Package to visualize LLM's Neural Networks activation regions
Project description
LLM-MRI: a brain scanner for LLMs
This repository contains the implementation from the paper LLM-MRI Python module: a brain scanner for LLMs
As the everyday use of large language models (LLMs) expands, so does the necessity of understanding how these models achieve their designated outputs. While many approaches focus on the interpretability of LLMs through visualizing different attention mechanisms and methods that explain the model's architecture, LLM-MRI focuses on the activations of the feed-forward layers in a transformer-based LLM.
By adopting this approach, the library examines the neuron activations produced by the model for each distinct label. Through a series of steps, such as dimensionality reduction and representing each layer as a grid, the tool provides various visualization methods for the activation patterns in the feed-forward layers. Accordingly, the objective of this library is to contribute to LLM interpretability research, enabling users to explore visualization methods, such as heatmaps and graph representations of the hidden layers' activations in transformer-based LLMs.
This model allows users to explore questions such as:
- How do different categories of text in the corpus activate different neural regions?
- What are the differences between the properties of graphs formed by activations from two distinct categories?
- Are there regions of activation in the model more related to specific aspects of a category?
We encourage you to not only use this toolkit but also to extend it as you see fit.
Index
Online Example
The link below runs an online example of our library, in the Jupyter platform running over the Binder server:
Instalation
To see LLM-MRI in action on your own data:
pip install llm_mri
Usage
Firstly, the user needs to import the LLM-MRI and matplotlib.pyplot packages:
from llm_mri import LLM_MRI
import matplotlib.pyplot as plt
The user also needs to specify the Hugging Face Dataset that will be used to process the model's activations. There are two ways to do this:
- Load the Dataset from Hugging Face Hub:
dataset_url = "https://huggingface.co/datasets/dataset_link" dataset = load_dataset("csv", data_files=dataset_url) - If you already have the dataset loaded on your machine, you can use the load_from_disk function:
dataset = load_from_disk(dataset_path) # Specify the Dataset's path
Make sure that the selected dataset is a HuggingFace Dataset, and contains the columns "text" and "label", the last one being "ClassLabel" type. For more instructions on how to make this conversion, check out some of the examples on the GitHub documentation.
Next, the user selects the model to be used as a string:
model_ckpt = "distilbert/distilbert-base-multilingual-cased"
Then, the user instantiates LLM-MRI, to apply the methods defined on Functions:
llm_mri = LLM_MRI(model=model_ckpt, device="cpu", dataset=dataset)
For now, we recommend to use "cpu" as device. Further tests are going to be executed to ensure full "gpu" compatibility.
Functions
The library's functionality is divided into the following sections:
Activation Extraction:
As the user inputs the model and corpus to be analyzed, the dimensionality of the model's hidden layers is reduced, enabling visualization as an NxN grid.
llm_mri.process_activation_areas(map_dimension)
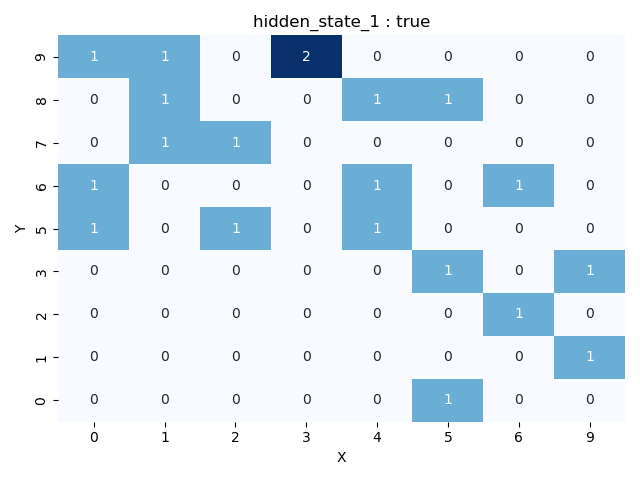
Heatmap representation of activations:
This includes the get_layer_image function, which transforms the NxN grid for a selected layer into a heatmap. In this heatmap, each cell represents the number of activations that different regions on a determined layer received for the provided corpus. Additionally, users can visualize activations for a specific label.
fig = llm_mri.get_layer_image(layer, category)
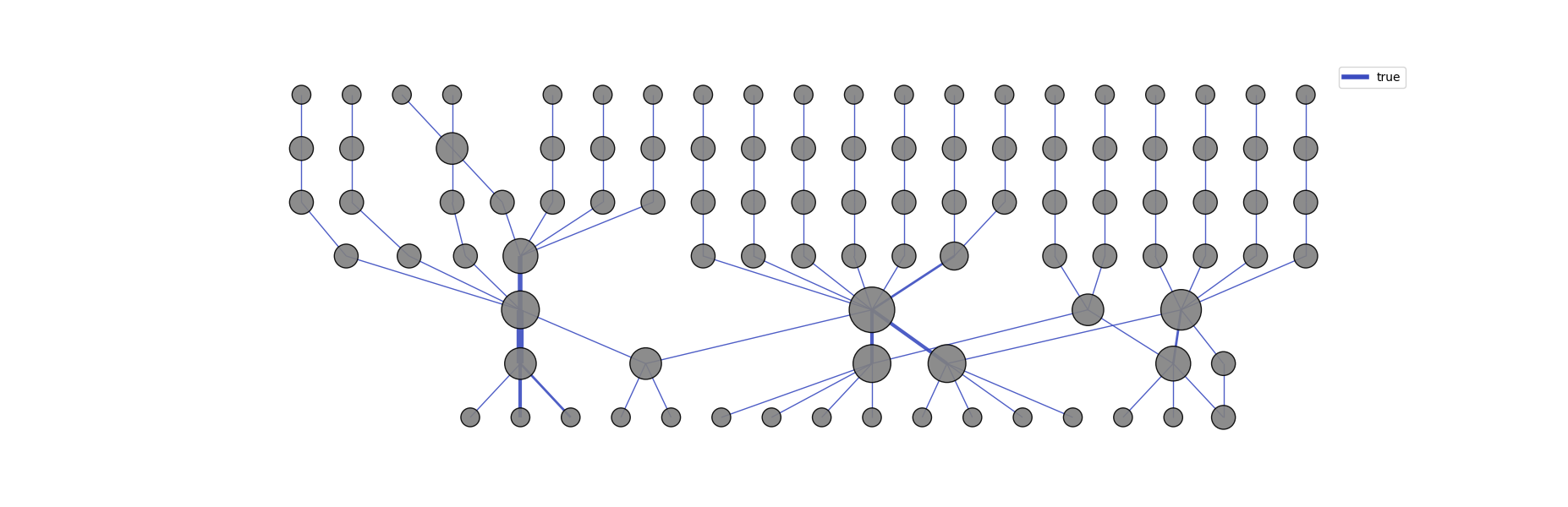
Graph Representation of Activations:
Using the get_graph function, the module connects regions from neighboring layers based on co-activations to form a graph representing the entire network. The graph's edges can also be colored according to different labels, allowing the user to identify the specific category that activated each neighboring node.
colormap: The default used colormap is the 'coolwarm'. More can be found on matplotlib.colors. We recommend the use of a 'Diverging' colormap for better visualization. fix_node_positions: 'True' keeps the nodes and edges at the same positions, independently of the categories. This could be useful for comparing activations between distinct categories. Setting to 'False' does not allow this comparison, although the graph will be more easily visualized.
graph = llm_mri.get_graph(category)
graph_image = llm_mri.get_graph_image(graph, colormap, fix_node_positions)
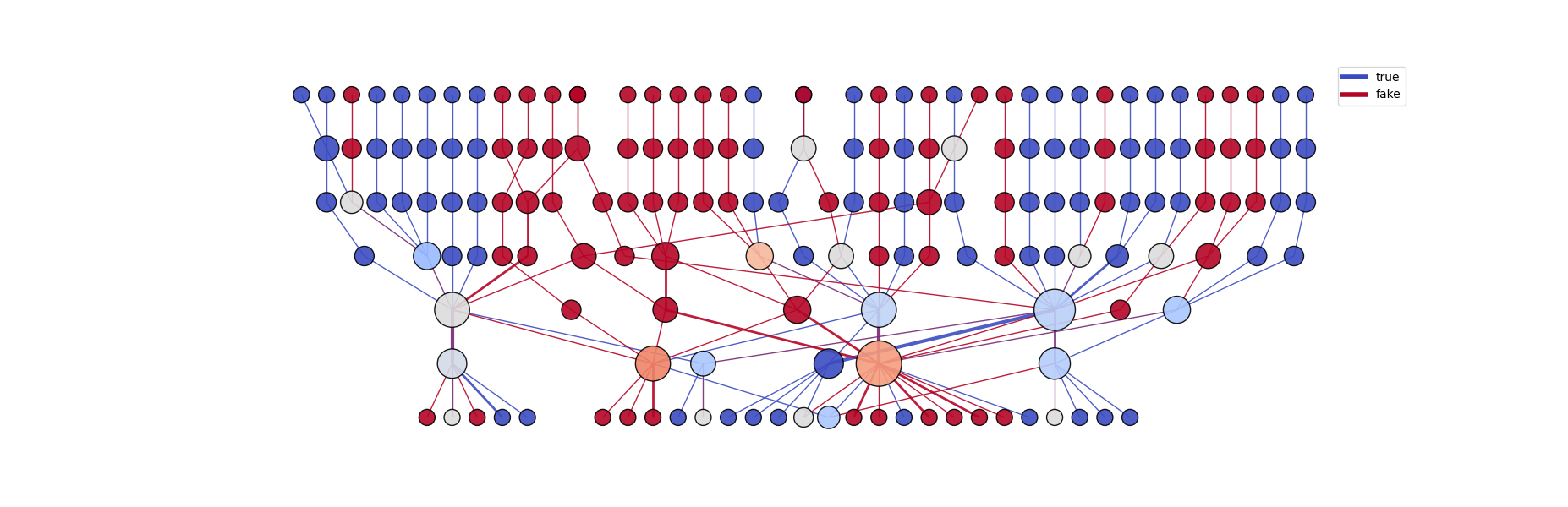
Composed Graph Visualization:
The user is also able to obtain a composed visualization of two different categories using the get_composed_graph function. By setting a category, each edge is colored based on the designated label, so the user is able to see which document label activated each region. Additionally, the user can select a colormap, where node colors reflect the label that most strongly activated them. Nodes colored white indicate equal activation by both categories, as white represents the midpoint of the color spectrum.
g_composed = llm_mri.get_composed_graph("true", "fake")
g_composed_img = llm_mri.get_graph_image(g_composed)
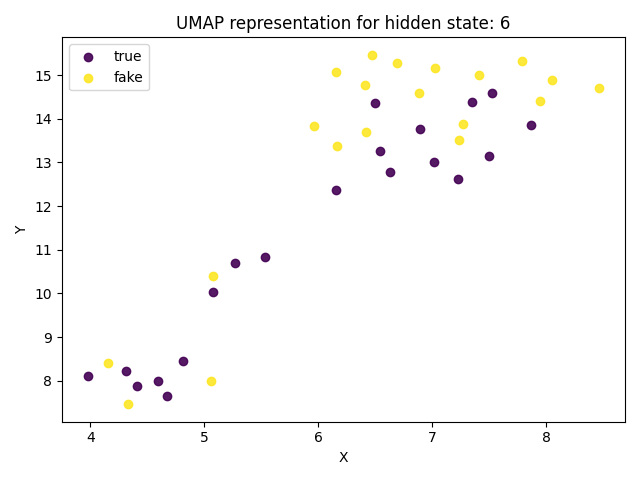
Reduced dimensionality representation of the documents
It is also possible to analyze where documents are disposed on a 2D space (after the dimensionality reduction) for a certain layer, divided by category. This could be essentialy more useful to visualize the impact of the DR algorithm on the activations disposal on the 2D space.
For comparison purpouses, this method differs from the heatmap representation of activations only by the cut of the 2D representation into a grid. With this approach, it is also posible to compare all documents on a single graph.
fig_scatter = llm_mri.get_original_map(layer=6)
Project details
Release history Release notifications | RSS feed
Download files
Download the file for your platform. If you're not sure which to choose, learn more about installing packages.
Source Distribution
Built Distribution
Filter files by name, interpreter, ABI, and platform.
If you're not sure about the file name format, learn more about wheel file names.
Copy a direct link to the current filters
File details
Details for the file llm_mri-0.1.10.tar.gz.
File metadata
- Download URL: llm_mri-0.1.10.tar.gz
- Upload date:
- Size: 15.7 kB
- Tags: Source
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.3 CPython/3.9.13 Linux/6.8.0-51-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
a6fd52f5dbc3e0480a49247ab0bac0f94fb18e975ee2c34b3a069eb309e0b299
|
|
| MD5 |
76661eec53fe677b7bcb200046e75843
|
|
| BLAKE2b-256 |
b4496e19849f6ef008304ec0f6edbaac1f243c6ad9d41e20423b363106c1df49
|
File details
Details for the file llm_mri-0.1.10-py3-none-any.whl.
File metadata
- Download URL: llm_mri-0.1.10-py3-none-any.whl
- Upload date:
- Size: 14.2 kB
- Tags: Python 3
- Uploaded using Trusted Publishing? No
- Uploaded via: poetry/1.8.3 CPython/3.9.13 Linux/6.8.0-51-generic
File hashes
| Algorithm | Hash digest | |
|---|---|---|
| SHA256 |
eb12051210a5613838bf0aa8270e727c917e1a73c4c1760769d96302a245fa90
|
|
| MD5 |
e283d2ce0f467549417fb33b03943a80
|
|
| BLAKE2b-256 |
12e9d12e50dc8f03b9b760a98b4f34984201e317de2421a2311ccf42e5b6474b
|