A python package for parsing, merging, and analyzing Kraken2 output files.
Project description
kraut




A python package for parsing, merging, and analyzing Kraken2 output files.
Installation
From PyPI:
pip install krautils
From a local checkout:
pip install .
For developer's dependencies:
pip install .[dev]
pytest
Command
kraut parses, filters, merges, tabulates, splits, and plots Kraken2 reports.
single-report: filter or reformat one Kraken report.make-table: build a configurable multi-sample abundance table.ma-export: export reports for MicrobiomeAnalyst as counts, taxonomy, metadata, and tree files.alpha: calculate alpha diversity from Kraken or Bracken reports.beta: calculate beta diversity distance matrices and heatmap/PCA plots.dendrogram: plot hierarchical clustering dendrograms from beta distances.split-combine-table: split a KrakenTools combined table into ALL/LVL tables.plot-single: plot one sample as an HTML or static composition chart.plot-multi: plot multiple samples as an HTML or static stacked/bubble chart.
kraut alpha reports/*.tsv -o alpha.tsv -p alpha.html --metrics core --add-metrics chao1,ace
kraut beta reports/*.tsv -o beta.tsv --plot beta.html --pca beta_pca.html
kraut dendrogram reports/*.tsv -o dendrogram.html --distance braycurtis --clustering ward
kraut ma-export reports/*.tsv -o microbiomeanalyst --metadata metadata.tsv --metadata-sample-col Sample
ma-export writes counts.csv, taxonomy.csv, metadata.csv, and tree.nwk.
It uses taxon-specific counts by default (--metric LVL); use --metric TOT
for cumulative clade counts.
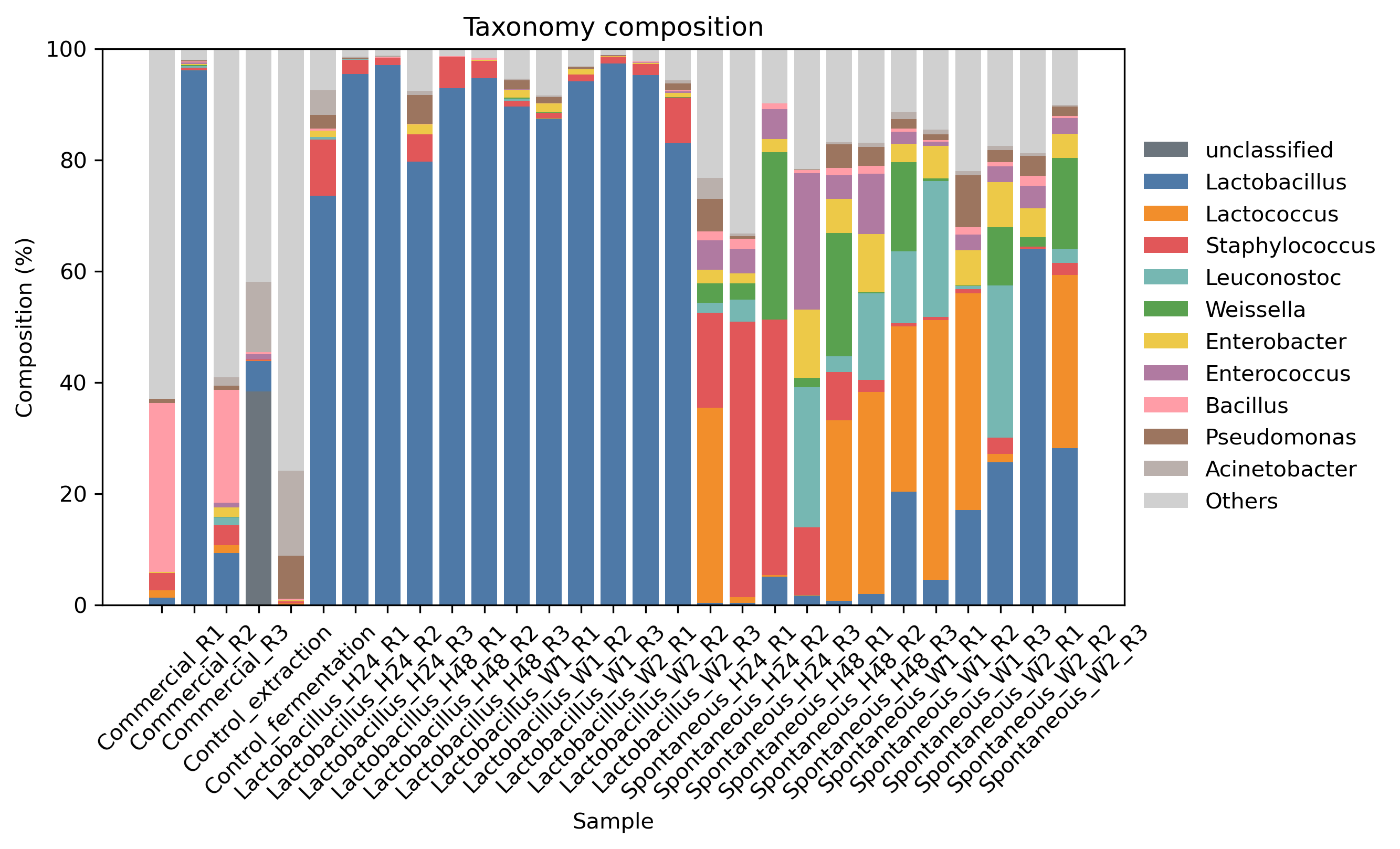
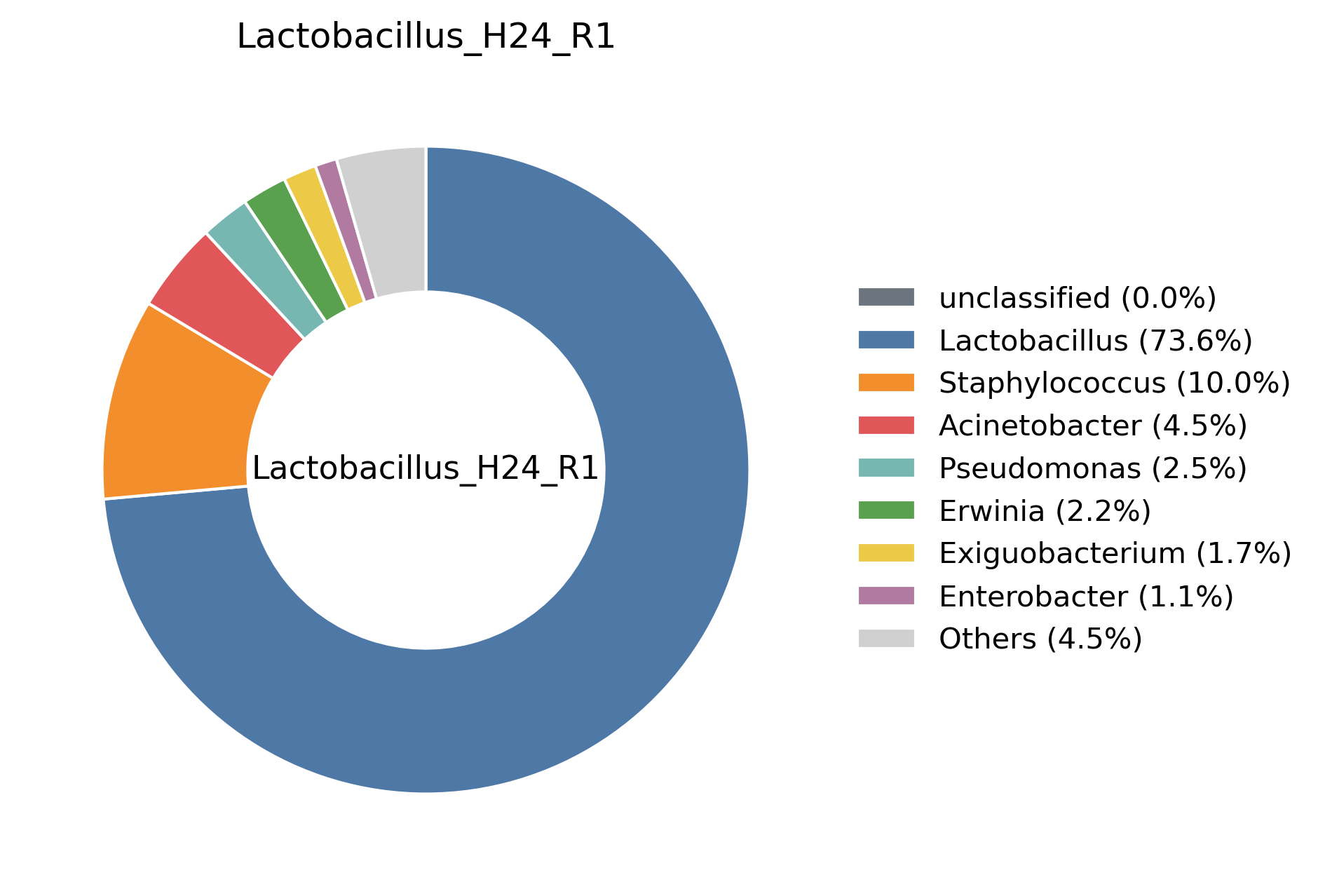
Example plots:
License
This project is licensed under the MIT License. See LICENSE.
