# overhang sequence of RNAi sequence 3' end (defined by user)
# recomended 'UU'
'overhang_3_prime':'UU',
# REQUIRED!
# name of the target gene for the RNAi searching algorithm (gene name for Homo sapien / Mus musculus / Rattus norvegicus)
# algorithm is working when the rnai_sequence is empty ''
# if the user defines 'rnai_sequence' this 'rnai_gene_name' is just a name for a user-supplied sequence
'rnai_gene_name':'',
# REQUIRED!
# sequence of the loop to create the structure of the hairpin of shRNA or siRNA depending on the loop sequence
# algorithm is working when the rnai_sequence is empty ''
# if the user defines 'rnai_sequence' this 'rnai_gene_name' is just a name for a user-supplied sequence
# sequence orientation 5' ---> 3' - sense
'loop_sequence':'',
# REQUIRED!
# sequence of provided selection marker
# sequence orientation 5' ---> 3' - sense
'selection_marker_sequence':'',
# REQUIRED!
# name of provided selection marker
'selection_marker_name':''
}
Example dictionary:
input_dict = {
'project_name':'test_invitro_transcription_RNAi',
'vector_type':'transcription',
'vector_function':'rnai',
'species':'human',
'rnai_sequence':'',
'rnai_length':20,
'overhang_3_prime':'UU',
'rnai_gene_name':'KIT',
'loop_sequence':'TAGTGAAGCCACAGATGTAC',
'selection_marker_sequence':'ATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATTTTGCCTTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGTTACATCGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTTTTAAAGTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTCTCAGAATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGTGCTGCCATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTTTTTTGCACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGAGCGTGACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCCCGGCAACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGTTTATTGCTGATAAATCTGGAGCCGGTGAGCGTGGAAGCCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTCCCGTATCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCCTCACTGATTAAGCATTGGTAA',
'selection_marker_name':'Ampicillin'
}
Output:
# Name of project
project['project']
# Graph of the designed vector
vector_plot = project['vector']['graph']
vector_plot.savefig('expression_vector.svg')
Example return:

# Complete FASTA file of the designed vecotr
project['vector']['full_fasta']
Example return:
>test_invitro_transcription_mRNA_Regular_plasmid_mrna_3676nc
TAATACGACTCACTATAGGGGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGATGCCACCATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTGTGACAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAATGAAGAGCCGTACGGGCGCGCCTAGGCGCGATTCCGCTTCCTCGCTCACTGACTCGCTGCGCTCGGTCGTTCGGCTGCGGCGAGCGGTATCAGCTCACTCAAAGGCGGTAATACGGTTATCCACAGAATCAGGGGATAACGCAGGAAAGAACATGTGAGCAAAAGGCCAGCAAAAGGCCAGGAACCGTAAAAAGGCCGCGTTGCTGGCGTTTTTCCATAGGCTCCGCCCCCCTGACGAGCATCACAAAAATCGACGCTCAAGTCAGAGGTGGCGAAACCCGACAGGACTATAAAGATACCAGGCGTTTCCCCCTGGAAGCTCCCTCGTGCGCTCTCCTGTTCCGACCCTGCCGCTTACCGGATACCTGTCCGCCTTTCTCCCTTCGGGAAGCGTGGCGCTTTCTCATAGCTCACGCTGTAGGTATCTCAGTTCGGTGTAGGTCGTTCGCTCCAAGCTGGGCTGTGTGCACGAACCCCCCGTTCAGCCCGACCGCTGCGCCTTATCCGGTAACTATCGTCTTGAGTCCAACCCGGTAAGACACGACTTATCGCCACTGGCAGCAGCCACTGGTAACAGGATTAGCAGAGCGAGGTATGTAGGCGGTGCTACAGAGTTCTTGAAGTGGTGGCCTAACTACGGCTACACTAGAAGAACAGTATTTGGTATCTGCGCTCTGCTGAAGCCAGTTACCTTCGGAAAAAGAGTTGGTAGCTCTTGATCCGGCAAACAAACCACCGCTGGTAGCGGTGGTTTTTTTGTTTGCAAGCAGCAGATTACGCGCAGAAAAAAAGGATCTCAAGAAGATCCTTTGATCTTTTCTACGGGGTCTGACGCTCAGTGGAACGAAAACTCACGTTAAGGGATTTTGGTCATGAGATTATCAAAAAGGATCTTCACCTAGATCCTTTTAAATTAAAAATGAAGTTTTAAATCAATCTAAAGTATATATGAGTAAACTTGGTCTGACAGAATGGTTACGAATTAGTCACTCCGTGGATAGAGTCGCTAGACAGATAAAGCAAGTAGGTATCAACGGACTGAGGGGCAGCACATCTATTGATGCTATGCCCTCCCGAATGGTAGACCGGGGTCACGACGTTACTATGGCGCCGAAGGTGCGAGTGGCCGAGGTCTAAATAGTCGTTATTTGGTCGGTCGGCCTTCCCGGCTCGCGTCTTCACCAGGACGTTGAAATAGGCGGAGGTAGGTCAGATAATTAACAACGGCCCTTCGATCTCATTCATCAAGCGGTCAATTATCAAACGCGTTGCAACAACGGTAACGATGTCCGTAGCACCACAGTGCGAGCAGCAAACCATACCGAAGTAAGTCGAGGCCAAGGGTTGCTAGTTCCGCTCAATGTACTAGGGGGTACAACACGTTTTTTCGCCAATCGAGGAAGCCAGGAGGCTAGCAACAGTCTTCATTCAACCGGCGTCACAATAGTGAGTACCAATACCGTCGTGACGTATTAAGAGAATGACAGTACGGTAGGCATTCTACGAAAAGACACTGACCACTCATGAGTTGGTTCAGTAAGACTCTTATCACATACGCCGCTGGCTCAACGAGAACGGGCCGCAGTTATGCCCTATTATGGCGCGGTGTATCGTCTTGAAATTTTCACGAGTAGTAACCTTTTGCAAGAAGCCCCGCTTTTGAGAGTTCCTAGAATGGCGACAACTCTAGGTCAAGCTACATTGGGTGAGCACGTGGGTTGACTAGAAGTCGTAGAAAATGAAAGTGGTCGCAAAGACCCACTCGTTTTTGTCCTTCCGTTTTACGGCGTTTTTTCCCTTATTCCCGCTGTGCCTTTACAACTTATGAGTAACTCTTCCTTTTTCAATATTATTGAAGCATTTATCAGGGTTATTGTCTCATGAGCGGATACATATTTGAATGTATTTAGAAAAATAAACAAATAGGGGTTCCGCGCACATTTCCCCGAAAAGTGCCACCTGACGTCTAAGAAACCATTATTATCATGACATTAACCTATAAAAATAGGCGTATCACGAGGCCCTTTCGTC
# The FASTA file is divided into particular elements of the designed vector
project['vector']['fasta']
Example return:
#test_invitro_transcription_mRNA_Regular_plasmid_mrna_3676nc
>T7
TAATACGACTCACTATAGG
>5`UTR : SMN1
GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGAT
>Kozak_sequence
GCCACC
>SEQ1 : SMN1
ATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTGTGA
>3`UTR : KIT
CAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTA
>PolyA_tail : x 50
AAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAAA
>backbone_element
T
>SapI
GAAGAGC
>BsiWI
CGTACG
>AscI
GGCGCGCC
>backbone_element
TAGGCGCGATTCCGCTTCCTCGCTCACTGACTCGCTGCGCTCGGTCGTTCGGCTGCGGCGAGCGGTATCAGCTCACTCAAAGGCGGTAATACGGTTATCCACAGAATCAGGGGATAACGCAGGAAAGAACATGTGAGCAAAAGGCCAGCAAAAGGCCAGGAACCGTAAAAAGGCCGCGTTGCTGGCGTT
>pUC_ori
TTTCCATAGGCTCCGCCCCCCTGACGAGCATCACAAAAATCGACGCTCAAGTCAGAGGTGGCGAAACCCGACAGGACTATAAAGATACCAGGCGTTTCCCCCTGGAAGCTCCCTCGTGCGCTCTCCTGTTCCGACCCTGCCGCTTACCGGATACCTGTCCGCCTTTCTCCCTTCGGGAAGCGTGGCGCTTTCTCATAGCTCACGCTGTAGGTATCTCAGTTCGGTGTAGGTCGTTCGCTCCAAGCTGGGCTGTGTGCACGAACCCCCCGTTCAGCCCGACCGCTGCGCCTTATCCGGTAACTATCGTCTTGAGTCCAACCCGGTAAGACACGACTTATCGCCACTGGCAGCAGCCACTGGTAACAGGATTAGCAGAGCGAGGTATGTAGGCGGTGCTACAGAGTTCTTGAAGTGGTGGCCTAACTACGGCTACACTAGAAGAACAGTATTTGGTATCTGCGCTCTGCTGAAGCCAGTTACCTTCGGAAAAAGAGTTGGTAGCTCTTGATCCGGCAAACAAACCACCGCTGGTAGCGGTGGTTTTTTTGTTTGCAAGCAGCAGATTACGCGCAGAAAAAAAGGATCTCAA
>backbone_element
GAAGATCCTTTGATCTTTTCTACGGGGTCTGACGCTCAGTGGAACGAAAACTCACGTTAAGGGATTTTGGTCATGAGATTATCAAAAAGGATCTTCACCTAGATCCTTTTAAATTAAAAATGAAGTTTTAAATCAATCTAAAGTATATATGAGTAAACTTGGTCTGACAG
>Resistance : Ampicillin
AATGGTTACGAATTAGTCACTCCGTGGATAGAGTCGCTAGACAGATAAAGCAAGTAGGTATCAACGGACTGAGGGGCAGCACATCTATTGATGCTATGCCCTCCCGAATGGTAGACCGGGGTCACGACGTTACTATGGCGCCGAAGGTGCGAGTGGCCGAGGTCTAAATAGTCGTTATTTGGTCGGTCGGCCTTCCCGGCTCGCGTCTTCACCAGGACGTTGAAATAGGCGGAGGTAGGTCAGATAATTAACAACGGCCCTTCGATCTCATTCATCAAGCGGTCAATTATCAAACGCGTTGCAACAACGGTAACGATGTCCGTAGCACCACAGTGCGAGCAGCAAACCATACCGAAGTAAGTCGAGGCCAAGGGTTGCTAGTTCCGCTCAATGTACTAGGGGGTACAACACGTTTTTTCGCCAATCGAGGAAGCCAGGAGGCTAGCAACAGTCTTCATTCAACCGGCGTCACAATAGTGAGTACCAATACCGTCGTGACGTATTAAGAGAATGACAGTACGGTAGGCATTCTACGAAAAGACACTGACCACTCATGAGTTGGTTCAGTAAGACTCTTATCACATACGCCGCTGGCTCAACGAGAACGGGCCGCAGTTATGCCCTATTATGGCGCGGTGTATCGTCTTGAAATTTTCACGAGTAGTAACCTTTTGCAAGAAGCCCCGCTTTTGAGAGTTCCTAGAATGGCGACAACTCTAGGTCAAGCTACATTGGGTGAGCACGTGGGTTGACTAGAAGTCGTAGAAAATGAAAGTGGTCGCAAAGACCCACTCGTTTTTGTCCTTCCGTTTTACGGCGTTTTTTCCCTTATTCCCGCTGTGCCTTTACAACTTATGAGTA
>backbone_element
ACTCTTCCTTTTTCAATATTATTGAAGCATTTATCAGGGTTATTGTCTCATGAGCGGATACATATTTGAATGTATTTAGAAAAATAAACAAATAGGGGTTCCGCGCACATTTCCCCGAAAAGTGCCACCTGACGTCTAAGAAACCATTATTATCATGACATTAACCTATAAAAATAGGCGTATCACGAGGCCCTTTCGTC
# Top 1 designed RNAi in shRNA form information
# RNAi name
project['rnai']['name']
# RNAi sequence
project['rnai']['sequence']
# RNAi prediction structure
rnai_prediction = project['rnai']['figure']
rnai_prediction.savefig('rnai_predicted_structure.svg')
Example return:

# other selected RNAi with statistics
pd.DataFrame(project['rnai']['full_data'])
Examples with structure description presented in:
- 1.23. Prediction of RNAi on the provided sequence
- 1.24. Correcting of RNAi_data for complementarity to the loop sequence
- 1.25. Correcting of RNAi_data for complementarity to the additional external sequence
2.3. Creating sequence for synthesis de novo
project = vb.create_sequence_from_dict(metadata, input_dict, show_plot=True)
This function change provided by user metadata into two types of predicted sequences:
-expression (artificial gene - mRNA)
-RNAi (silencing - siRNA/shRNA)
Args:
metadata (dict) - matadata loaded in the load_metadata() function
input_dict (dict) - dictionary of metadata provided by the user
Examples:
-expression (mRNA)
-RNAi (siRNA/shRNA)
Avaiable on https://github.com/jkubis96/JBioSeqTools
If you have any problem, don't hesitate to contact us!
Args
show_plot (bool) - if True the plot will be displayed, if False only the graph will be returned to the project. Default: True
Returns:
dict: Dictionary including all vector data (graphs, sequences, fasta) created based on user definition
2.3.1 Creating sequence prediction of mRNA for de novo synthesis
Empty input dictionary schema:
input_dict = {
# REQUIRED!
# name of current project (defined by user)
'project_name':'',
# REQUIRED!
# avaiable options (human / mouse / rat / both (mouse + human) / both2 (rat + human) / multi (mouse + rat + human))
# 'both / both2 / multi' - creating vector function adjusted for all species taking into consideration most adjustments for Homo sapiens
'species':'',
# REQUIRED!
# string of coding sequence (CDS) provided to optimize sequence
# sequences orientation 5' ---> 3' - sense
'sequences':[''],
# REQUIRED!
# name of coding sequence
'sequence_name':'',
# OPTIONAL!
# restriction enzymes protection of transcript sequence
# if the user does not need any restriction places protection, provide empty list []
'restriction_list':[],
# REQUIRED!
# available options (True / False)
# decision; if the user wants the transcription sequence optimized based on the provided species
'optimize':True,
# REQUIRED; if optimize == True!
# user-defined percent of GC% content in predicted/optimized sequence
'transcript_GC':58,
# REQUIRED; if optimize == True!
# user-defined maximum number of consecutive repeats of a single nucleotide (A, C, T(U), G) in the predicted/optimized sequence, eg. AAAAAA
'poly_len':7
}
Example dictionary:
input_dict = {
'project_name':'test_mRNA',
'species':'human',
'sequence':'ATGTTAACTAGTGGACTTGTAGTAAGCAACATGTTCTCCTATCATCTCGCAGCCTTGGGACTCATGCCGTCATTCCAGATGGAAGGGCGAGGTCGAGTTAATCAGCTAGGAGGTGTTTTCATTAATGGACGGCCACTGCCCAACCATATACGACTGAAGATTGTCGAGCTAGCGGCCCAGGGCGTCCGTCCGTGCGTCATCAGTAGACAGCTGCGGGTGTCACATGGCTGTGTCAGTAAAATACTCCAACGATATCAAGAAACCGGAAGTATCCGACCTGGGGTTATTGGCGGAAGTAAACCAAGGGTCGCAACTCCGGAAGTTGAGAAAAAGATAGAACAATACAAAAAAGATAATCCGGGAATTTTCAGTTGGGAGATTCGGGATCGGCTGCTGAAGGAGGGGATTTGTGACCGCAGCACCGTGCCAAGTGTGAGCTCCATCAGTCGAGTATTACGGAGCAGGTTCCAGAAATGTGATTCTGATGACAATGACAATGACAATGACAATGAGGACGACGATGGCGATGACGGCAGTAACAGTAGTGTGGCAGACAGGTCTGTTAACTTCTCTGTCAGCGGTCTGCTGTCCGACAATAAAAGCGACAAAAGCGACAACGATTCCGATTGTGAATCAGAGCCGGGGCTATCTGTAAAACGGAAGCAACGCCGCAGTCGAACTACTTTCACCGCGGAGCAGTTGGAGGAACTGGAAAGAGCCTTTGAACGAACTCACTATCCGGATATATATACGCGAGAGGAATTAGCACAAAGAACAAAGCTAACCGAGGCAAGAGTCCAAGTATGGTTTAGTAACCGAAGAGCGAGATGGCGGAAACAGATGGGTAGCAATCAGCTGACAGCCTTGAACAGTATATTACAAGTGCCACAGGGTATGGGAACGCCCTCTTATATGCTGCACGAGCCTGGGTATCCACTCTCACATAATGCAGACAATCTTTGGCATAGATCGTCTATGGCCCAGTCATTACAGTCATTTGGTCAGACAATAAAACCAGAGAATTCCTACGCCGGTCTTATGGAAAACTATTTATCTCATTCATCACAGCTTCATGGTCTTCCTACACATAGTTCATCCGATCCCCTCTCATCCACTTGGTCATCTCCCGTGTCCACTTCCGTTCCTGCGCTAGGATACACGCCATCTAGTGGCCATTACCATCATTACTCTGATGTCACCAAAAGTACTCTTCATTCATATAACGCTCATATTCCTTCAGTCACAAACATGGAGAGATGTTCAGTTGATGACAGTTTGGTTGCTTTACGTATGAAGTCACGTGAGCATTCCGCCGCTCTCAGTTTGATGCAGGTGGCAGACAACAAAATGGCTACCTCATTTTGA',
'sequence_name':'SMN1',
'restriction_list':['RsaI', 'MnlI', 'AciI', 'AluI', 'BmrI'],
'optimize':True,
'transcript_GC':58,
'poly_len':7
}
Output:
# Name of project
project['project']
# Sequences
# Input sequence
project['transcripts']['sequences']['sequence']
# Predicted structure - input
project['transcripts']['sequences']['sequence_figure']
# Optimized sequence
project['transcripts']['sequences']['optimized_sequence']
# Predicted structure - optimized
project['transcripts']['sequences']['optimized_sequence_figure']
Example return:
- Otimized sequence structure:

- Input sequence structure:

- Alternative options:
# Sequences results and metadata
# optimized sequence - main
project['transcripts']['sequences']
# optimized sequence - alternatives
project['transcripts']['alternative']
# additional variant 1
project['transcripts']['alternative']['var0]
# additional variant 2
project['transcripts']['alternative']['var1]
Example return:
>Optimized sequence - main
ATGCTGACCAGCGGATTGGTTGTGAGCAACATGTTCAGCTACCACCTGGCCGCCCTGGGATTGATGCCTTCTTTTCAGATGGAGGGCAGGGGCAGAGTGAACCAGTTGGGAGGAGTGTTTATCAACGGCCGGCCTCTGCCTAATCACATCCGGCTGAAAATCGTGGAGTTGGCCGCTCAGGGAGTTAGGCCTTGTGTGATTAGCAGGCAGCTGAGGGTGTCTCACGGATGTGTGAGCAAGATCCTGCAGCGGTACCAGGAGACCGGCTCTATCAGGCCTGGAGTGATTGGCGGCTCTAAACCCAGGGTGGCTACACCTGAAGTGGAAAAGAAGATCGAGCAGTACAAGAAGGACAACCCCGGCATCTTCAGCTGGGAGATCCGGGACCGGCTGCTGAAGGAGGGCATCTGCGACAGAAGCACCGTGCCTTCTGTGTCTAGCATCAGCAGGGTGTTGAGGAGCAGATTTCAGAAGTGCGACAGCGACGACAACGACAACGACAACGACAACGAGGACGACGACGGCGACGACGGCAGCAACAGCAGCGTGGCCGATAGGTCTGTGAACTTCAGCGTGAGCGGCCTGCTGTCTGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGAGCGAGCCCGGACTGTCTGTGAAACGGAAGCAGAGAAGGAGCAGAACAACCTTCACAGCTGAACAGCTGGAGGAACTGGAGAGGGCCTTTGAGAGGACACACTACCCTGATATCTACACCCGGGAAGAGCTGGCTCAGAGAACAAAGTTGACCGAGGCCAGAGTTCAGGTGTGGTTCAGCAACCGGAGGGCCAGATGGAGAAAGCAGATGGGCTCTAACCAGTTGACCGCTCTGAATAGCATCCTGCAGGTGCCCCAGGGAATGGGAACACCTAGCTACATGCTGCACGAGCCCGGATACCCTTTGTCTCACAACGCCGATAACCTGTGGCACAGGTCTAGCATGGCTCAGTCTCTGCAGAGCTTCGGACAGACAATCAAGCCCGAGAACAGCTACGCCGGATTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGACTGCCTACACATAGCAGCAGCGACCCTTTGTCTTCTACCTGGTCTAGCCCTGTGTCTACCTCTGTGCCTGCTCTGGGATATACCCCTTCTAGCGGACACTATCACCACTACAGCGACGTGACCAAGTCTACCCTGCACAGCTACAACGCCCACATCCCCTCTGTGACCAACATGGAGCGGTGCAGCGTGGACGACTCTCTGGTGGCTCTGAGAATGAAAAGCCGGGAGCACAGCGCCGCCCTGTCTTTGATGCAGGTGGCCGATAACAAGATGGCCACCTCTTTCTGA
>Optimized sequence - alternative variant 1
ATGCTGACCAGCGGCCTGGTTGTGAGCAACATGTTCTCTTACCACCTGGCCGCTCTGGGATTAATGCCTTCTTTTCAGATGGAGGGCAGGGGCAGAGTGAATCAGTTAGGAGGAGTGTTTATTAACGGCAGGCCTTTACCTAATCACATTCGGCTGAAAATCGTGGAGTTAGCTGCTCAGGGAGTTAGACCTTGTGTGATCAGCAGGCAGTTGAGAGTGTCTCACGGATGTGTGTCTAAAATCCTGCAGAGATACCAGGAGACAGGCAGCATCAGGCCTGGAGTGATTGGAGGCTCTAAACCCAGAGTGGCTACACCTGAAGTGGAGAAAAAGATCGAGCAATACAAGAAGGACAACCCCGGCATCTTCTCTTGGGAGATCCGGGACAGGTTGCTGAAGGAGGGAATCTGTGATAGGTCTACAGTGCCTTCTGTGTCTAGCATCAGCAGGGTGTTGAGAAGCAGATTCCAGAAATGCGACAGCGACGATAACGACAACGACAACGACAACGAGGACGACGACGGCGATGATGGCTCTAATAGCAGCGTGGCCGATAGATCTGTGAACTTCAGCGTGAGCGGATTGCTGTCTGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGTCTGAGCCTGGACTGTCTGTTAAACGGAAGCAGAGGAGGAGCAGAACAACATTCACAGCCGAGCAGTTGGAGGAACTGGAGAGGGCCTTTGAGAGAACCCACTATCCCGATATCTACACCAGGGAAGAACTGGCTCAGAGAACAAAACTGACCGAGGCCAGAGTTCAGGTGTGGTTTAGCAACCGGAGGGCCAGATGGAGAAAACAGATGGGCAGCAACCAGTTAACAGCCCTGAACAGCATCCTGCAGGTGCCTCAAGGAATGGGAACACCTTCTTATATGCTGCACGAGCCTGGATATCCTTTATCTCATAACGCCGATAACCTGTGGCACAGGTCTTCTATGGCTCAATCTCTGCAGTCTTTCGGACAGACAATTAAGCCCGAGAACTCTTACGCCGGATTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGACTGCCTACACATTCTTCTAGCGACCCTTTATCTTCTACATGGTCTAGCCCTGTGTCTACATCTGTGCCTGCTCTGGGATATACACCTTCTTCTGGCCACTATCACCACTACAGCGACGTGACCAAGAGCACCCTGCATTCTTACAACGCCCACATTCCCAGCGTGACCAACATGGAGCGGTGCTCTGTGGATGACAGCTTAGTGGCTCTGAGAATGAAAAGCAGGGAACACTCTGCTGCTCTGTCTTTAATGCAGGTGGCCGATAATAAGATGGCCACCTCTTTCTGA
>Optimized sequence - alternative variant 2
ATGCTGACCAGCGGATTGGTGGTGAGCAACATGTTCTCTTACCACCTGGCTGCTCTGGGCCTGATGCCCTCTTTCCAGATGGAGGGCCGGGGCAGGGTGAATCAGTTGGGCGGAGTGTTTATCAACGGCCGGCCCCTGCCTAATCACATCCGGCTGAAGATCGTGGAGTTGGCTGCTCAGGGAGTGAGACCCTGCGTGATCAGCCGGCAGTTGAGGGTGTCTCACGGATGTGTGAGCAAGATCCTGCAGAGATACCAGGAGACCGGCAGCATCCGGCCCGGAGTGATTGGCGGCTCTAAGCCCAGGGTGGCTACACCTGAAGTGGAGAAGAAGATCGAGCAGTATAAGAAGGACAACCCCGGCATCTTCTCTTGGGAGATCCGGGACCGGCTGCTGAAGGAGGGCATCTGCGACCGGAGCACCGTGCCCAGCGTGAGCAGCATCAGCCGGGTGCTGCGGAGCCGGTTCCAGAAGTGCGACAGCGACGACAACGACAACGACAACGACAACGAGGACGACGACGGCGACGACGGCAGCAACAGCAGCGTGGCCGACCGGAGCGTGAACTTCAGCGTGAGCGGCCTGCTGAGCGACAACAAGAGCGACAAGAGCGACAACGACAGCGACTGCGAGAGCGAGCCCGGCCTGAGCGTGAAGCGGAAGCAGCGGCGGAGCCGGACCACCTTCACCGCTGAGCAGTTGGAGGAGTTGGAGCGGGCCTTCGAGCGGACCCACTACCCCGACATCTACACCCGGGAGGAGTTGGCCCAGCGGACCAAGTTGACCGAGGCCCGGGTGCAGGTGTGGTTCAGCAACCGGCGGGCCAGATGGAGGAAACAGATGGGCAGCAACCAGTTGACCGCTCTGAACAGCATCCTGCAGGTGCCCCAGGGCATGGGCACCCCTTCTTACATGCTGCACGAGCCCGGATACCCTTTGAGCCACAACGCCGACAACCTGTGGCACCGGAGCAGCATGGCCCAGAGCCTGCAGTCTTTCGGCCAGACCATCAAGCCCGAGAACTCTTACGCCGGCCTGATGGAGAACTACCTGAGCCACAGCAGCCAGTTGCACGGCCTGCCCACCCACAGCAGCAGCGACCCCCTGAGCAGCACCTGGAGCAGCCCTGTGTCTACCAGCGTGCCTGCTCTGGGATATACCCCCAGCAGCGGACACTATCACCACTACAGCGACGTGACCAAGAGCACCCTGCACTCTTACAACGCCCACATCCCCAGCGTGACCAACATGGAGCGGTGCAGCGTGGACGACAGCCTGGTGGCCCTGCGGATGAAGAGCCGGGAGCACAGCGCTGCTCTGAGCCTGATGCAGGTGGCCGACAACAAGATGGCCACCTCTTTCTGA
2.3.2 Creating plasmid vector of in-vitro transcription of RNAi
Empty input dictionary schema:
input_dict = {
# REQUIRED!
# name of current project (defined by user)
'project_name':'',
# REQUIRED!
# avaiable options (human / mouse / rat / both (mouse + human) / both2 (rat + human) / multi (mouse + rat + human))
# 'both / both2 / multi' - creating vector function adjusted for all species taking into consideration most adjustments for Homo sapiens
'species':'human',
# REQUIRED!
# type of RNAi (defined by user)
# avaiable options 'sh' - for shRNA and 'sirna' for siRNA
'rnai_type':'',
# OPTIONAL!
# sequence of custom RNAi, which can be provided by user
# if provided, then the algorithm of RNAi estimation is off
# if empt '' the algorithm share the best possible RNAi based on 'rnai_gene_name'
# sequence orientation 5' ---> 3' - sense
'rnai_sequence':'',
# REQUIRED; if rnai_sequence is empty!
# length of RNAi (defined by user)
# recomended range: 20–25
'rnai_length':20,
# OPTIONAL; if rnai_sequence is empty!
# overhang sequence of RNAi sequence 3' end (defined by user)
# recomended 'UU'
'overhang_3_prime':'UU',
# REQUIRED!
# name of the target gene for the RNAi searching algorithm (gene name for Homo sapien / Mus musculus / Rattus norvegicus)
# algorithm is working when the rnai_sequence is empty ''
# if the user defines 'rnai_sequence' this 'rnai_gene_name' is just a name for a user-supplied sequence
'rnai_gene_name':'',
# REQUIRED, if rnai_type = 'sh'!
# sequence of the loop to create the structure of the hairpin of shRNA
# sequence orientation 5' ---> 3' - sense
'loop_sequence':''
}
Example dictionary:
# siRNA
input_dict = {
'project_name':'test_siRNA',
'species':'human',
'rnai_type':'sirna',
'rnai_sequence':'',
'rnai_length':20,
'overhang_3_prime':'UU',
'rnai_gene_name':'KIT',
'loop_sequence':''
}
# shRNA
input_dict = {
'project_name':'test_shRNA',
'species':'human',
'rnai_type':'sh',
'rnai_sequence':'',
'rnai_length':20,
'overhang_3_prime':'UU',
'rnai_gene_name':'KIT',
'loop_sequence':'TAGTGAAGCCACAGATGTAC'
}
Output:
# Name of project
project['project']
# Top 1 designed RNAi
# RNAi name
project['rnai']['name']
# RNAi sequence
project['rnai']['sequence']
# RNAi prediction structure
rnai_prediction = project['rnai']['figure']
rnai_prediction.savefig('rnai_predicted_structure.svg')
Example return:
- shRNA

- siRNA

# other selected RNAi with statistics
pd.DataFrame(project['rnai']['full_data'])
Examples with structure description presented in:
- 1.23. Prediction of RNAi on the provided sequence
- 1.24. Correcting of RNAi_data for complementarity to the loop sequence
2.4. Creating vector plasmid from FASTA - display existing or custom editing FASTA file
FASTA stucture for prepare custom vector
>name1
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACC
>name2
TCTAGACAACTTTGTATAGAAAAGTTG
>name3
GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTAC
Header explanation:
name1,2,3,... - the name of the sequence element
- FASTA file can be prepared in any text editor without any additional tools. The user must only remember that the file extension should be *.fasta!
- User can also modify previously obtained plasmid vector fasta file obtained in vector_create_on_dict() function. Read above!
2.4.1 Loading fasta from the file
fasta_string = vb.load_fasta(path)
This function finds and removes restriction places inside the sequence.
Args:
path (str) - path to the FASTA file *.FASTA
Returns:
str: Loaded FASTA file to the string object
2.4.2 Converting the FASTA string to the data frame
df_fasta = vb.decode_fasta_to_dataframe(fasta_string)
This function decodes the FASTA file from the string to the data frame
Args:
fasta_file (str) - FASTA file (string) loaded by load_fasta() from external source
Returns:
DataFrame: Data frame containing in separate columns FASTA headers and sequences
2.4.3 Decoding FASTA information
df_fasta = vb.extract_fasta_info(df_fasta)
This function extracts the necessary information from headers for vector plotting using plot_vector()
For decoding headers the FASTA structure should be:
>name1
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACC
>name2
TCTAGACAACTTTGTATAGAAAAGTTG
>name3
GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTAC
Header explanation:
name1,2,3,... - the name of the sequence element
Args:
df_fasta (DataFrame) - FASTA data frame obtained from decode_fasta_to_dataframe()
ommit_pattern (str) - string pattern. Any header containing this pattern will be marked as
False for visibility, meaning that the corresponding part of the plasmid
vector sequence will not be displayed.
Returns:
DataFrame: Data frame with additional columns of decoded headers for plot_vector() function
2.4.4 Creating graph of the plasmid vector
graph = vb.plot_vector(df_fasta, title = None, title_size = 20, show_plot = True)
This function displays a plot of the vector plasmid provided in the DataFrame of the FASTA file derived from load_fasta(path) -> decode_fasta_to_dataframe(fasta) -> extract_fasta_info(df_fasta) pipeline.
Args:
df_fasta (DataFrame) - dataframe obtained from load_fasta(path) -> decode fasta to_dataframe(fasta) -> extract_fasta_info(df_fasta) pipeline which prepare decoded FASTA file of vector plasmid.
*dedicated FASTA structure:
>name1
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACC
>name2
TCTAGACAACTTTGTATAGAAAAGTTG
>name3
GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTAC
Header explanation:
name1,2,3,... - the name of the sequence element
title (str | None) - a title that will display in the middle of the plasmid vector. Default: None
title_size (int | float) - font size of the title. Default: 20
show_plot (bool) - if True the plot will be displayed, if False only the graph will be returned to the variable. Default: True
Returns:
Graph: The vector plot based on the provided DataFrame FASTA data
Full pipeline example:
# Exaple FASTA file is loaded from the library repository
import pkg_resources
from jbst import vector_build as vb
fasta_string = vb.load_fasta(pkg_resources.resource_filename("jbst", "tests/fasta_vector_test.fasta"))
Example FASTA file content:
# test_expression_ssAAV_expression_8717nc
>5`ITR
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTCAGTGAGCGAGCGAGCGCGCAGAGAGGGAGTGGCCAACTCCATCACTAGGGGTTCCT
>backbone_element
TCTAGACAACTTTGTATAGAAAAGTTG
>Promoter:TBG
GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGAT
>backbone_element
CAAGTTTGTACAAAAAAGCAGGCT
>Kozak_sequence
GCCACC
>SEQ1:SMN1
ATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTG
>SEQ2:SMN2
ATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTGTGA
>PolyA_signal:SV40_late
CAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTA
>2nd_promoter:CAG
CTCGACATTGATTATTGACTAGTTATTAATAGTAATCAATTACGGGGTCATTAGTTCATAGCCCATATATGGAGTTCCGCGTTACATAACTTACGGTAAATGGCCCGCCTGGCTGACCGCCCAACGACCCCCGCCCATTGACGTCAATAATGACGTATGTTCCCATAGTAACGCCAATAGGGACTTTCCATTGACGTCAATGGGTGGAGTATTTACGGTAAACTGCCCACTTGGCAGTACATCAAGTGTATCATATGCCAAGTACGCCCCCTATTGACGTCAATGACGGTAAATGGCCCGCCTGGCATTATGCCCAGTACATGACCTTATGGGACTTTCCTACTTGGCAGTACATCTACGTATTAGTCATCGCTATTACCATGGTCGAGGTGAGCCCCACGTTCTGCTTCACTCTCCCCATCTCCCCCCCCTCCCCACCCCCAATTTTGTATTTATTTATTTTTTAATTATTTTGTGCAGCGATGGGGGCGGGGGGGGGGGGGGGGCGCGCGCCAGGCGGGGCGGGGCGGGGCGAGGGGCGGGGCGGGGCGAGGCGGAGAGGTGCGGCGGCAGCCAATCAGAGCGGCGCGCTCCGAAAGTTTCCTTTTATGGCGAGGCGGCGGCGGCGGCGGCCCTATAAAAAGCGAAGCGCGCGGCGGGCGGGAGTCGCTGCGCGCTGCCTTCGCCCCGTGCCCCGCTCCGCCGCCGCCTCGCGCCGCCCGCCCCGGCTCTGACTGACCGCGTTACTCCCACAGGTGAGCGGGCGGGACGGCCCTTCTCCTCCGGGCTGTAATTAGCGCTTGGTTTAATGACGGCTTGTTTCTTTTCTGTGGCTGCGTGAAAGCCTTGAGGGGCTCCGGGAGGGCCCTTTGTGCGGGGGGAGCGGCTCGGGGGGTGCGTGCGTGTGTGTGTGCGTGGGGAGCGCCGCGTGCGGCTCCGCGCTGCCCGGCGGCTGTGAGCGCTGCGGGCGCGGCGCGGGGCTTTGTGCGCTCCGCAGTGTGCGCGAGGGGAGCGCGGCCGGGGGCGGTGCCCCGCGGTGCGGGGGGGGCTGCGAGGGGAACAAAGGCTGCGTGCGGGGTGTGTGCGTGGGGGGGTGAGCAGGGGGTGTGGGCGCGTCGGTCGGGCTGCAACCCCCCCTGCACCCCCCTCCCCGAGTTGCTGAGCACGGCCCGGCTTCGGGTGCGGGGCTCCGTACGGGGCGTGGCGCGGGGCTCGCCGTGCCGGGCGGGGGGTGGCGGCAGGTGGGGGTGCCGGGCGGGGCGGGGCCGCCTCGGGCCGGGGAGGGCTCGGGGGAGGGGCGCGGCGGCCCCCGGAGCGCCGGCGGCTGTCGAGGCGCGGCGAGCCGCAGCCATTGCCTTTTATGGTAATCGTGCGAGAGGGCGCAGGGACTTCCTTTGTCCCAAATCTGTGCGGAGCCGAAATCTGGGAGGCGCCGCCGCACCCCCTCTAGCGGGCGCGGGGCGAAGCGGTGCGGCGCCGGCAGGAAGGAAATGGGCGGGGAGGGCCTTCGTGCGTCGCCGCGCCGCCGTCCCCTTCTCCCTCTCCAGCCTCGGGGCTGTCCGCGGGGGGACGGCTGCCTTCGGGGGGGACGGGGCAGGGCGGGGTTCGGCTTCTGGCGTGTGACCGGCGGCTCTAGAGCCTCTGCTAACCATGTTCATGCCTTCTTCTTTTTCCTACAGCTCCTGGGCAACGTGCTGGTTATTGTGCTGTCTCATCATTTTGGCAAAGAATTG
>Fluorescent_tag:EGFP
ATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGGGCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCATCTGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGACCTACGGCGTGCAGTGCTTCAGCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTCTTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGGACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCATGGCCGACAAGCAGAAGAACGGCATCAAGGTGAACTTCAAGATCCGCCACAACATCGAGGACGGCAGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCACTACCTGAGCACCCAGTCCGCCCTGAGCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCGTGACCGCCGCCGGGATCACTCTCGGCATGGACGAGCTGTACAAGTAA
>backbone_element
ACCCAGCTTTCTTGTACAAAGTGGGAATTC
>Enhancer:WPRE
CGATAATCAACCTCTGGATTACAAAATTTGTGAAAGATTGACTGGTATTCTTAACTATGTTGCTCCTTTTACGCTATGTGGATACGCTGCTTTAATGCCTTTGTATCATGCTATTGCTTCCCGTATGGCTTTCATTTTCTCCTCCTTGTATAAATCCTGGTTGCTGTCTCTTTATGAGGAGTTGTGGCCCGTTGTCAGGCAACGTGGCGTGGTGTGCACTGTGTTTGCTGACGCAACCCCCACTGGTTGGGGCATTGCCACCACCTGTCAGCTCCTTTCCGGGACTTTCGCTTTCCCCCTCCCTATTGCCACGGCGGAACTCATCGCCGCCTGCCTTGCCCGCTGCTGGACAGGGGCTCGGCTGTTGGGCACTGACAATTCCGTGGTGTTGTCGGGGAAGCTGACGTCCTTTCCATGGCTGCTCGCCTGTGTTGCCACCTGGATTCTGCGCGGGACGTCCTTCTGCTACGTCCCTTCGGCCCTCAATCCAGCGGACCTTCCTTCCCGCGGCCTGCTGCCGGCTCTGCGGCCTCTTCCGCGTCTTCGCCTTCGCCCTCAGACGAGTCGGATCTCCCTTTGGGCCGCCTCCCCGCATCGG
>backbone_element
GAATTCCTAGAGCTCGCTGATCAGCCTCGA
>2nd_polyA_signal:bGH
CTGTGCCTTCTAGTTGCCAGCCATCTGTTGTTTGCCCCTCCCCCGTGCCTTCCTTGACCCTGGAAGGTGCCACTCCCACTGTCCTTTCCTAATAAAATGAGGAAATTGCATCGCATTGTCTGAGTAGGTGTCATTCTATTCTGGGGGGTGGGGTGGGGCAGGACAGCAAGGGGGAGGATTGGGAAGAGAATAGCAGGCATGCTGGGGA
>backbone_element
GGGCCGC
>3`ITR
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTCAGTGAGCGAGCGAGCGCGCAGAGAGGGAGTGGCCAACTCCATCACTAGGGGTTCCT
>backbone_element
CTGCCTGCAGGGGCGCCTGATGCGGTATTTTCTCCTTACGCATCTGTGCGGTATTTCACACCGCATACGTCAAAGCAACCATAGTACGCGCCCTGTAGCGGCGCATTAAGCGCGGCGGGGGTGGTGGTTACGCGCAGCGTGACCGCTACACTTGCCAGCGCCTTAGCGCCCGCTCCTTTCGCTTTCTTCCCTTCCTTTCTCGCCACGTTCGCCGGCTTTCCCCGTCAAGCTCTAAATCGGGGGCTCCCTTTAGGGTTCCGATTTAGTGCTTTACGGCACCTCGACCCCAAAAAACTTGATTTGGGTGATGGTTCACGTAGTGGGCCATCGCCCTGATAGACGGTTTTTCGCCCTTTGACGTTGGAGTCCACGTTCTTTAATAGTGGACTCTTGTTCCAAACTGGAACAACACTCAACTCTATCTCGGGCTATTCTTTTGATTTATAAGGGATTTTGCCGATTTCGGTCTATTGGTTAAAAAATGAGCTGATTTAACAAAAATTTAACGCGAATTTTAACAAAATATTAACGTTTACAATTTTATGGTGCACTCTCAGTACAATCTGCTCTGATGCCGCATAGTTAAGCCAGCCCCGACACCCGCCAACACCCGCTGACGCGCCCTGACGGGCTTGTCTGCTCCCGGCATCCGCTTACAGACAAGCTGTGACCGTCTCCGGGAGCTGCATGTGTCAGAGGTTTTCACCGTCATCACCGAAACGCGCGAGACGAAAGGGCCTCGTGATACGCCTATTTTTATAGGTTAATGTCATGATAATAATGGTTTCTTAGACGTCAGGTGGCACTTTTCGGGGAAATGTGCGCGGAACCCCTATTTGTTTATTTTTCTAAATACATTCAAATATGTATCCGCTCATGAGACAATAACCCTGATAAATGCTTCAATAATATTGAAAAAGGAAGAGT
>Resistance:Ampicillin
ATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATTTTGCCTTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGTTACATCGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTTTTAAAGTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTCTCAGAATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGTGCTGCCATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTTTTTTGCACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGAGCGTGACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCCCGGCAACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGTTTATTGCTGATAAATCTGGAGCCGGTGAGCGTGGAAGCCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTCCCGTATCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCCTCACTGATTAAGCATTGGTAA
>backbone_element
CTGTCAGACCAAGTTTACTCATATATACTTTAGATTGATTTAAAACTTCATTTTTAATTTAAAAGGATCTAGGTGAAGATCCTTTTTGATAATCTCATGACCAAAATCCCTTAACGTGAGTTTTCGTTCCACTGAGCGTCAGACCCCGTAGAAAAGATCAAAGGATCTTC
>pUC_ori
TTGAGATCCTTTTTTTCTGCGCGTAATCTGCTGCTTGCAAACAAAAAAACCACCGCTACCAGCGGTGGTTTGTTTGCCGGATCAAGAGCTACCAACTCTTTTTCCGAAGGTAACTGGCTTCAGCAGAGCGCAGATACCAAATACTGTTCTTCTAGTGTAGCCGTAGTTAGGCCACCACTTCAAGAACTCTGTAGCACCGCCTACATACCTCGCTCTGCTAATCCTGTTACCAGTGGCTGCTGCCAGTGGCGATAAGTCGTGTCTTACCGGGTTGGACTCAAGACGATAGTTACCGGATAAGGCGCAGCGGTCGGGCTGAACGGGGGGTTCGTGCACACAGCCCAGCTTGGAGCGAACGACCTACACCGAACTGAGATACCTACAGCGTGAGCTATGAGAAAGCGCCACGCTTCCCGAAGGGAGAAAGGCGGACAGGTATCCGGTAAGCGGCAGGGTCGGAACAGGAGAGCGCACGAGGGAGCTTCCAGGGGGAAACGCCTGGTATCTTTATAGTCCTGTCGGGTTTCGCCACCTCTGACTTGAGCGTCGATTTTTGTGATGCTCGTCAGGGGGGCGGAGCCTATGGAAA
>backbone_element
AACGCCAGCAACGCGGCCTTTTTACGGTTCCTGGCCTTTTGCTGGCCTTTTGCTCACATGTCCTGCAGGCAG
df_fasta = vb.decode_fasta_to_dataframe(fasta_string)
df_fasta = vb.extract_fasta_info(df_fasta)
graph = vb.plot_vector(df_fasta, title = None, title_size = 20, show_plot = True)
graph.savefig('example_graph.svg)
Output:

2.4.5 Writing FASTA format of the plasmid vector
vb.write_fasta(fasta_string, path = None, name = 'fasta_file')
This function saves into FASTA *.fasta
Args:
fasta_string (str/FASTA) - sequences provided in FASTA format from generate_fasta_string() or loaded from external sources
path (str | None) - the path to save. If None save it to the current working directory. Default: None
name (str) - the name of the saving file. Default: 'fasta_file'
2.4.6 Converting FASTA format to GeneBank format
gb_format = vb.get_genebank(df_fasta,
name = 'viral_vector',
definition = 'Synthetic viral plasmid vector')
Generate a GenBank (.gb) file based on a FASTA DataFrame.
This function takes a DataFrame containing parsed FASTA data from a plasmid vector
and converts it into a GenBank-formatted file. It is designed to work with
outputs generated by the pipeline:
`load_fasta(path) -> decode_fasta_to_dataframe(fasta) -> extract_fasta_info(df_fasta)`.
Args:
df_fasta (DataFrame) - dataframe obtained from load_fasta(path) -> decode fasta_to_dataframe(fasta) -> extract_fasta_info(df_fasta) pipeline which prepare decoded FASTA file of vector plasmid.
*dedicated FASTA structure:
>name1
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACC
>name2
TCTAGACAACTTTGTATAGAAAAGTTG
>name3
GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTAC
Header explanation:
name1,2,3,... - the name of the sequence element
name (str) - name of the GenBank data. Default is 'viral_vector'.
definition (str) - description of the plasmid/vector for the GenBank `DEFINITION` field.
Default is 'Synthetic viral plasmid vector'.
Returns:
Str: The GeneBank text format based on the provided DataFrame FASTA data
Full pipeline example:
# Exaple FASTA file is loaded from the library repository
import pkg_resources
from jbst import vector_build as vb
fasta_string = vb.load_fasta(pkg_resources.resource_filename("jbst", "tests/fasta_vector_test.fasta"))
Example FASTA file content:
# test_expression_ssAAV_expression_8717nc
>5`ITR
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTCAGTGAGCGAGCGAGCGCGCAGAGAGGGAGTGGCCAACTCCATCACTAGGGGTTCCT
>backbone_element
TCTAGACAACTTTGTATAGAAAAGTTG
>Promoter:TBG
GGGCTGGAAGCTACCTTTGACATCATTTCCTCTGCGAATGCATGTATAATTTCTACAGAACCTATTAGAAAGGATCACCCAGCCTCTGCTTTTGTACAACTTTCCCTTAAAAAACTGCCAATTCCACTGCTGTTTGGCCCAATAGTGAGAACTTTTTCCTGCTGCCTCTTGGTGCTTTTGCCTATGGCCCCTATTCTGCCTGCTGAAGACACTCTTGCCAGCATGGACTTAAACCCCTCCAGCTCTGACAATCCTCTTTCTCTTTTGTTTTACATGAAGGGTCTGGCAGCCAAAGCAATCACTCAAAGTTCAAACCTTATCATTTTTTGCTTTGTTCCTCTTGGCCTTGGTTTTGTACATCAGCTTTGAAAATACCATCCCAGGGTTAATGCTGGGGTTAATTTATAACTAAGAGTGCTCTAGTTTTGCAATACAGGACATGCTATAAAAATGGAAAGAT
>backbone_element
CAAGTTTGTACAAAAAAGCAGGCT
>Kozak_sequence
GCCACC
>SEQ1:SMN1
ATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTG
>SEQ2:SMN2
ATGGCTATGTCTAGCGGAGGCTCTGGAGGAGGAGTTCCTGAACAGGAGGACTCTGTGCTGTTCCGGAGGGGCACAGGACAAAGCGATGACAGCGACATCTGGGACGACACAGCTCTGATTAAGGCCTACGACAAGGCCGTGGCCAGCTTCAAGCACGCCCTGAAGAACGGCGACATCTGCGAGACCAGCGGAAAGCCTAAAACCACCCCTAAGAGAAAGCCTGCTAAAAAGAACAAGAGCCAGAAGAAGAACACCGCTGCCAGCCTGCAGCAGTGGAAGGTGGGCGACAAGTGCAGCGCCATTTGGAGCGAGGACGGATGTATCTACCCTGCCACAATCGCCAGCATCGACTTCAAGCGGGAGACCTGCGTGGTGGTGTATACCGGCTACGGCAACAGGGAAGAGCAGAACCTGAGCGACCTGCTGAGCCCTATTTGCGAGGTGGCCAATAACATCGAGCAGAACGCCCAGGAGAACGAGAACGAGAGCCAGGTGAGCACCGACGAGAGCGAGAACAGCCGGAGCCCCGGCAATAAGAGCGACAACATCAAGCCCAAGAGCGCCCCCTGGAACTCTTTCCTGCCCCCCCCCCCCCCCATGCCTGGACCTAGATTGGGACCTGGAAAACCTGGACTGAAATTCAACGGCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCATTTGCTGTCTTGTTGGCTGCCCCCCTTCCCTTCTGGACCCCCCATTATCCCCCCCCCCCCCCCCATCTGTCCTGATTCTCTGGACGACGCCGATGCTTTGGGCTCTATGCTGATCTCTTGGTATATGAGCGGCTACCACACCGGCTACTACATGTTCCCCGAGGCCAGCCTGAAGGCCGAGCAGATGCCCGCTCCTTGTTTTCTGTGA
>PolyA_signal:SV40_late
CAGACATGATAAGATACATTGATGAGTTTGGACAAACCACAACTAGAATGCAGTGAAAAAAATGCTTTATTTGTGAAATTTGTGATGCTATTGCTTTATTTGTAACCATTATAAGCTGCAATAAACAAGTTAACAACAACAATTGCATTCATTTTATGTTTCAGGTTCAGGGGGAGGTGTGGGAGGTTTTTTAAAGCAAGTAAAACCTCTACAAATGTGGTA
>2nd_promoter:CAG
CTCGACATTGATTATTGACTAGTTATTAATAGTAATCAATTACGGGGTCATTAGTTCATAGCCCATATATGGAGTTCCGCGTTACATAACTTACGGTAAATGGCCCGCCTGGCTGACCGCCCAACGACCCCCGCCCATTGACGTCAATAATGACGTATGTTCCCATAGTAACGCCAATAGGGACTTTCCATTGACGTCAATGGGTGGAGTATTTACGGTAAACTGCCCACTTGGCAGTACATCAAGTGTATCATATGCCAAGTACGCCCCCTATTGACGTCAATGACGGTAAATGGCCCGCCTGGCATTATGCCCAGTACATGACCTTATGGGACTTTCCTACTTGGCAGTACATCTACGTATTAGTCATCGCTATTACCATGGTCGAGGTGAGCCCCACGTTCTGCTTCACTCTCCCCATCTCCCCCCCCTCCCCACCCCCAATTTTGTATTTATTTATTTTTTAATTATTTTGTGCAGCGATGGGGGCGGGGGGGGGGGGGGGGCGCGCGCCAGGCGGGGCGGGGCGGGGCGAGGGGCGGGGCGGGGCGAGGCGGAGAGGTGCGGCGGCAGCCAATCAGAGCGGCGCGCTCCGAAAGTTTCCTTTTATGGCGAGGCGGCGGCGGCGGCGGCCCTATAAAAAGCGAAGCGCGCGGCGGGCGGGAGTCGCTGCGCGCTGCCTTCGCCCCGTGCCCCGCTCCGCCGCCGCCTCGCGCCGCCCGCCCCGGCTCTGACTGACCGCGTTACTCCCACAGGTGAGCGGGCGGGACGGCCCTTCTCCTCCGGGCTGTAATTAGCGCTTGGTTTAATGACGGCTTGTTTCTTTTCTGTGGCTGCGTGAAAGCCTTGAGGGGCTCCGGGAGGGCCCTTTGTGCGGGGGGAGCGGCTCGGGGGGTGCGTGCGTGTGTGTGTGCGTGGGGAGCGCCGCGTGCGGCTCCGCGCTGCCCGGCGGCTGTGAGCGCTGCGGGCGCGGCGCGGGGCTTTGTGCGCTCCGCAGTGTGCGCGAGGGGAGCGCGGCCGGGGGCGGTGCCCCGCGGTGCGGGGGGGGCTGCGAGGGGAACAAAGGCTGCGTGCGGGGTGTGTGCGTGGGGGGGTGAGCAGGGGGTGTGGGCGCGTCGGTCGGGCTGCAACCCCCCCTGCACCCCCCTCCCCGAGTTGCTGAGCACGGCCCGGCTTCGGGTGCGGGGCTCCGTACGGGGCGTGGCGCGGGGCTCGCCGTGCCGGGCGGGGGGTGGCGGCAGGTGGGGGTGCCGGGCGGGGCGGGGCCGCCTCGGGCCGGGGAGGGCTCGGGGGAGGGGCGCGGCGGCCCCCGGAGCGCCGGCGGCTGTCGAGGCGCGGCGAGCCGCAGCCATTGCCTTTTATGGTAATCGTGCGAGAGGGCGCAGGGACTTCCTTTGTCCCAAATCTGTGCGGAGCCGAAATCTGGGAGGCGCCGCCGCACCCCCTCTAGCGGGCGCGGGGCGAAGCGGTGCGGCGCCGGCAGGAAGGAAATGGGCGGGGAGGGCCTTCGTGCGTCGCCGCGCCGCCGTCCCCTTCTCCCTCTCCAGCCTCGGGGCTGTCCGCGGGGGGACGGCTGCCTTCGGGGGGGACGGGGCAGGGCGGGGTTCGGCTTCTGGCGTGTGACCGGCGGCTCTAGAGCCTCTGCTAACCATGTTCATGCCTTCTTCTTTTTCCTACAGCTCCTGGGCAACGTGCTGGTTATTGTGCTGTCTCATCATTTTGGCAAAGAATTG
>Fluorescent_tag:EGFP
ATGGTGAGCAAGGGCGAGGAGCTGTTCACCGGGGTGGTGCCCATCCTGGTCGAGCTGGACGGCGACGTAAACGGCCACAAGTTCAGCGTGTCCGGCGAGGGCGAGGGCGATGCCACCTACGGCAAGCTGACCCTGAAGTTCATCTGCACCACCGGCAAGCTGCCCGTGCCCTGGCCCACCCTCGTGACCACCCTGACCTACGGCGTGCAGTGCTTCAGCCGCTACCCCGACCACATGAAGCAGCACGACTTCTTCAAGTCCGCCATGCCCGAAGGCTACGTCCAGGAGCGCACCATCTTCTTCAAGGACGACGGCAACTACAAGACCCGCGCCGAGGTGAAGTTCGAGGGCGACACCCTGGTGAACCGCATCGAGCTGAAGGGCATCGACTTCAAGGAGGACGGCAACATCCTGGGGCACAAGCTGGAGTACAACTACAACAGCCACAACGTCTATATCATGGCCGACAAGCAGAAGAACGGCATCAAGGTGAACTTCAAGATCCGCCACAACATCGAGGACGGCAGCGTGCAGCTCGCCGACCACTACCAGCAGAACACCCCCATCGGCGACGGCCCCGTGCTGCTGCCCGACAACCACTACCTGAGCACCCAGTCCGCCCTGAGCAAAGACCCCAACGAGAAGCGCGATCACATGGTCCTGCTGGAGTTCGTGACCGCCGCCGGGATCACTCTCGGCATGGACGAGCTGTACAAGTAA
>backbone_element
ACCCAGCTTTCTTGTACAAAGTGGGAATTC
>Enhancer:WPRE
CGATAATCAACCTCTGGATTACAAAATTTGTGAAAGATTGACTGGTATTCTTAACTATGTTGCTCCTTTTACGCTATGTGGATACGCTGCTTTAATGCCTTTGTATCATGCTATTGCTTCCCGTATGGCTTTCATTTTCTCCTCCTTGTATAAATCCTGGTTGCTGTCTCTTTATGAGGAGTTGTGGCCCGTTGTCAGGCAACGTGGCGTGGTGTGCACTGTGTTTGCTGACGCAACCCCCACTGGTTGGGGCATTGCCACCACCTGTCAGCTCCTTTCCGGGACTTTCGCTTTCCCCCTCCCTATTGCCACGGCGGAACTCATCGCCGCCTGCCTTGCCCGCTGCTGGACAGGGGCTCGGCTGTTGGGCACTGACAATTCCGTGGTGTTGTCGGGGAAGCTGACGTCCTTTCCATGGCTGCTCGCCTGTGTTGCCACCTGGATTCTGCGCGGGACGTCCTTCTGCTACGTCCCTTCGGCCCTCAATCCAGCGGACCTTCCTTCCCGCGGCCTGCTGCCGGCTCTGCGGCCTCTTCCGCGTCTTCGCCTTCGCCCTCAGACGAGTCGGATCTCCCTTTGGGCCGCCTCCCCGCATCGG
>backbone_element
GAATTCCTAGAGCTCGCTGATCAGCCTCGA
>2nd_polyA_signal:bGH
CTGTGCCTTCTAGTTGCCAGCCATCTGTTGTTTGCCCCTCCCCCGTGCCTTCCTTGACCCTGGAAGGTGCCACTCCCACTGTCCTTTCCTAATAAAATGAGGAAATTGCATCGCATTGTCTGAGTAGGTGTCATTCTATTCTGGGGGGTGGGGTGGGGCAGGACAGCAAGGGGGAGGATTGGGAAGAGAATAGCAGGCATGCTGGGGA
>backbone_element
GGGCCGC
>3`ITR
CTGCGCGCTCGCTCGCTCACTGAGGCCGCCCGGGCAAAGCCCGGGCGTCGGGCGACCTTTGGTCGCCCGGCCTCAGTGAGCGAGCGAGCGCGCAGAGAGGGAGTGGCCAACTCCATCACTAGGGGTTCCT
>backbone_element
CTGCCTGCAGGGGCGCCTGATGCGGTATTTTCTCCTTACGCATCTGTGCGGTATTTCACACCGCATACGTCAAAGCAACCATAGTACGCGCCCTGTAGCGGCGCATTAAGCGCGGCGGGGGTGGTGGTTACGCGCAGCGTGACCGCTACACTTGCCAGCGCCTTAGCGCCCGCTCCTTTCGCTTTCTTCCCTTCCTTTCTCGCCACGTTCGCCGGCTTTCCCCGTCAAGCTCTAAATCGGGGGCTCCCTTTAGGGTTCCGATTTAGTGCTTTACGGCACCTCGACCCCAAAAAACTTGATTTGGGTGATGGTTCACGTAGTGGGCCATCGCCCTGATAGACGGTTTTTCGCCCTTTGACGTTGGAGTCCACGTTCTTTAATAGTGGACTCTTGTTCCAAACTGGAACAACACTCAACTCTATCTCGGGCTATTCTTTTGATTTATAAGGGATTTTGCCGATTTCGGTCTATTGGTTAAAAAATGAGCTGATTTAACAAAAATTTAACGCGAATTTTAACAAAATATTAACGTTTACAATTTTATGGTGCACTCTCAGTACAATCTGCTCTGATGCCGCATAGTTAAGCCAGCCCCGACACCCGCCAACACCCGCTGACGCGCCCTGACGGGCTTGTCTGCTCCCGGCATCCGCTTACAGACAAGCTGTGACCGTCTCCGGGAGCTGCATGTGTCAGAGGTTTTCACCGTCATCACCGAAACGCGCGAGACGAAAGGGCCTCGTGATACGCCTATTTTTATAGGTTAATGTCATGATAATAATGGTTTCTTAGACGTCAGGTGGCACTTTTCGGGGAAATGTGCGCGGAACCCCTATTTGTTTATTTTTCTAAATACATTCAAATATGTATCCGCTCATGAGACAATAACCCTGATAAATGCTTCAATAATATTGAAAAAGGAAGAGT
>Resistance:Ampicillin
ATGAGTATTCAACATTTCCGTGTCGCCCTTATTCCCTTTTTTGCGGCATTTTGCCTTCCTGTTTTTGCTCACCCAGAAACGCTGGTGAAAGTAAAAGATGCTGAAGATCAGTTGGGTGCACGAGTGGGTTACATCGAACTGGATCTCAACAGCGGTAAGATCCTTGAGAGTTTTCGCCCCGAAGAACGTTTTCCAATGATGAGCACTTTTAAAGTTCTGCTATGTGGCGCGGTATTATCCCGTATTGACGCCGGGCAAGAGCAACTCGGTCGCCGCATACACTATTCTCAGAATGACTTGGTTGAGTACTCACCAGTCACAGAAAAGCATCTTACGGATGGCATGACAGTAAGAGAATTATGCAGTGCTGCCATAACCATGAGTGATAACACTGCGGCCAACTTACTTCTGACAACGATCGGAGGACCGAAGGAGCTAACCGCTTTTTTGCACAACATGGGGGATCATGTAACTCGCCTTGATCGTTGGGAACCGGAGCTGAATGAAGCCATACCAAACGACGAGCGTGACACCACGATGCCTGTAGCAATGGCAACAACGTTGCGCAAACTATTAACTGGCGAACTACTTACTCTAGCTTCCCGGCAACAATTAATAGACTGGATGGAGGCGGATAAAGTTGCAGGACCACTTCTGCGCTCGGCCCTTCCGGCTGGCTGGTTTATTGCTGATAAATCTGGAGCCGGTGAGCGTGGAAGCCGCGGTATCATTGCAGCACTGGGGCCAGATGGTAAGCCCTCCCGTATCGTAGTTATCTACACGACGGGGAGTCAGGCAACTATGGATGAACGAAATAGACAGATCGCTGAGATAGGTGCCTCACTGATTAAGCATTGGTAA
>backbone_element
CTGTCAGACCAAGTTTACTCATATATACTTTAGATTGATTTAAAACTTCATTTTTAATTTAAAAGGATCTAGGTGAAGATCCTTTTTGATAATCTCATGACCAAAATCCCTTAACGTGAGTTTTCGTTCCACTGAGCGTCAGACCCCGTAGAAAAGATCAAAGGATCTTC
>pUC_ori
TTGAGATCCTTTTTTTCTGCGCGTAATCTGCTGCTTGCAAACAAAAAAACCACCGCTACCAGCGGTGGTTTGTTTGCCGGATCAAGAGCTACCAACTCTTTTTCCGAAGGTAACTGGCTTCAGCAGAGCGCAGATACCAAATACTGTTCTTCTAGTGTAGCCGTAGTTAGGCCACCACTTCAAGAACTCTGTAGCACCGCCTACATACCTCGCTCTGCTAATCCTGTTACCAGTGGCTGCTGCCAGTGGCGATAAGTCGTGTCTTACCGGGTTGGACTCAAGACGATAGTTACCGGATAAGGCGCAGCGGTCGGGCTGAACGGGGGGTTCGTGCACACAGCCCAGCTTGGAGCGAACGACCTACACCGAACTGAGATACCTACAGCGTGAGCTATGAGAAAGCGCCACGCTTCCCGAAGGGAGAAAGGCGGACAGGTATCCGGTAAGCGGCAGGGTCGGAACAGGAGAGCGCACGAGGGAGCTTCCAGGGGGAAACGCCTGGTATCTTTATAGTCCTGTCGGGTTTCGCCACCTCTGACTTGAGCGTCGATTTTTGTGATGCTCGTCAGGGGGGCGGAGCCTATGGAAA
>backbone_element
AACGCCAGCAACGCGGCCTTTTTACGGTTCCTGGCCTTTTGCTGGCCTTTTGCTCACATGTCCTGCAGGCAG
df_fasta = vb.decode_fasta_to_dataframe(fasta_string)
df_fasta = vb.extract_fasta_info(df_fasta)
gb_format = vb.get_genebank(df_fasta,
name = 'viral_vector',
definition = ' Synthetic viral plasmid vector')
Output:
LOCUS viral_vector 8717 bp DNA circular 11-JUL-2025
DEFINITION Synthetic viral plasmid vector.
FEATURES Location/Qualifiers
misc_feature 1..130
/note="5`ITR"
misc_feature 158..617
/note="Promoter:TBG"
misc_feature 642..647
/note="Kozak_sequence"
misc_feature 648..1532
/note="SEQ1:SMN1"
misc_feature 1533..2420
/note="SEQ2:SMN2"
misc_feature 2421..2642
/note="PolyA_signal:SV40"
misc_feature 2643..4375
/note="2nd_promoter:CAG"
misc_feature 4376..5095
/note="Fluorescent_tag:EGFP"
misc_feature 5126..5723
/note="Enhancer:WPRE"
misc_feature 5754..5961
/note="2nd_polyA_signal:bGH"
misc_feature 5969..6098
/note="3`ITR"
misc_feature 7026..7886
/note="Resistance:Ampicillin"
misc_feature 8057..8645
/note="pUC_ori"
ORIGIN
1 ctgcgcgctc gctcgctcac tgaggccgcc cgggcaaagc ccgggcgtcg ggcgaccttt
61 ggtcgcccgg cctcagtgag cgagcgagcg cgcagagagg gagtggccaa ctccatcact
121 aggggttcct tctagacaac tttgtataga aaagttgggg ctggaagcta cctttgacat
181 catttcctct gcgaatgcat gtataatttc tacagaacct attagaaagg atcacccagc
241 ctctgctttt gtacaacttt cccttaaaaa actgccaatt ccactgctgt ttggcccaat
301 agtgagaact ttttcctgct gcctcttggt gcttttgcct atggccccta ttctgcctgc
361 tgaagacact cttgccagca tggacttaaa cccctccagc tctgacaatc ctctttctct
421 tttgttttac atgaagggtc tggcagccaa agcaatcact caaagttcaa accttatcat
481 tttttgcttt gttcctcttg gccttggttt tgtacatcag ctttgaaaat accatcccag
541 ggttaatgct ggggttaatt tataactaag agtgctctag ttttgcaata caggacatgc
601 tataaaaatg gaaagatcaa gtttgtacaa aaaagcaggc tgccaccatg gctatgtcta
661 gcggaggctc tggaggagga gttcctgaac aggaggactc tgtgctgttc cggaggggca
721 caggacaaag cgatgacagc gacatctggg acgacacagc tctgattaag gcctacgaca
781 aggccgtggc cagcttcaag cacgccctga agaacggcga catctgcgag accagcggaa
841 agcctaaaac cacccctaag agaaagcctg ctaaaaagaa caagagccag aagaagaaca
901 ccgctgccag cctgcagcag tggaaggtgg gcgacaagtg cagcgccatt tggagcgagg
961 acggatgtat ctaccctgcc acaatcgcca gcatcgactt caagcgggag acctgcgtgg
1021 tggtgtatac cggctacggc aacagggaag agcagaacct gagcgacctg ctgagcccta
1081 tttgcgaggt ggccaataac atcgagcaga acgcccagga gaacgagaac gagagccagg
1141 tgagcaccga cgagagcgag aacagccgga gccccggcaa taagagcgac aacatcaagc
1201 ccaagagcgc cccctggaac tctttcctgc cccccccccc ccccatgcct ggacctagat
1261 tgggacctgg aaaacctgga ctgaaattca acggcccccc cccccccccc cccccccccc
1321 ccccccattt gctgtcttgt tggctgcccc ccttcccttc tggacccccc attatccccc
1381 cccccccccc catctgtcct gattctctgg acgacgccga tgctttgggc tctatgctga
1441 tctcttggta tatgagcggc taccacaccg gctactacat gttccccgag gccagcctga
1501 aggccgagca gatgcccgct ccttgttttc tgatggctat gtctagcgga ggctctggag
1561 gaggagttcc tgaacaggag gactctgtgc tgttccggag gggcacagga caaagcgatg
1621 acagcgacat ctgggacgac acagctctga ttaaggccta cgacaaggcc gtggccagct
1681 tcaagcacgc cctgaagaac ggcgacatct gcgagaccag cggaaagcct aaaaccaccc
1741 ctaagagaaa gcctgctaaa aagaacaaga gccagaagaa gaacaccgct gccagcctgc
1801 agcagtggaa ggtgggcgac aagtgcagcg ccatttggag cgaggacgga tgtatctacc
1861 ctgccacaat cgccagcatc gacttcaagc gggagacctg cgtggtggtg tataccggct
1921 acggcaacag ggaagagcag aacctgagcg acctgctgag ccctatttgc gaggtggcca
1981 ataacatcga gcagaacgcc caggagaacg agaacgagag ccaggtgagc accgacgaga
2041 gcgagaacag ccggagcccc ggcaataaga gcgacaacat caagcccaag agcgccccct
2101 ggaactcttt cctgcccccc ccccccccca tgcctggacc tagattggga cctggaaaac
2161 ctggactgaa attcaacggc cccccccccc cccccccccc cccccccccc catttgctgt
2221 cttgttggct gccccccttc ccttctggac cccccattat cccccccccc ccccccatct
2281 gtcctgattc tctggacgac gccgatgctt tgggctctat gctgatctct tggtatatga
2341 gcggctacca caccggctac tacatgttcc ccgaggccag cctgaaggcc gagcagatgc
2401 ccgctccttg ttttctgtga cagacatgat aagatacatt gatgagtttg gacaaaccac
2461 aactagaatg cagtgaaaaa aatgctttat ttgtgaaatt tgtgatgcta ttgctttatt
2521 tgtaaccatt ataagctgca ataaacaagt taacaacaac aattgcattc attttatgtt
2581 tcaggttcag ggggaggtgt gggaggtttt ttaaagcaag taaaacctct acaaatgtgg
2641 tactcgacat tgattattga ctagttatta atagtaatca attacggggt cattagttca
2701 tagcccatat atggagttcc gcgttacata acttacggta aatggcccgc ctggctgacc
2761 gcccaacgac ccccgcccat tgacgtcaat aatgacgtat gttcccatag taacgccaat
2821 agggactttc cattgacgtc aatgggtgga gtatttacgg taaactgccc acttggcagt
2881 acatcaagtg tatcatatgc caagtacgcc ccctattgac gtcaatgacg gtaaatggcc
2941 cgcctggcat tatgcccagt acatgacctt atgggacttt cctacttggc agtacatcta
3001 cgtattagtc atcgctatta ccatggtcga ggtgagcccc acgttctgct tcactctccc
3061 catctccccc ccctccccac ccccaatttt gtatttattt attttttaat tattttgtgc
3121 agcgatgggg gcgggggggg ggggggggcg cgcgccaggc ggggcggggc ggggcgaggg
3181 gcggggcggg gcgaggcgga gaggtgcggc ggcagccaat cagagcggcg cgctccgaaa
3241 gtttcctttt atggcgaggc ggcggcggcg gcggccctat aaaaagcgaa gcgcgcggcg
3301 ggcgggagtc gctgcgcgct gccttcgccc cgtgccccgc tccgccgccg cctcgcgccg
3361 cccgccccgg ctctgactga ccgcgttact cccacaggtg agcgggcggg acggcccttc
3421 tcctccgggc tgtaattagc gcttggttta atgacggctt gtttcttttc tgtggctgcg
3481 tgaaagcctt gaggggctcc gggagggccc tttgtgcggg gggagcggct cggggggtgc
3541 gtgcgtgtgt gtgtgcgtgg ggagcgccgc gtgcggctcc gcgctgcccg gcggctgtga
3601 gcgctgcggg cgcggcgcgg ggctttgtgc gctccgcagt gtgcgcgagg ggagcgcggc
3661 cgggggcggt gccccgcggt gcgggggggg ctgcgagggg aacaaaggct gcgtgcgggg
3721 tgtgtgcgtg ggggggtgag cagggggtgt gggcgcgtcg gtcgggctgc aaccccccct
3781 gcacccccct ccccgagttg ctgagcacgg cccggcttcg ggtgcggggc tccgtacggg
3841 gcgtggcgcg gggctcgccg tgccgggcgg ggggtggcgg caggtggggg tgccgggcgg
3901 ggcggggccg cctcgggccg gggagggctc gggggagggg cgcggcggcc cccggagcgc
3961 cggcggctgt cgaggcgcgg cgagccgcag ccattgcctt ttatggtaat cgtgcgagag
4021 ggcgcaggga cttcctttgt cccaaatctg tgcggagccg aaatctggga ggcgccgccg
4081 caccccctct agcgggcgcg gggcgaagcg gtgcggcgcc ggcaggaagg aaatgggcgg
4141 ggagggcctt cgtgcgtcgc cgcgccgccg tccccttctc cctctccagc ctcggggctg
4201 tccgcggggg gacggctgcc ttcggggggg acggggcagg gcggggttcg gcttctggcg
4261 tgtgaccggc ggctctagag cctctgctaa ccatgttcat gccttcttct ttttcctaca
4321 gctcctgggc aacgtgctgg ttattgtgct gtctcatcat tttggcaaag aattgatggt
4381 gagcaagggc gaggagctgt tcaccggggt ggtgcccatc ctggtcgagc tggacggcga
4441 cgtaaacggc cacaagttca gcgtgtccgg cgagggcgag ggcgatgcca cctacggcaa
4501 gctgaccctg aagttcatct gcaccaccgg caagctgccc gtgccctggc ccaccctcgt
4561 gaccaccctg acctacggcg tgcagtgctt cagccgctac cccgaccaca tgaagcagca
4621 cgacttcttc aagtccgcca tgcccgaagg ctacgtccag gagcgcacca tcttcttcaa
4681 ggacgacggc aactacaaga cccgcgccga ggtgaagttc gagggcgaca ccctggtgaa
4741 ccgcatcgag ctgaagggca tcgacttcaa ggaggacggc aacatcctgg ggcacaagct
4801 ggagtacaac tacaacagcc acaacgtcta tatcatggcc gacaagcaga agaacggcat
4861 caaggtgaac ttcaagatcc gccacaacat cgaggacggc agcgtgcagc tcgccgacca
4921 ctaccagcag aacaccccca tcggcgacgg ccccgtgctg ctgcccgaca accactacct
4981 gagcacccag tccgccctga gcaaagaccc caacgagaag cgcgatcaca tggtcctgct
5041 ggagttcgtg accgccgccg ggatcactct cggcatggac gagctgtaca agtaaaccca
5101 gctttcttgt acaaagtggg aattccgata atcaacctct ggattacaaa atttgtgaaa
5161 gattgactgg tattcttaac tatgttgctc cttttacgct atgtggatac gctgctttaa
5221 tgcctttgta tcatgctatt gcttcccgta tggctttcat tttctcctcc ttgtataaat
5281 cctggttgct gtctctttat gaggagttgt ggcccgttgt caggcaacgt ggcgtggtgt
5341 gcactgtgtt tgctgacgca acccccactg gttggggcat tgccaccacc tgtcagctcc
5401 tttccgggac tttcgctttc cccctcccta ttgccacggc ggaactcatc gccgcctgcc
5461 ttgcccgctg ctggacaggg gctcggctgt tgggcactga caattccgtg gtgttgtcgg
5521 ggaagctgac gtcctttcca tggctgctcg cctgtgttgc cacctggatt ctgcgcggga
5581 cgtccttctg ctacgtccct tcggccctca atccagcgga ccttccttcc cgcggcctgc
5641 tgccggctct gcggcctctt ccgcgtcttc gccttcgccc tcagacgagt cggatctccc
5701 tttgggccgc ctccccgcat cgggaattcc tagagctcgc tgatcagcct cgactgtgcc
5761 ttctagttgc cagccatctg ttgtttgccc ctcccccgtg ccttccttga ccctggaagg
5821 tgccactccc actgtccttt cctaataaaa tgaggaaatt gcatcgcatt gtctgagtag
5881 gtgtcattct attctggggg gtggggtggg gcaggacagc aagggggagg attgggaaga
5941 gaatagcagg catgctgggg agggccgcct gcgcgctcgc tcgctcactg aggccgcccg
6001 ggcaaagccc gggcgtcggg cgacctttgg tcgcccggcc tcagtgagcg agcgagcgcg
6061 cagagaggga gtggccaact ccatcactag gggttcctct gcctgcaggg gcgcctgatg
6121 cggtattttc tccttacgca tctgtgcggt atttcacacc gcatacgtca aagcaaccat
6181 agtacgcgcc ctgtagcggc gcattaagcg cggcgggggt ggtggttacg cgcagcgtga
6241 ccgctacact tgccagcgcc ttagcgcccg ctcctttcgc tttcttccct tcctttctcg
6301 ccacgttcgc cggctttccc cgtcaagctc taaatcgggg gctcccttta gggttccgat
6361 ttagtgcttt acggcacctc gaccccaaaa aacttgattt gggtgatggt tcacgtagtg
6421 ggccatcgcc ctgatagacg gtttttcgcc ctttgacgtt ggagtccacg ttctttaata
6481 gtggactctt gttccaaact ggaacaacac tcaactctat ctcgggctat tcttttgatt
6541 tataagggat tttgccgatt tcggtctatt ggttaaaaaa tgagctgatt taacaaaaat
6601 ttaacgcgaa ttttaacaaa atattaacgt ttacaatttt atggtgcact ctcagtacaa
6661 tctgctctga tgccgcatag ttaagccagc cccgacaccc gccaacaccc gctgacgcgc
6721 cctgacgggc ttgtctgctc ccggcatccg cttacagaca agctgtgacc gtctccggga
6781 gctgcatgtg tcagaggttt tcaccgtcat caccgaaacg cgcgagacga aagggcctcg
6841 tgatacgcct atttttatag gttaatgtca tgataataat ggtttcttag acgtcaggtg
6901 gcacttttcg gggaaatgtg cgcggaaccc ctatttgttt atttttctaa atacattcaa
6961 atatgtatcc gctcatgaga caataaccct gataaatgct tcaataatat tgaaaaagga
7021 agagtatgag tattcaacat ttccgtgtcg cccttattcc cttttttgcg gcattttgcc
7081 ttcctgtttt tgctcaccca gaaacgctgg tgaaagtaaa agatgctgaa gatcagttgg
7141 gtgcacgagt gggttacatc gaactggatc tcaacagcgg taagatcctt gagagttttc
7201 gccccgaaga acgttttcca atgatgagca cttttaaagt tctgctatgt ggcgcggtat
7261 tatcccgtat tgacgccggg caagagcaac tcggtcgccg catacactat tctcagaatg
7321 acttggttga gtactcacca gtcacagaaa agcatcttac ggatggcatg acagtaagag
7381 aattatgcag tgctgccata accatgagtg ataacactgc ggccaactta cttctgacaa
7441 cgatcggagg accgaaggag ctaaccgctt ttttgcacaa catgggggat catgtaactc
7501 gccttgatcg ttgggaaccg gagctgaatg aagccatacc aaacgacgag cgtgacacca
7561 cgatgcctgt agcaatggca acaacgttgc gcaaactatt aactggcgaa ctacttactc
7621 tagcttcccg gcaacaatta atagactgga tggaggcgga taaagttgca ggaccacttc
7681 tgcgctcggc ccttccggct ggctggttta ttgctgataa atctggagcc ggtgagcgtg
7741 gaagccgcgg tatcattgca gcactggggc cagatggtaa gccctcccgt atcgtagtta
7801 tctacacgac ggggagtcag gcaactatgg atgaacgaaa tagacagatc gctgagatag
7861 gtgcctcact gattaagcat tggtaactgt cagaccaagt ttactcatat atactttaga
7921 ttgatttaaa acttcatttt taatttaaaa ggatctaggt gaagatcctt tttgataatc
7981 tcatgaccaa aatcccttaa cgtgagtttt cgttccactg agcgtcagac cccgtagaaa
8041 agatcaaagg atcttcttga gatccttttt ttctgcgcgt aatctgctgc ttgcaaacaa
8101 aaaaaccacc gctaccagcg gtggtttgtt tgccggatca agagctacca actctttttc
8161 cgaaggtaac tggcttcagc agagcgcaga taccaaatac tgttcttcta gtgtagccgt
8221 agttaggcca ccacttcaag aactctgtag caccgcctac atacctcgct ctgctaatcc
8281 tgttaccagt ggctgctgcc agtggcgata agtcgtgtct taccgggttg gactcaagac
8341 gatagttacc ggataaggcg cagcggtcgg gctgaacggg gggttcgtgc acacagccca
8401 gcttggagcg aacgacctac accgaactga gatacctaca gcgtgagcta tgagaaagcg
8461 ccacgcttcc cgaagggaga aaggcggaca ggtatccggt aagcggcagg gtcggaacag
8521 gagagcgcac gagggagctt ccagggggaa acgcctggta tctttatagt cctgtcgggt
8581 ttcgccacct ctgacttgag cgtcgatttt tgtgatgctc gtcagggggg cggagcctat
8641 ggaaaaacgc cagcaacgcg gcctttttac ggttcctggc cttttgctgg ccttttgctc
8701 acatgtcctg caggcag
//
2.4.7 Writing GeneBank format of the plasmid vector
vb.write_genebank(gb_format, path = None, name = 'viral_vector')
This function saves into GeneBank format *.gb
Args:
fasta_string (str/GeneBank) - sequences provided in GeneBank format from get_genebank()
path (str | None) - the path to save. If None save it to the current working directory. Default: None
name (str) - the name of the saving file. Default: 'viral_vector'
Have fun JBS
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file jbioseqtools-2.1.4.tar.gz.
File metadata
- Download URL: jbioseqtools-2.1.4.tar.gz
- Upload date:
- Size: 25.2 MB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
7c3f7e0f279319ce06ce6e008989b71223bcc56f7f2ac2ce9e6d7e344750f0ea
|
|
| MD5 |
2bc78b052d05021147ee8a7908c9a29a
|
|
| BLAKE2b-256 |
d4777a9e238aeaadca7ea187258116737a337925ceb458003400b9119cf0626f
|
File details
Details for the file jbioseqtools-2.1.4-py3-none-any.whl.
File metadata
- Download URL: jbioseqtools-2.1.4-py3-none-any.whl
- Upload date:
- Size: 25.9 MB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: twine/6.2.0 CPython/3.11.9
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
28627ac7c620b6636837459b3b945e08cf3571785543ceced1aaa6ed02cb68b5
|
|
| MD5 |
e64046aad908fbb73e59ad710f3410a5
|
|
| BLAKE2b-256 |
eb15fa294feea0e206c847afc5e155665aae6eba5ff3bdf4e8e298f4f50eeeda
|