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file krautils-0.5.0.tar.gz.
File metadata
- Download URL: krautils-0.5.0.tar.gz
- Upload date:
- Size: 50.5 kB
- Tags: Source
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
432efd5272dff24fe4c297c7230cf855b6aa4251b15cb2c9f953849dbb4160cf
|
|
| MD5 |
4b2a9f74d7b53ffd258a037410b467fb
|
|
| BLAKE2b-256 |
e80c813304ede2556b75e4a91405ffcbc7afa92b1d2a7027afa2569dfa7a9a6c
|
Provenance
The following attestation bundles were made for krautils-0.5.0.tar.gz:
Publisher:
publish.yml on quadram-institute-bioscience/kraut
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
krautils-0.5.0.tar.gz -
Subject digest:
432efd5272dff24fe4c297c7230cf855b6aa4251b15cb2c9f953849dbb4160cf - Sigstore transparency entry: 1449983313
- Sigstore integration time:
-
Permalink:
quadram-institute-bioscience/kraut@766fea2c54ba7acb6840f811f2e5a34c05e8408e -
Branch / Tag:
refs/tags/v0.5.0 - Owner: https://github.com/quadram-institute-bioscience
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@766fea2c54ba7acb6840f811f2e5a34c05e8408e -
Trigger Event:
release
-
Statement type:
File details
Details for the file krautils-0.5.0-py3-none-any.whl.
File metadata
- Download URL: krautils-0.5.0-py3-none-any.whl
- Upload date:
- Size: 48.9 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? Yes
- Uploaded via: twine/6.1.0 CPython/3.13.12
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
3dec27eef8a0f839d913a6e98ef53d4a1b359baea812d13cb1cbdf40c5724ef1
|
|
| MD5 |
65f857b05c3a5a527a45bc2c5ed43209
|
|
| BLAKE2b-256 |
08af4b7ce8bdafc2a5411e601a7cd461c9d003aefa06586d8cf0d556832e5fd6
|
Provenance
The following attestation bundles were made for krautils-0.5.0-py3-none-any.whl:
Publisher:
publish.yml on quadram-institute-bioscience/kraut
-
Statement:
-
Statement type:
https://in-toto.io/Statement/v1 -
Predicate type:
https://docs.pypi.org/attestations/publish/v1 -
Subject name:
krautils-0.5.0-py3-none-any.whl -
Subject digest:
3dec27eef8a0f839d913a6e98ef53d4a1b359baea812d13cb1cbdf40c5724ef1 - Sigstore transparency entry: 1449983600
- Sigstore integration time:
-
Permalink:
quadram-institute-bioscience/kraut@766fea2c54ba7acb6840f811f2e5a34c05e8408e -
Branch / Tag:
refs/tags/v0.5.0 - Owner: https://github.com/quadram-institute-bioscience
-
Access:
public
-
Token Issuer:
https://token.actions.githubusercontent.com -
Runner Environment:
github-hosted -
Publication workflow:
publish.yml@766fea2c54ba7acb6840f811f2e5a34c05e8408e -
Trigger Event:
release
-
Statement type: